

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC136286.2 - phase: 0
(251 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
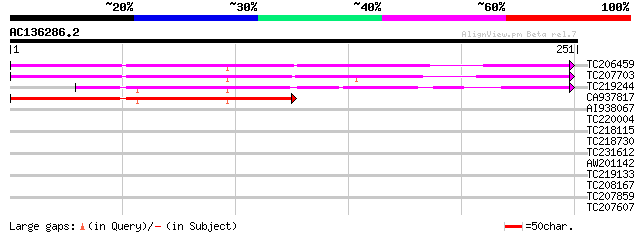
Sequences producing significant alignments: (bits) Value
TC206459 weakly similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial ... 195 2e-50
TC207703 similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (57%) 192 2e-49
TC219244 similar to GB|AAO11608.1|27363378|BT002692 At3g53950/F5... 154 5e-38
CA937817 131 3e-31
AI938067 similar to GP|9369366|gb|A F10A5.18 {Arabidopsis thalia... 35 0.025
TC220004 28 3.0
TC218115 similar to UP|Q94B60 (Q94B60) ATP-dependent Clp proteas... 28 4.0
TC218730 28 4.0
TC231612 28 5.2
AW201142 similar to PIR|T47568|T47 fructokinase-like protein - A... 27 6.7
TC219133 UP|Q39879 (Q39879) Mitotic cyclin a2-type, complete 27 6.7
TC208167 weakly similar to UP|Q9M6E7 (Q9M6E7) UDP-glucose:salicy... 27 6.7
TC207859 27 6.7
TC207607 similar to UP|Q9LJH9 (Q9LJH9) Gb|AAD25781.1, partial (20%) 27 8.8
>TC206459 weakly similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (37%)
Length = 970
Score = 195 bits (495), Expect = 2e-50
Identities = 111/251 (44%), Positives = 144/251 (57%), Gaps = 1/251 (0%)
Frame = +3
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
ME MP RVM DM+LLP G+V+I+NGAA GTAGWE NPVL P LYKP M+FE+
Sbjct: 120 METMPGARVMSDMVLLPNGDVLIVNGAAVGTAGWELGRNPVLNPFLYKPN-KRVGMRFEV 296
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDF-QAKYPTELSLDAYYPDYLRPELDT 119
P+ PRMYHS AVLL DGR+L+ GSNPH Y+F + +PTEL L+A+ P YL P
Sbjct: 297 QNPSHIPRMYHSGAVLLRDGRVLLAGSNPHTYYNFTKVLFPTELRLEAFSPWYLEPGFSN 476
Query: 120 LRPVIVAVEVVNSTLSYESLFSVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFLE 179
+RP IV+ + L Y + F + + + V+M+AP F THSF+MNQRLL L+
Sbjct: 477 VRPAIVS-PASQTKLKYGQTLRLRFKVSATLVGDSVSVTMLAPPFNTHSFSMNQRLLVLK 653
Query: 180 VTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIHV 239
L V S ++ V P S +APPG+Y+LFV+H
Sbjct: 654 PHHLSGV-----------------------GESTHEVEVTAPASAVLAPPGFYLLFVVHQ 764
Query: 240 GIPSVATWVHV 250
+PS WV +
Sbjct: 765 EVPSHGIWVQM 797
>TC207703 similar to UP|Q9LR03 (Q9LR03) F10A5.18, partial (57%)
Length = 1396
Score = 192 bits (487), Expect = 2e-49
Identities = 111/252 (44%), Positives = 144/252 (57%), Gaps = 2/252 (0%)
Frame = +3
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPFMKFEL 60
ME MP RVM DM+LLP G+V+I+NGAA GTAGWE NP+L P LYKP + +FE+
Sbjct: 510 METMPGARVMSDMVLLPNGDVLIVNGAAVGTAGWELGRNPLLSPFLYKPN-NRVGSRFEV 686
Query: 61 LAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDF-QAKYPTELSLDAYYPDYLRPELDT 119
+ PRMYHSSAVLL DGR+LV GSNPH Y F +PTEL L+A+ P YL P +
Sbjct: 687 QTSSDIPRMYHSSAVLLRDGRVLVAGSNPHIYYKFTNVLFPTELRLEAFSPWYLEPGFSS 866
Query: 120 LRPVIVAVEVVNSTLSYESLFSVSFLLREVKDV-NRIRVSMVAPSFTTHSFAMNQRLLFL 178
+RP IV + L Y + F + V + + V+M++P F THSF+MNQR+L L
Sbjct: 867 VRPTIV-FPASQTKLKYGQTLRLRFEVMSATLVGDSVSVTMLSPPFNTHSFSMNQRMLVL 1043
Query: 179 EVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVYKATVRGPPSLNVAPPGYYMLFVIH 238
E L K+ S Y+ V P S +APPG+Y+LF++H
Sbjct: 1044EPHDL-----------------------SKVGESTYEVEVTAPVSAVLAPPGFYLLFLVH 1154
