

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135801.5 - phase: 0
(736 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
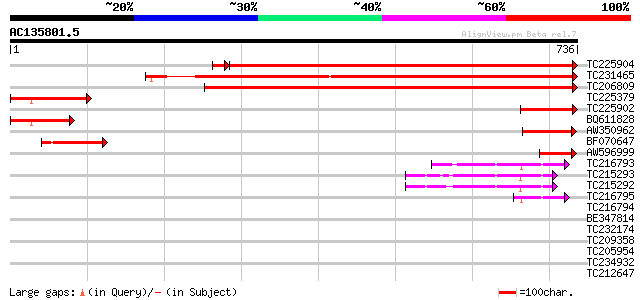
Sequences producing significant alignments: (bits) Value
TC225904 similar to UP|O48937 (O48937) NADPH cytochrome P450 red... 826 0.0
TC231465 UP|Q8GUS1 (Q8GUS1) NADPH:P450 reductase, partial (77%) 763 0.0
TC206809 homologue to UP|NCPR_PHAAU (P37116) NADPH--cytochrome P... 726 0.0
TC225379 similar to UP|Q9AU06 (Q9AU06) NADPH-cytochrome P450 oxy... 165 6e-41
TC225902 similar to UP|O48937 (O48937) NADPH cytochrome P450 red... 133 2e-31
BQ611828 weakly similar to PIR|JE0230|JE02 NADPH-cytochrome P450... 123 3e-28
AW350962 homologue to GP|27261128|gb| NADPH:P450 reductase {Glyc... 110 2e-24
BF070647 homologue to PIR|A47298|A47 NADPH-ferrihemoprotein redu... 89 9e-18
AW596999 73 5e-13
TC216793 similar to UP|FENS_PEA (Q41014) Ferredoxin--NADP reduct... 71 2e-12
TC215293 similar to UP|FENR_TOBAC (O04977) Ferredoxin--NADP redu... 63 5e-10
TC215292 similar to UP|FENR_PEA (P10933) Ferredoxin--NADP reduct... 57 3e-08
TC216795 homologue to UP|FENS_PEA (Q41014) Ferredoxin--NADP redu... 46 5e-05
TC216794 homologue to UP|FENS_PEA (Q41014) Ferredoxin--NADP redu... 41 0.002
BE347814 similar to SP|P41346|FENR Ferredoxin--NADP reductase c... 40 0.005
TC232174 36 0.054
TC209358 similar to UP|P92987 (P92987) Myosin heavy chain-like p... 33 0.59
TC205954 similar to UP|Q9FK50 (Q9FK50) Brn1-like protein, partia... 32 1.0
TC234932 31 2.3
TC212647 weakly similar to UP|Q8L9A6 (Q8L9A6) Thioredoxin, parti... 31 2.3
>TC225904 similar to UP|O48937 (O48937) NADPH cytochrome P450 reductase ,
partial (68%)
Length = 1704
Score = 826 bits (2134), Expect(2) = 0.0
Identities = 401/451 (88%), Positives = 428/451 (93%)
Frame = +1
Query: 286 LRDEDDATVATPYTASVLEYRVVIRDQLDATVDEKKQLNGNGHAVVDAHHPVRANVAVRK 345
LRDEDDATV+TPYTA+VLEYRVVI D L+A+VDEKK N NGHA+VDA HPVRANVAVRK
Sbjct: 70 LRDEDDATVSTPYTAAVLEYRVVIHDPLEASVDEKKWHNVNGHAIVDAQHPVRANVAVRK 249
Query: 346 ELHTPASDRSCTHLEFDISGTGVVYETGDHVGVYCENLSDTVEEAERILGLSPDTYFSVH 405
ELHTPASDRSCTHLEFDISGTGV YETGDHVGVYCENLS+TVEEA R++GLSPDTYFS+H
Sbjct: 250 ELHTPASDRSCTHLEFDISGTGVTYETGDHVGVYCENLSETVEEAIRLIGLSPDTYFSIH 429
Query: 406 TDDEDGKPLGGSSLPPPFPPCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRH 465
