

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135801.4 + phase: 0
(61 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
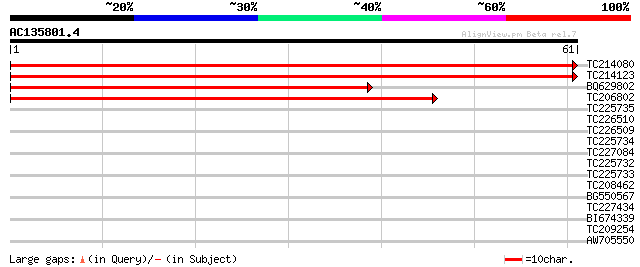
TC214080 homologue to UP|RUXF_ARATH (Q9SUM2) Probable small nucl... 122 4e-29
TC214123 homologue to UP|RUXF_ARATH (Q9SUM2) Probable small nucl... 120 1e-28
BQ629802 79 4e-16
TC206802 similar to PIR|T47805|T47805 U6 snRNA-associated Sm-lik... 50 3e-07
TC225735 homologue to UP|Q8LAK5 (Q8LAK5) Small nuclear ribonucle... 30 0.19
TC226510 similar to GB|BAC55689.1|27817924|AP003940 contains EST... 30 0.19
TC226509 similar to GB|BAC55689.1|27817924|AP003940 contains EST... 30 0.19
TC225734 homologue to UP|Q8LAK5 (Q8LAK5) Small nuclear ribonucle... 30 0.25
TC227084 homologue to UP|LSM4_TOBAC (Q43582) Probable U6 snRNA-a... 30 0.33
TC225732 homologue to UP|Q8LAK5 (Q8LAK5) Small nuclear ribonucle... 30 0.33
TC225733 homologue to UP|Q8LAK5 (Q8LAK5) Small nuclear ribonucle... 30 0.33
TC208462 homologue to UP|LSM4_TOBAC (Q43582) Probable U6 snRNA-a... 29 0.43
BG550567 similar to GP|21593343|gb|A small nuclear ribonucleopro... 28 1.2
TC227434 26 4.7
BI674339 homologue to SP|P80572|ADHX_ Alcohol dehydrogenase clas... 25 8.0
TC209254 homologue to UP|Q9FUN3 (Q9FUN3) QM-like protein, partia... 25 8.0
AW705550 GP|21666457|g polyprotein 1 {Bean pod mottle virus}, pa... 25 8.0
>TC214080 homologue to UP|RUXF_ARATH (Q9SUM2) Probable small nuclear
ribonucleoprotein F (snRNP-F) (Sm protein F) (Sm-F)
(SmF), partial (97%)
Length = 666
Score = 122 bits (306), Expect = 4e-29
Identities = 58/61 (95%), Positives = 60/61 (98%)
Frame = +2
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEEIEDAAD 60
MEYKGYLVSVDSYMNLQLANTEEYIEG FTGNLGEILIRCNNVLY+RGVPEDEEIEDAA+
Sbjct: 245 MEYKGYLVSVDSYMNLQLANTEEYIEGQFTGNLGEILIRCNNVLYLRGVPEDEEIEDAAE 424
Query: 61 D 61
D
Sbjct: 425 D 427
>TC214123 homologue to UP|RUXF_ARATH (Q9SUM2) Probable small nuclear
ribonucleoprotein F (snRNP-F) (Sm protein F) (Sm-F)
(SmF), partial (95%)
Length = 649
Score = 120 bits (302), Expect = 1e-28
Identities = 57/61 (93%), Positives = 59/61 (96%)
Frame = +3
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYMRGVPEDEEIEDAAD 60
MEYKGYLVSVDSYMNLQLANTEEYIEG FTGNLGEILIRCNNVLY+RGVPEDEEIED A+
Sbjct: 219 MEYKGYLVSVDSYMNLQLANTEEYIEGQFTGNLGEILIRCNNVLYLRGVPEDEEIEDVAE 398
Query: 61 D 61
D
Sbjct: 399 D 401
>BQ629802
Length = 439
Score = 79.3 bits (194), Expect = 4e-16
Identities = 38/39 (97%), Positives = 38/39 (97%)
Frame = +1
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIR 39
MEYKGYLVSVDSYMNLQLANTEEYIEG FTGNLGEILIR
Sbjct: 100 MEYKGYLVSVDSYMNLQLANTEEYIEGQFTGNLGEILIR 216
>TC206802 similar to PIR|T47805|T47805 U6 snRNA-associated Sm-like protein -
Arabidopsis thaliana {Arabidopsis thaliana;} , complete
Length = 705
Score = 49.7 bits (117), Expect = 3e-07
Identities = 20/46 (43%), Positives = 29/46 (62%)
Frame = +3
Query: 1 MEYKGYLVSVDSYMNLQLANTEEYIEGNFTGNLGEILIRCNNVLYM 46
