

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135848.1 + phase: 0
(64 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
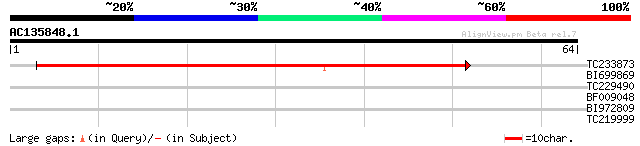
Score E
Sequences producing significant alignments: (bits) Value
TC233873 weakly similar to UP|Q6ZJC8 (Q6ZJC8) BHLH protein famil... 46 3e-06
BI699869 38 0.001
TC229490 similar to UP|Q8S9M5 (Q8S9M5) At2g41130/T3K9.10, partia... 36 0.004
BF009048 25 6.0
BI972809 25 6.0
TC219999 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (5%) 25 7.9
>TC233873 weakly similar to UP|Q6ZJC8 (Q6ZJC8) BHLH protein family-like,
partial (25%)
Length = 817
Score = 46.2 bits (108), Expect = 3e-06
Identities = 24/50 (48%), Positives = 37/50 (74%), Gaps = 1/50 (2%)
Frame = +3
Query: 4 ISRALLSMKAKVVKVEMVTVGGRTRSVLWVQ-GVENESLGMLKSTLKVVM 52
+S+A+ S++AK V+ EM+TVGGRT+SV+ V+ E E +G L+ LK V+
Sbjct: 483 VSQAIRSVRAKAVRAEMMTVGGRTKSVVVVEWEKEEEEVGALERALKAVV 632
>BI699869
Length = 421
Score = 37.7 bits (86), Expect = 0.001
Identities = 21/52 (40%), Positives = 34/52 (65%), Gaps = 3/52 (5%)
Frame = +1
Query: 8 LLSMKAKVVKVEMVTVGGRTRSVLWV---QGVENESLGMLKSTLKVVMHKPS 56
L S++ K +K EM T+GGRTR+VL V + ES+ L+++LK ++ + S
Sbjct: 31 LRSLRLKTLKAEMATLGGRTRNVLVVATDKDHSGESIQFLQNSLKSLVERSS 186
>TC229490 similar to UP|Q8S9M5 (Q8S9M5) At2g41130/T3K9.10, partial (56%)
Length = 1006
Score = 35.8 bits (81), Expect = 0.004
Identities = 20/52 (38%), Positives = 33/52 (63%), Gaps = 3/52 (5%)
Frame = +1
Query: 8 LLSMKAKVVKVEMVTVGGRTRSVLWVQGVEN---ESLGMLKSTLKVVMHKPS 56
L S+ K +K EM T+GGRTR+VL V + ES+ L+++L+ ++ + S
Sbjct: 661 LNSLHLKTLKAEMATLGGRTRNVLVVAADKEHSIESIHFLQNSLRSILDRSS 816
>BF009048
Length = 421
Score = 25.4 bits (54), Expect = 6.0
Identities = 10/36 (27%), Positives = 21/36 (57%), Gaps = 1/36 (2%)
Frame = +3
Query: 25 GRTRSV-LWVQGVENESLGMLKSTLKVVMHKPSFKM 59
GR + + LW+ ++ E+ K +L +V+H P+ +
Sbjct: 96 GREKPIKLWLLSIQREARATKKQSLGIVLHLPTHSL 203
>BI972809
Length = 286
Score = 25.4 bits (54), Expect = 6.0
Identities = 12/29 (41%), Positives = 18/29 (61%)
Frame = -1
Query: 12 KAKVVKVEMVTVGGRTRSVLWVQGVENES 40
+ +VV E+ T+GG + S WV G +ES
Sbjct: 262 RRRVVSSEVKTIGGSSSS*KWV*GNSSES 176
>TC219999 weakly similar to UP|Q7T9Y7 (Q7T9Y7) Odv-e66, partial (5%)
Length = 641
Score = 25.0 bits (53), Expect = 7.9
Identities = 9/20 (45%), Positives = 12/20 (60%)
Frame = +2
Query: 23 VGGRTRSVLWVQGVENESLG 42
VGGR +W +GVE +G
Sbjct: 278 VGGRRERNVWTEGVEGSGMG 337
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.129 0.343
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,022,456
Number of Sequences: 63676
Number of extensions: 13926
Number of successful extensions: 75
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 75
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 75
length of query: 64
length of database: 12,639,632
effective HSP length: 40
effective length of query: 24
effective length of database: 10,092,592
effective search space: 242222208
effective search space used: 242222208
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC135848.1