TDDEDGKP GSSLPP FPPCTLR AL +YADVLSSPKKSALLALAAHASDPSEADRLRH
Sbjct: 430 TDDEDGKPRSGSSLPPTFPPCTLRKALTQYADVLSSPKKSALLALAAHASDPSEADRLRH 609
Query: 466 LASPAGKDEYAEWVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPRVA 525
LASPAGKDEY+EWVIASQRSLLEVMAEF SAKPPIGVFFA+VAPRLQPR+YSISSSPR+
Sbjct: 610 LASPAGKDEYSEWVIASQRSLLEVMAEFPSAKPPIGVFFAAVAPRLQPRFYSISSSPRMV 789
Query: 526 PSRIHVTCALVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNK 585
P+RIHVTCALVH+KMPTGRIH+GVCSTWMKNS PLEKSQDCSWAPIFVR SNFRLP+DNK
Sbjct: 790 PNRIHVTCALVHEKMPTGRIHKGVCSTWMKNSVPLEKSQDCSWAPIFVRTSNFRLPSDNK 969
Query: 586 VPIIMIGPGTGLAPFRGFLQERLALKEEGAELGPSVLFFGCRNRQVDYIYEDELNHFVHG 645
VPIIMIGPGTGLAPFRGFLQERLALKE GAELGPSVLFFGCRNRQ+DYIYEDEL+HFV+
Sbjct: 970 VPIIMIGPGTGLAPFRGFLQERLALKEGGAELGPSVLFFGCRNRQMDYIYEDELSHFVNT 1149
Query: 646 GALSELIVAFSREGPTKEYVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHT 705
GAL ELI+AFSREGPTKEYVQHKM+EKAS+IW+MISQGAYIYVCGDAKGMA+DVHR LHT
Sbjct: 1150GALDELILAFSREGPTKEYVQHKMMEKASEIWSMISQGAYIYVCGDAKGMARDVHRALHT 1329
Query: 706 ILQEQGSLDNSKTESMVKNLQMTGRYLRDVW 736
ILQEQGSLD+SK ESMVKNLQ TGRYLRDVW
Sbjct: 1330ILQEQGSLDSSKAESMVKNLQTTGRYLRDVW 1422
Score = 52.4 bits (124), Expect(2) = 0.0
Identities = 21/22 (95%), Positives = 22/22 (99%)
Frame = +3
Query: 264 DQCIEDDFTAWKEELWPALDQL 285
DQCIEDDFTAWKEELWPALD+L
Sbjct: 3 DQCIEDDFTAWKEELWPALDEL 68
>TC231465 UP|Q8GUS1 (Q8GUS1) NADPH:P450 reductase, partial (77%)
Length = 1933
Score = 763 bits (1969), Expect = 0.0
Identities = 371/565 (65%), Positives = 447/565 (78%), Gaps = 5/565 (0%)
Frame = +1
Query: 177 EGEEDS---FKNLQYGVFGLGNRQYEHFNKVCTQALMHSCGFLSFDYSDGVASFRAFSDI 233
EG+++ + L YGVFGLGNRQYEHFNK+
Sbjct: 7 EGKDERGIWLQQLTYGVFGLGNRQYEHFNKI----------------------------- 99
Query: 234 YHSTQVAKIVDDKLLEKGGNRLVPVGLGDDDQCIEDDFTAWKEELWPALDQLLRDEDDA- 292
KIVD++L E+G RLVP+GLGDDDQ IEDDF AWKE LW LDQLLRDEDD
Sbjct: 100 ------GKIVDEELSEQGAKRLVPLGLGDDDQSIEDDFVAWKESLWSELDQLLRDEDDVN 261
Query: 293 TVATPYTASVLEYRVVIRDQLDATVDEKKQLNGNGHAVVDAHHPVRANVAVRKELHTPAS 352
TV+TPY A++ EYRVVI D + ++ NG+AV D HHP R N+A ++ELH P S
Sbjct: 262 TVSTPYKAAIPEYRVVIHDSTVTSCNDNHLNVANGNAVFDIHHPCRVNIAAQRELHKPES 441
Query: 353 DRSCTHLEFDISGTGVVYETGDHVGVYCENLSDTVEEAERILGLSPDTYFSVHTDDEDGK 412
DRSC HLEFDISGTG++YETGDHVGV+ EN +TVEEA ++LG D FS+HT++EDG
Sbjct: 442 DRSCIHLEFDISGTGIIYETGDHVGVFAENGDETVEEAGKLLGQDLDLVFSIHTNNEDGT 621