++Y+G L +D YMN+ + TEEY+ G G+ IR NNVLY+
Sbjct: 291 VDYRGILACLDGYMNIAMEQTEEYVNGQLKNKYGDAFIRGNNVLYI 428
>TC225735 homologue to UP|Q8LAK5 (Q8LAK5) Small nuclear ribonucleoprotein
homolog, complete
Length = 431
Score = 30.4 bits (67), Expect = 0.19
Identities = 16/47 (34%), Positives = 27/47 (57%), Gaps = 1/47 (2%)
Frame = +1
Query: 1 MEYKGYLVSVDSYMNLQLANTEEY-IEGNFTGNLGEILIRCNNVLYM 46
+ +G ++ D YMNL L + EE I+ LG+IL++ +N+ M
Sbjct: 253 LRIEGRIIGFDEYMNLVLDDAEEVNIKKKSRKTLGKILLKGDNITLM 393
>TC226510 similar to GB|BAC55689.1|27817924|AP003940 contains ESTs
AU077521(C52420),AU077522(C52420) snRNP core Sm protein
Sm-X5-like protein {Oryza sativa (japonica
cultivar-group);} , complete
Length = 562
Score = 30.4 bits (67), Expect = 0.19
Identities = 20/55 (36%), Positives = 29/55 (52%), Gaps = 2/55 (3%)
Frame = +1
Query: 4 KGYLVSVDSYMNLQLANTEEYIEGNFTGNLG--EILIRCNNVLYMRGVPEDEEIE 56
KG L SVD Y+N++L NT + + L IR + V Y++ PE +IE
Sbjct: 160 KGTLHSVDQYLNIKLENTSVVNQEKYPHMLSVRNCFIRGSVVRYVQLPPEGVDIE 324
>TC226509 similar to GB|BAC55689.1|27817924|AP003940 contains ESTs
AU077521(C52420),AU077522(C52420) snRNP core Sm protein
Sm-X5-like protein {Oryza sativa (japonica
cultivar-group);} , complete
Length = 660
Score = 30.4 bits (67), Expect = 0.19
Identities = 20/55 (36%), Positives = 29/55 (52%), Gaps = 2/55 (3%)
Frame = +2
Query: 4 KGYLVSVDSYMNLQLANTEEYIEGNFTGNLG--EILIRCNNVLYMRGVPEDEEIE 56
KG L SVD Y+N++L NT + + L IR + V Y++ PE +IE
Sbjct: 182 KGTLHSVDQYLNIKLENTSVVNQEKYPHMLSVRNCFIRGSVVRYVQLPPEGVDIE 346
>TC225734 homologue to UP|Q8LAK5 (Q8LAK5) Small nuclear ribonucleoprotein
homolog, complete
Length = 837
Score = 30.0 bits (66), Expect = 0.25
Identities = 16/47 (34%), Positives = 26/47 (55%), Gaps = 1/47 (2%)
Frame = +2
Query: 1 MEYKGYLVSVDSYMNLQLANTEEY-IEGNFTGNLGEILIRCNNVLYM 46
+ +G ++ D YMNL L + EE I+ LG IL++ +N+ M
Sbjct: 215 LRIEGRIIGFDEYMNLVLDDAEEVNIKKKSRKTLGRILLKGDNITLM 355
>TC227084 homologue to UP|LSM4_TOBAC (Q43582) Probable U6 snRNA-associated
Sm-like protein LSm4 (Glycine-rich protein 10) (GRP 10),
partial (83%)
Length = 926
Score = 29.6 bits (65), Expect = 0.33
Identities = 23/68 (33%), Positives = 36/68 (52%), Gaps = 10/68 (14%)
Frame = +3
Query: 3 YKGYLVSVDSYMNLQLANTEEYI----EGNFTGNLGEILIRCNNVLYMRGVPED------ 52
Y G+LV+ D++MN+ L E I +G+ + E IR N + Y+R VP++
Sbjct: 222 YNGHLVNCDTWMNIHL---REVICTSKDGDRFWRMPECYIRGNTIKYLR-VPDEVIDKVQ 389
Query: 53 EEIEDAAD 60
EE + AD
Sbjct: 390 EETKSRAD 413
>TC225732 homologue to UP|Q8LAK5 (Q8LAK5) Small nuclear ribonucleoprotein
homolog, complete
Length = 715
Score = 29.6 bits (65), Expect = 0.33
Identities = 15/47 (31%), Positives = 26/47 (54%), Gaps = 1/47 (2%)
Frame = +1
Query: 1 MEYKGYLVSVDSYMNLQLANTEEY-IEGNFTGNLGEILIRCNNVLYM 46
+ +G ++ D YMNL L + EE ++ LG IL++ +N+ M
Sbjct: 208 LRIEGRIIGFDEYMNLVLDDAEEVNVKKKSRKTLGRILLKGDNITLM 348
>TC225733 homologue to UP|Q8LAK5 (Q8LAK5) Small nuclear ribonucleoprotein
homolog, complete
Length = 530
Score = 29.6 bits (65), Expect = 0.33