Query: 239 VGIPSVATWVHV 250
IPS WV +
Sbjct: 1155QEIPSQGIWVQM 1190
>TC219244 similar to GB|AAO11608.1|27363378|BT002692 At3g53950/F5K20_250
{Arabidopsis thaliana;} , partial (34%)
Length = 858
Score = 154 bits (388), Expect = 5e-38
Identities = 91/223 (40%), Positives = 131/223 (57%), Gaps = 2/223 (0%)
Frame = +1
Query: 30 GTAGWENAANPVLYPVLYKPGLDNPF-MKFELLAPASTPRMYHSSAVLLPDGRILVGGSN 88
GT G+E A++P L PVLY+P D P ++F +L P + PRMYH++A LLPD R+L+ GSN
Sbjct: 1 GTQGFEMASDPCLNPVLYRP--DQPVGLRFMVLNPGTVPRMYHATANLLPDARVLLAGSN 174
Query: 89 PHRLYDF-QAKYPTELSLDAYYPDYLRPELDTLRPVIVAVEVVNSTLSYESLFSVSFLLR 147
PH LY F ++PTEL L+A+ P+YL + LRPVI E V T+ + F V +
Sbjct: 175 PHVLYRFDDVEFPTELRLEAFSPEYLSADRANLRPVI---EEVPQTVRFGGKFDVVVSV- 342
Query: 148 EVKDVNRIRVSMVAPSFTTHSFAMNQRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPG 207
++ V + V++ + F THSF+ QRL+ L V+ +++ D + G + G
Sbjct: 343 DLPVVGIVEVNLASAPFATHSFSQGQRLVKLTVS------SAVPDGSDGRYRIG------ 486
Query: 208 KIANSVYKATVRGPPSLNVAPPGYYMLFVIHVGIPSVATWVHV 250
V PPS VAPPGYYM F ++ G+PS+A W+HV
Sbjct: 487 ----------VTAPPSGAVAPPGYYMAFAVNQGVPSIAKWIHV 585
>CA937817
Length = 427
Score = 131 bits (330), Expect = 3e-31
Identities = 67/129 (51%), Positives = 93/129 (71%), Gaps = 2/129 (1%)
Frame = +1
Query: 1 MEVMPVPRVMPDMLLLPTGNVIILNGAANGTAGWENAANPVLYPVLYKPGLDNPF-MKFE 59
ME MP R+M DM++LP G+V+I+NGA +GT G+E A++P L PVLY+P D P ++F
Sbjct: 19 MEDMPFGRIMGDMVMLPNGDVLIINGAMSGTQGFEMASDPCLNPVLYRP--DQPVGLRFL 192
Query: 60 LLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDF-QAKYPTELSLDAYYPDYLRPELD 118
+L P + PRMYH++A LLPD R+L+ GSNPH LY F ++PTEL ++A+ P+YL +
Sbjct: 193 VLNPGTVPRMYHATANLLPDARVLLAGSNPHVLYRFNDVEFPTELRVEAFSPEYLSADRA 372
Query: 119 TLRPVIVAV 127
LRPVI V
Sbjct: 373 NLRPVIEEV 399
>AI938067 similar to GP|9369366|gb|A F10A5.18 {Arabidopsis thaliana}, partial
(4%)
Length = 196
Score = 35.4 bits (80), Expect = 0.025
Identities = 27/82 (32%), Positives = 33/82 (39%)
Frame = -1
Query: 155 IRVSMVAPSFTTHSFAMNQRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGPGKIANSVY 214
+ V+M+AP F HSF+M QRLL LE L V +
Sbjct: 178 VYVTMLAPPFNQHSFSMKQRLLVLEPHHLSGVGETRP----------------------- 68
Query: 215 KATVRGPPSLNVAPPGYYMLFV 236
K V P S + P G YM FV
Sbjct: 67 KVKVTAPASNVLGPRGNYMRFV 2
>TC220004
Length = 465
Score = 28.5 bits (62), Expect = 3.0
Identities = 29/102 (28%), Positives = 45/102 (43%), Gaps = 10/102 (9%)
Frame = +2
Query: 157 VSMVAPSFTTHS----------FAMNQRLLFLEVTALEEVVNSMQDQNFGEFGFGSSLGP 206
+S+ AP F+ S +NQRL VT NS+ +++ E
Sbjct: 89 LSIAAPKFSPSSSTIRCKGFSRLIVNQRL----VTVFATNPNSVGEKSPTE--------- 229
Query: 207 GKIANSVYKATVRGPPSLNVAPPGYYMLFVIHVGIPSVATWV 248
G N A +GPP L + G+++LFV+ I S+ TW+
Sbjct: 230 GSDVNETSNAA-QGPPLLTILA-GFFVLFVVCWSIWSIITWL 349
>TC218115 similar to UP|Q94B60 (Q94B60) ATP-dependent Clp protease-like
protein (At5g45390), partial (77%)
Length = 1087
Score = 28.1 bits (61), Expect = 4.0
Identities = 14/30 (46%), Positives = 16/30 (52%)
Frame = +2
Query: 50 GLDNPFMKFELLAPASTPRMYHSSAVLLPD 79
GL NP K +L P S P H A+L PD
Sbjct: 188 GLANPLQKPKLWVPVSEPYEPHLRALLAPD 277
>TC218730
Length = 747