Query: 413 PLGGSSLPPPFP-PCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLASPAG 471
PLG SSLPPPFP PCTLR ALA YAD+L+ P+K++L+ALAAH S+PSEADRL L+SP G
Sbjct: 622 PLG-SSLPPPFPGPCTLRFALAHYADLLNPPRKASLVALAAHTSEPSEADRLTFLSSPQG 798
Query: 472 KDEYAEWVIASQRSLLEVMAEFSSAKPPIGVFFASVAPRLQPRYYSISSSPRVAPSRIHV 531
KDEY++W++ SQRSLLEVMAEF SAKPP+GVFFA+VAP LQPRYYSISSSPR +P ++HV
Sbjct: 799 KDEYSKWLVGSQRSLLEVMAEFPSAKPPLGVFFAAVAPHLQPRYYSISSSPRFSPQKVHV 978
Query: 532 TCALVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPIIMI 591
TCALV PTGRIH+GVCSTWMKN+ PLEKS+DCSWAPIF+R SNF+LPAD+ +PIIM+
Sbjct: 979 TCALVCGPTPTGRIHKGVCSTWMKNAIPLEKSRDCSWAPIFIRTSNFKLPADHSIPIIMV 1158
Query: 592 GPGTGLAPFRGFLQERLALKEEGAELGPSVLFFGCRNRQVDYIYEDELNHFVHGGALSEL 651
GPGTGLAPFRGFLQERLALKE+ +LGP++LFFGCRNRQ+D+IYEDEL +F+ GALSEL
Sbjct: 1159GPGTGLAPFRGFLQERLALKEDAVQLGPALLFFGCRNRQMDFIYEDELKNFMEQGALSEL 1338
Query: 652 IVAFSREGPTKEYVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQG 711
IV FSREGP KEYVQHKM++KA+++WN+ISQG Y+YVCGDAKGMA+DVHRTLHTI+Q+Q
Sbjct: 1339IVTFSREGPEKEYVQHKMMDKAANLWNLISQGGYLYVCGDAKGMARDVHRTLHTIVQQQE 1518
Query: 712 SLDNSKTESMVKNLQMTGRYLRDVW 736
++D+SK E++VK LQM GRYLRDVW
Sbjct: 1519NVDSSKAEAIVKKLQMDGRYLRDVW 1593
>TC206809 homologue to UP|NCPR_PHAAU (P37116) NADPH--cytochrome P450
reductase (CPR) (P450R) , partial (70%)
Length = 1867
Score = 726 bits (1874), Expect = 0.0
Identities = 344/484 (71%), Positives = 411/484 (84%), Gaps = 1/484 (0%)
Frame = +2
Query: 254 RLVPVGLGDDDQCIEDDFTAWKEELWPALDQLLRDEDDA-TVATPYTASVLEYRVVIRDQ 312
RLV GLGDDDQ IEDDF+AWKE LWP LDQLLRDEDDA TV+TPYTA++LEYRVVI D
Sbjct: 5 RLVTFGLGDDDQSIEDDFSAWKETLWPELDQLLRDEDDANTVSTPYTAAILEYRVVIHDP 184
Query: 313 LDATVDEKKQLNGNGHAVVDAHHPVRANVAVRKELHTPASDRSCTHLEFDISGTGVVYET 372
+ + NG+AV D HHP R NVAV++ELH P SDRSC HLEFDISGTG+ YET
Sbjct: 185 TFTSSYDNHVTVANGNAVFDIHHPCRVNVAVQRELHKPESDRSCIHLEFDISGTGLTYET 364
Query: 373 GDHVGVYCENLSDTVEEAERILGLSPDTYFSVHTDDEDGKPLGGSSLPPPFPPCTLRTAL 432
GDHVGVY +N +TVEE ++LG + D FS+HTD EDG LGGS LPP PCTLRTAL
Sbjct: 365 GDHVGVYADNCDETVEETGKLLGQNLDLLFSLHTDKEDGTSLGGSLLPPFPGPCTLRTAL 544
Query: 433 AKYADVLSSPKKSALLALAAHASDPSEADRLRHLASPAGKDEYAEWVIASQRSLLEVMAE 492
A+YAD+L+ P+K+AL+ALA+HAS+PSEA+RL+ L+SP GKDEY++WV+ SQRSLLEVMAE
Sbjct: 545 ARYADLLNPPRKAALVALASHASEPSEAERLKFLSSPQGKDEYSKWVVGSQRSLLEVMAE 724
Query: 493 FSSAKPPIGVFFASVAPRLQPRYYSISSSPRVAPSRIHVTCALVHDKMPTGRIHQGVCST 552