Identities = 15/47 (31%), Positives = 26/47 (54%), Gaps = 1/47 (2%)
Frame = +2
Query: 1 MEYKGYLVSVDSYMNLQLANTEEY-IEGNFTGNLGEILIRCNNVLYM 46
+ +G ++ D YMNL L + EE ++ LG IL++ +N+ M
Sbjct: 188 LRIEGRIIGFDEYMNLVLDDAEEVNVKKKSRKTLGRILLKGDNITLM 328
>TC208462 homologue to UP|LSM4_TOBAC (Q43582) Probable U6 snRNA-associated
Sm-like protein LSm4 (Glycine-rich protein 10) (GRP 10),
partial (94%)
Length = 794
Score = 29.3 bits (64), Expect = 0.43
Identities = 22/63 (34%), Positives = 35/63 (54%), Gaps = 4/63 (6%)
Frame = +3
Query: 3 YKGYLVSVDSYMNLQLANTEEYI----EGNFTGNLGEILIRCNNVLYMRGVPEDEEIEDA 58
Y G+LV+ D++MN+ L E I +G+ + E IR N + Y+R VP DE I+
Sbjct: 201 YNGHLVNCDTWMNIHL---REVICTSKDGDRFWRMPECYIRGNTIKYLR-VP-DEVIDKV 365
Query: 59 ADD 61
++
Sbjct: 366 QEE 374
>BG550567 similar to GP|21593343|gb|A small nuclear ribonucleoprotein homolog
{Arabidopsis thaliana}, partial (82%)
Length = 379
Score = 27.7 bits (60), Expect = 1.2
Identities = 15/44 (34%), Positives = 25/44 (56%), Gaps = 1/44 (2%)
Frame = +3
Query: 4 KGYLVSVDSYMNLQLANTEEY-IEGNFTGNLGEILIRCNNVLYM 46
+G ++ D YMNL L + E+ I+ LG IL++ +N+ M
Sbjct: 198 EGRIIGFDEYMNLVLDDKEKVNIKKKSRKTLGRILLKGDNITLM 329
>TC227434
Length = 1250
Score = 25.8 bits (55), Expect = 4.7
Identities = 10/22 (45%), Positives = 15/22 (67%)
Frame = +3
Query: 24 YIEGNFTGNLGEILIRCNNVLY 45
Y+EG F N+G+++ R N LY
Sbjct: 681 YMEGAFIVNIGDLMERWTNCLY 746
>BI674339 homologue to SP|P80572|ADHX_ Alcohol dehydrogenase class III (EC
1.1.1.1) (Glutathione-dependent formaldehyde
dehydrogenase), partial (23%)
Length = 379
Score = 25.0 bits (53), Expect = 8.0
Identities = 10/27 (37%), Positives = 17/27 (62%)
Frame = +3
Query: 4 KGYLVSVDSYMNLQLANTEEYIEGNFT 30
+GY+VS+ + ++LQ +EYI T
Sbjct: 252 QGYIVSLSALLDLQEIKVDEYITHTLT 332
>TC209254 homologue to UP|Q9FUN3 (Q9FUN3) QM-like protein, partial (96%)
Length = 1060
Score = 25.0 bits (53), Expect = 8.0
Identities = 8/17 (47%), Positives = 13/17 (76%)
Frame = -1
Query: 25 IEGNFTGNLGEILIRCN 41
+EG F+G LG +L+ C+
Sbjct: 406 VEGIFSGELGHVLVACD 356
>AW705550 GP|21666457|g polyprotein 1 {Bean pod mottle virus}, partial (7%)
Length = 413
Score = 25.0 bits (53), Expect = 8.0
Identities = 12/26 (46%), Positives = 17/26 (65%), Gaps = 1/26 (3%)
Frame = +3
Query: 16 LQLANTEEYIEGNF-TGNLGEILIRC 40
L LANT+E+ +GNF T + I + C
Sbjct: 24 LVLANTQEFFDGNFPTDSWLPIFVNC 101
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.137 0.385
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,915,223
Number of Sequences: 63676
Number of extensions: 15477
Number of successful extensions: 62
Number of sequences better than 10.0: 34
Number of HSP's better than 10.0 without gapping: 62
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 62
length of query: 61
length of database: 12,639,632
effective HSP length: 37
effective length of query: 24
effective length of database: 10,283,620
effective search space: 246806880
effective search space used: 246806880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC135801.4