Score = 28.1 bits (61), Expect = 4.0
Identities = 14/38 (36%), Positives = 18/38 (46%)
Frame = -1
Query: 90 HRLYDFQAKYPTELSLDAYYPDYLRPELDTLRPVIVAV 127
H Y F +P +LS Y P P L TL+P V +
Sbjct: 429 HNSYSFIPLFPPQLSTPLYGPLPQHPTLATLKPYFVGL 316
>TC231612
Length = 659
Score = 27.7 bits (60), Expect = 5.2
Identities = 14/29 (48%), Positives = 16/29 (54%)
Frame = -1
Query: 63 PASTPRMYHSSAVLLPDGRILVGGSNPHR 91
P TPR HS+ L RIL+ SN HR
Sbjct: 182 PVCTPRTCHSTHNLHSRPRILLAASNDHR 96
>AW201142 similar to PIR|T47568|T47 fructokinase-like protein - Arabidopsis
thaliana, partial (22%)
Length = 374
Score = 27.3 bits (59), Expect = 6.7
Identities = 13/45 (28%), Positives = 23/45 (50%)
Frame = -3
Query: 207 GKIANSVYKATVRGPPSLNVAPPGYYMLFVIHVGIPSVATWVHVH 251
GK+ K +RG P+ P +++ +HVG P A +++H
Sbjct: 204 GKVEFICKKRVLRGLPNRLHLHPPFFLKLHLHVGAPRPALRINLH 70
>TC219133 UP|Q39879 (Q39879) Mitotic cyclin a2-type, complete
Length = 1804
Score = 27.3 bits (59), Expect = 6.7
Identities = 31/108 (28%), Positives = 46/108 (41%), Gaps = 7/108 (6%)
Frame = +2
Query: 55 FMKFELLAPASTPRMYHSSAVLLPDGRILVGGSNPHRLYDFQAKYPTELSLDAYYPDYLR 114
F+KFE+ AP T + + V G V P + Y ELSL Y
Sbjct: 1070 FLKFEMTAP--TVKCFLRRFVRAAQGVDEV----PSLQLECLTNYIAELSLMEYSMLGYA 1231
Query: 115 PELDTLRPVIVAVEVV-------NSTLSYESLFSVSFLLREVKDVNRI 155
P L + +A ++ NSTL + +L+ S L VKD++R+
Sbjct: 1232 PSLVAASAIFLAKFILFPSKKPWNSTLQHYTLYQPSDLCVCVKDLHRL 1375
>TC208167 weakly similar to UP|Q9M6E7 (Q9M6E7) UDP-glucose:salicylic acid
glucosyltransferase, partial (23%)
Length = 890
Score = 27.3 bits (59), Expect = 6.7
Identities = 16/32 (50%), Positives = 19/32 (59%)
Frame = -2
Query: 214 YKATVRGPPSLNVAPPGYYMLFVIHVGIPSVA 245
Y A VRG S+N YMLFV+HV + VA
Sbjct: 523 YSACVRGN-SINFL*TWKYMLFVVHV*VKKVA 431
>TC207859
Length = 751
Score = 27.3 bits (59), Expect = 6.7
Identities = 17/47 (36%), Positives = 22/47 (46%)
Frame = -1
Query: 87 SNPHRLYDFQAKYPTELSLDAYYPDYLRPELDTLRPVIVAVEVVNST 133
+NP L YP LS D +PD R L LR A++ NS+
Sbjct: 427 TNPLDLCPAPIIYPMRLSFDLLFPDRARCCLTNLRFTFSALKHANSS 287
>TC207607 similar to UP|Q9LJH9 (Q9LJH9) Gb|AAD25781.1, partial (20%)
Length = 1260
Score = 26.9 bits (58), Expect = 8.8
Identities = 25/88 (28%), Positives = 38/88 (42%)
Frame = -1
Query: 141 SVSFLLREVKDVNRIRVSMVAPSFTTHSFAMNQRLLFLEVTALEEVVNSMQDQNFGEFGF 200
++S LL+ ++ S PSF++ S + R F EEV NS+
Sbjct: 477 NMS*LLKLQNSISLHSTSPFNPSFSSPSQVLGNRNPF------EEVQNSIL--------- 343
Query: 201 GSSLGPGKIANSVYKATVRGPPSLNVAP 228
++ N V +R PP+LNVAP
Sbjct: 342 -------QVLNRVLDVILRMPPTLNVAP 280
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,242,712
Number of Sequences: 63676
Number of extensions: 134201
Number of successful extensions: 634
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 618
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 619
length of query: 251
length of database: 12,639,632
effective HSP length: 95
effective length of query: 156
effective length of database: 6,590,412
effective search space: 1028104272
effective search space used: 1028104272
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 58 (26.9 bits)
Medicago: description of AC136286.2