F SAKPP+GVFFA+VAPRLQPRYYSISSSPR AP R+HVTCALV+ PTGRIH+GVCST
Sbjct: 725 FPSAKPPLGVFFAAVAPRLQPRYYSISSSPRFAPQRVHVTCALVYGPTPTGRIHKGVCST 904
Query: 553 WMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRGFLQERLALKE 612
WMKN+ PLEKS DCSWAPIF+R SNF+LP D+ +PIIM+GPGTGLAPFRGFLQER ALKE
Sbjct: 905 WMKNAIPLEKSPDCSWAPIFIRPSNFKLPVDHSIPIIMVGPGTGLAPFRGFLQERFALKE 1084
Query: 613 EGAELGPSVLFFGCRNRQVDYIYEDELNHFVHGGALSELIVAFSREGPTKEYVQHKMIEK 672
G + GP++LFFGCRNR++D+IYE+EL +FV G+LSELIVAFSREG KEYVQHKM+++
Sbjct: 1085AGVQQGPAILFFGCRNRRLDFIYEEELKNFVEQGSLSELIVAFSREGAEKEYVQHKMMDQ 1264
Query: 673 ASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTESMVKNLQMTGRYL 732
A+ +W++ISQG Y+YVCGDAKGMA+DVHRTLHTI+Q+Q ++D++K E++VK LQM GRYL
Sbjct: 1265AAQLWSLISQGGYLYVCGDAKGMARDVHRTLHTIVQQQENVDSTKAEAIVKKLQMDGRYL 1444
Query: 733 RDVW 736
RDVW
Sbjct: 1445RDVW 1456
>TC225379 similar to UP|Q9AU06 (Q9AU06) NADPH-cytochrome P450 oxydoreductase
isoform 3, partial (13%)
Length = 737
Score = 165 bits (418), Expect = 6e-41
Identities = 81/112 (72%), Positives = 95/112 (84%), Gaps = 6/112 (5%)
Frame = +1
Query: 1 MQDSSSMKFSPLDLMSALIKGKIDPSN------GTVPASLILENREFVMILTTSIAVLIG 54
MQDSSSMK SPLDLMSA+IKGK+DPSN A+ +LENR+F+M+LTTS+AVLIG
Sbjct: 400 MQDSSSMKISPLDLMSAIIKGKLDPSNVSSSTNAAAAAAFLLENRDFLMLLTTSVAVLIG 579
Query: 55 CVVVLIWRRSNSQKPKPIEVPKRVIEKLPELEIDDGTKKVTVFFGTQTGTAE 106
CVVV IWRRS+S K KP+E PKRV+EKLPE+E+DDGTKKVT+ FGTQTGTAE
Sbjct: 580 CVVVFIWRRSSSPKAKPLEPPKRVVEKLPEIEVDDGTKKVTILFGTQTGTAE 735
Score = 29.3 bits (64), Expect = 6.6
Identities = 16/53 (30%), Positives = 25/53 (46%)
Frame = +2
Query: 420 PPPFPPCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLASPAGK 472
PPP PP + RTA + + SP SA + ++ A P L L + + +
Sbjct: 500 PPPPPPSSSRTATSSCSSPPPSPSSSAASSSSSGADPPPPRPSLSSLLNASSR 658
>TC225902 similar to UP|O48937 (O48937) NADPH cytochrome P450 reductase ,
partial (11%)
Length = 581
Score = 133 bits (335), Expect = 2e-31
Identities = 62/73 (84%), Positives = 68/73 (92%)
Frame = +2
Query: 664 YVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTESMVK 723
YVQHKM+EKAS+IW+MIS GAYIYVCGDAKGMA++VHR LHTILQEQGSLD+SK ESMV
Sbjct: 2 YVQHKMMEKASEIWSMISXGAYIYVCGDAKGMARNVHRALHTILQEQGSLDSSKAESMVN 181
Query: 724 NLQMTGRYLRDVW 736
NLQ TGRYLRDVW
Sbjct: 182 NLQTTGRYLRDVW 220
>BQ611828 weakly similar to PIR|JE0230|JE02 NADPH-cytochrome P450
oxidoreductase (EC 1.-.-.-) - common tobacco, partial
(7%)
Length = 308
Score = 123 bits (309), Expect = 3e-28
Identities = 61/89 (68%), Positives = 73/89 (81%), Gaps = 5/89 (5%)
Frame = +2
Query: 1 MQDSSSMKFSPLDLMSALIKGKIDPSN-----GTVPASLILENREFVMILTTSIAVLIGC 55
MQDS SMK SPLDLMSA+IKGK+DPSN A+++LENR+F+M+LTTS+AVL+GC
Sbjct: 41 MQDSGSMKISPLDLMSAIIKGKLDPSNVSSSSSNAAAAVLLENRQFLMLLTTSVAVLVGC 220
Query: 56 VVVLIWRRSNSQKPKPIEVPKRVIEKLPE 84
VV IWRRS+S K KP+E PKRVIEKLPE
Sbjct: 221 FVVFIWRRSSSPKAKPLEPPKRVIEKLPE 307
>AW350962 homologue to GP|27261128|gb| NADPH:P450 reductase {Glycine max},
partial (10%)
Length = 519
Score = 110 bits (275), Expect = 2e-24
Identities = 49/70 (70%), Positives = 62/70 (88%)
Frame = -2
Query: 666 QHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTESMVKNL 725
QHKM++KA+++WN+ISQG Y+YVCGDAKGMA DV RTLHTI+Q+Q ++D+SK E +VK L
Sbjct: 518 QHKMMDKAANLWNLISQGGYLYVCGDAKGMAXDVXRTLHTIVQQQENVDSSKAEGIVKKL 339
Query: 726 QMTGRYLRDV 735
QM GRYLRDV
Sbjct: 338 QMDGRYLRDV 309
>BF070647 homologue to PIR|A47298|A47 NADPH-ferrihemoprotein reductase (EC
1.6.2.4) - mung bean, partial (14%)
Length = 340
Score = 88.6 bits (218), Expect = 9e-18
Identities = 48/89 (53%), Positives = 69/89 (76%), Gaps = 3/89 (3%)
Frame = +3
Query: 42 VMILTTSIAVLIGCVVVLIWRRSN--SQKPKPIEVPKRVI-EKLPELEIDDGTKKVTVFF 98
++I TTS A++IG ++V +W++S+ S++ KP+ VPK + + E+++ DG KVT+FF
Sbjct: 75 LLIATTSAALIIG-LLVFLWKKSSDRSKELKPVIVPKGLPKDDDDEVDVADGKTKVTIFF 251
Query: 99 GTQTGTAEGFAKAIAEEAKARYEKAKFKV 127
GTQTGTAEGFAKA+AEE KARY+KA KV
Sbjct: 252 GTQTGTAEGFAKALAEEIKARYDKAAVKV 338
>AW596999
Length = 145
Score = 72.8 bits (177), Expect = 5e-13
Identities = 37/48 (77%), Positives = 39/48 (81%)
Frame = -1
Query: 688 VCGDAKGMAKDVHRTLHTILQEQGSLDNSKTESMVKNLQMTGRYLRDV 735
VCGDAKGMA+D HR TILQEQGSLD+SK ESMV LQ TGRYL DV
Sbjct: 145 VCGDAKGMARDDHRACTTILQEQGSLDSSKAESMV*YLQTTGRYLPDV 2
>TC216793 similar to UP|FENS_PEA (Q41014) Ferredoxin--NADP reductase, root
isozyme, chloroplast precursor (FNR) , partial (97%)
Length = 1408
Score = 71.2 bits (173), Expect = 2e-12
Identities = 57/186 (30%), Positives = 90/186 (47%), Gaps = 7/186 (3%)
Frame = +3
Query: 548 GVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADN-KVPIIMIGPGTGLAPFRGFLQE 606
G+CS ++ NS P +K Q + + LP D+ IMI GTG+APFRG+L+
Sbjct: 783 GICSNFLCNSKPGDKIQITGPSGKIML-----LPEDDPNATHIMIATGTGVAPFRGYLRR 947
Query: 607 RLALKEEGAELGPSV-LFFGCRNRQVDYIYEDELNHFVHGGALS-ELIVAFSREGPTKE- 663
+ G LF G N +Y++E + +++ + + A SRE K
Sbjct: 948 MFMESVPTYKFGGLAWLFLGVANTD-SLLYDEEFSKYLNDYSDNFRYDRALSREQKNKNG 1124
Query: 664 ---YVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQEQGSLDNSKTES 720
YVQ K+ E + +I+ ++ GA+IY CG KGM + TL + +++G K
Sbjct: 1125GKMYVQDKIEEYSDEIFKLLDNGAHIYFCG-LKGMMPGIQDTLKKVAEQRGESWEEKLSQ 1301
Query: 721 MVKNLQ 726
+ KN Q
Sbjct: 1302LKKNKQ 1319
>TC215293 similar to UP|FENR_TOBAC (O04977) Ferredoxin--NADP reductase,
leaf-type isozyme, chloroplast precursor (FNR) ,
complete
Length = 1488
Score = 62.8 bits (151), Expect = 5e-10
Identities = 60/209 (28%), Positives = 96/209 (45%), Gaps = 11/209 (5%)
Frame = +1
Query: 514 RYYSISSSP--RVAPSRIHVTCA--LVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWA 569
R YSI+SS S+ C LV+ G I +GVCS ++ + P ++
Sbjct: 559 RLYSIASSAIGDFGDSKTVSLCVKRLVYTN-ENGEIVKGVCSNFLCDLKP--GAEVTITG 729
Query: 570 PIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRGFLQERLALKEEGAEL-GPSVLFFGCRN 628
P+ +P D IIM+G GTG+APFR FL + K E + G + LF G
Sbjct: 730 PV---GKEMLMPKDPNATIIMLGTGTGIAPFRSFLWKMFFEKHEDYKFNGLAWLFLGVPT 900
Query: 629 RQVDYIYEDELNHFVHGGALS-ELIVAFSREGPT----KEYVQHKMIEKASDIWNMISQ- 682
+Y++E + L A SRE K Y+Q +M + A ++W ++ +
Sbjct: 901 SS-SLLYKEEFEKMQEKSPENFRLDFAVSREQTNEKGEKMYIQTRMAQYAEELWELLKKD 1077
Query: 683 GAYIYVCGDAKGMAKDVHRTLHTILQEQG 711
++Y+CG KGM K + + ++ + G
Sbjct: 1078NTFVYMCG-LKGMEKGIDDIMVSLAAKDG 1161
>TC215292 similar to UP|FENR_PEA (P10933) Ferredoxin--NADP reductase, leaf
isozyme, chloroplast precursor (FNR) , complete
Length = 1435
Score = 57.0 bits (136), Expect = 3e-08
Identities = 56/209 (26%), Positives = 95/209 (44%), Gaps = 11/209 (5%)
Frame = +2
Query: 514 RYYSISSSPR--VAPSRIHVTCA--LVHDKMPTGRIHQGVCSTWMKNSAPLEKSQDCSWA 569
R YSI+SS S+ C LV+ G + +GVCS ++ + P + +
Sbjct: 551 RLYSIASSALGDFGDSKTVSLCVKRLVYTN-ENGELVKGVCSNFLCDLKPGAEVKITG-- 721
Query: 570 PIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRGFLQERLALKEEGAEL-GPSVLFFGCRN 628
P+ +P D +IM+ GTG+APFR FL + K + + G + LF G
Sbjct: 722 PV---GKEMLMPKDPNATVIMLATGTGIAPFRSFLWKMFFEKHDDYKFNGLAWLFLGVPT 892
Query: 629 RQVDYIYEDELNHFVHGGALS-ELIVAFSREGPT----KEYVQHKMIEKASDIWNMISQ- 682
+Y++E + L A SRE K Y+Q +M + A ++W ++ +
Sbjct: 893 SS-SLLYKEEFEKMKEKAPDNFRLDFAVSREQTNEKGEKMYIQTRMAQYAEELWELLKKD 1069
Query: 683 GAYIYVCGDAKGMAKDVHRTLHTILQEQG 711
++Y+CG KGM K + + ++ + G
Sbjct: 1070NTFVYMCG-LKGMEKGIDDIMVSLAAKDG 1153
>TC216795 homologue to UP|FENS_PEA (Q41014) Ferredoxin--NADP reductase, root
isozyme, chloroplast precursor (FNR) , partial (34%)
Length = 764
Score = 46.2 bits (108), Expect = 5e-05
Identities = 27/77 (35%), Positives = 40/77 (51%), Gaps = 4/77 (5%)
Frame = +1
Query: 654 AFSREGPTKE----YVQHKMIEKASDIWNMISQGAYIYVCGDAKGMAKDVHRTLHTILQE 709
A SRE K YVQ K+ E + +I+ ++ GA+IY CG KGM + TL + ++
Sbjct: 172 ALSREQKNKSGGKMYVQDKIEEYSDEIFKLLDNGAHIYFCG-LKGMMPGIQDTLKKVAEQ 348
Query: 710 QGSLDNSKTESMVKNLQ 726
+G K + KN Q
Sbjct: 349 RGESWEEKLSQLKKNKQ 399
>TC216794 homologue to UP|FENS_PEA (Q41014) Ferredoxin--NADP reductase, root
isozyme, chloroplast precursor (FNR) , partial (37%)
Length = 422
Score = 41.2 bits (95), Expect = 0.002
Identities = 31/98 (31%), Positives = 48/98 (48%), Gaps = 2/98 (2%)
Frame = +2
Query: 548 GVCSTWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADN-KVPIIMIGPGTGLAPFRGFLQE 606
G+CS ++ NS P +K Q + + LP D+ IMI GTG+APFRG+L+
Sbjct: 65 GICSNFLCNSKPGDKIQITGPSGKIML-----LPEDDPNATHIMIATGTGVAPFRGYLRR 229
Query: 607 RLALKEEGAELGP-SVLFFGCRNRQVDYIYEDELNHFV 643
+ G + LF G N +Y+DE + ++
Sbjct: 230 MFMESVPAYKFGGLAWLFLGVANTD-SLLYDDEFSKYL 340
>BE347814 similar to SP|P41346|FENR Ferredoxin--NADP reductase chloroplast
precursor (EC 1.18.1.2) (FNR). [Broad bean] {Vicia
faba}, partial (51%)
Length = 613
Score = 39.7 bits (91), Expect = 0.005
Identities = 39/136 (28%), Positives = 61/136 (44%), Gaps = 8/136 (5%)
Frame = +2
Query: 498 PPIGVFFASVAPRLQP---RYYSISSSPRVAPSRIHVTCALVHDKM----PTGRIHQGVC 550
P IGV + +P R YSI+SS + T +L ++ G + GVC
Sbjct: 17 PSIGVIPDGMDKNGEPH*LRLYSIASSA-IGDFGDSKTVSLCVKRLGFTNENGEMVYGVC 193
Query: 551 STWMKNSAPLEKSQDCSWAPIFVRQSNFRLPADNKVPIIMIGPGTGLAPFRGFLQERLAL 610
S ++ + P ++ P+ +P D IIM+G GTG+AP+R FL +
Sbjct: 194 SNFLCDLKPA--AEVTITVPV---GKEMLMPKDPNATIIMLGTGTGIAPYRSFLCKMFFA 358
Query: 611 KEEGAEL-GPSVLFFG 625
K + + G + LF G
Sbjct: 359 KHQDYKFNGEAWLFLG 406
>TC232174
Length = 796
Score = 36.2 bits (82), Expect = 0.054
Identities = 32/117 (27%), Positives = 52/117 (44%), Gaps = 4/117 (3%)
Frame = +1
Query: 624 FGCRNRQVDYIYEDELNHFVHGGALSELIVAFSREGPTKEYVQHKMIEKASDIWNMISQG 683
FGCR+ QVD ED N L E VAF ++ ++++ E SDI ++ S
Sbjct: 298 FGCRSEQVDATIEDLQN------KLKETTVAFELVTDERDLNKNRVSELESDIQSLQS-- 453
Query: 684 AYIYVCGDAKGMAKDVHRTLHTILQEQ----GSLDNSKTESMVKNLQMTGRYLRDVW 736
C + K +D H L L+E+ S+ N+ +N + +RD++
Sbjct: 454 ----ACSELKDKLEDYH-ALEEKLEEKEAEISSMHNAMLAKEEENSLLPASQMRDLF 609
>TC209358 similar to UP|P92987 (P92987) Myosin heavy chain-like protein,
partial (17%)
Length = 740
Score = 32.7 bits (73), Expect = 0.59
Identities = 14/42 (33%), Positives = 27/42 (63%)
Frame = +2
Query: 42 VMILTTSIAVLIGCVVVLIWRRSNSQKPKPIEVPKRVIEKLP 83
+MI+T+S+ + + C+V++IW R N +K E P ++ + P
Sbjct: 356 LMIMTSSLMISVTCLVLMIWTRWNCRKWMKQEKPIWLLFRFP 481
>TC205954 similar to UP|Q9FK50 (Q9FK50) Brn1-like protein, partial (89%)
Length = 958
Score = 32.0 bits (71), Expect = 1.0
Identities = 24/57 (42%), Positives = 27/57 (47%), Gaps = 2/57 (3%)
Frame = -1
Query: 415 GGSSLPPPFP--PCTLRTALAKYADVLSSPKKSALLALAAHASDPSEADRLRHLASP 469
G S LPPP P P T RT + +SS + SAL A S P R RH SP
Sbjct: 601 GSSPLPPPPPSTPRTPRTPASAPPPAMSSVE*SALRRRA*GGSRPPWRTRRRHARSP 431
>TC234932
Length = 440
Score = 30.8 bits (68), Expect = 2.3
Identities = 23/79 (29%), Positives = 42/79 (53%), Gaps = 3/79 (3%)
Frame = +3
Query: 248 LEKGGNRLVPVGLGDDDQCIEDDFTAWKEELWPAL-DQL-LRDEDDATVA-TPYTASVLE 304
+ GG+ +V G D+ C+E++F PAL D L L+DED+ A T S +
Sbjct: 105 INMGGDSIVSHGT-DEGSCLEENFPLLSSHSSPALRDNLGLQDEDNRMKAENGLTCSETD 281
Query: 305 YRVVIRDQLDATVDEKKQL 323
++ ++D V+E++++
Sbjct: 282 EEIIHPSRVDFEVNERRRV 338
>TC212647 weakly similar to UP|Q8L9A6 (Q8L9A6) Thioredoxin, partial (64%)
Length = 470
Score = 30.8 bits (68), Expect = 2.3
Identities = 16/56 (28%), Positives = 26/56 (45%)
Frame = +1
Query: 86 EIDDGTKKVTVFFGTQTGTAEGFAKAIAEEAKARYEKAKFKVVDMDDYAADDDEYE 141
EI D K V +FF F + E A+Y A + +D+++ + +EYE
Sbjct: 148 EIKDSPKPVVIFFTASWCGPCKFITPLFHEMAAKYPNADYVKIDVEELSGVAEEYE 315
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 30,268,778
Number of Sequences: 63676
Number of extensions: 429018
Number of successful extensions: 2323
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 2292
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2313
length of query: 736
length of database: 12,639,632
effective HSP length: 104
effective length of query: 632
effective length of database: 6,017,328
effective search space: 3802951296
effective search space used: 3802951296
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 63 (28.9 bits)
Medicago: description of AC135801.5


