

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135799.3 + phase: 0
(242 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
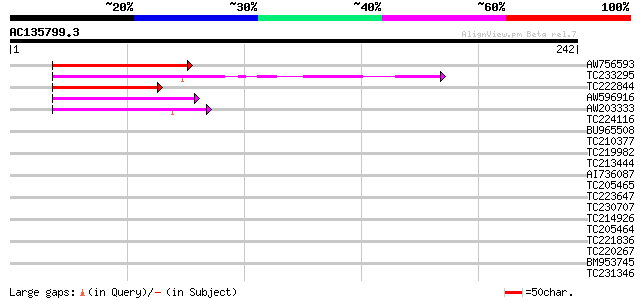
Score E
Sequences producing significant alignments: (bits) Value
AW756593 66 2e-11
TC233295 similar to UP|Q7KVQ0 (Q7KVQ0) CG4038-PA, partial (6%) 65 4e-11
TC222844 weakly similar to UP|Q9FFW4 (Q9FFW4) Similarity to heat... 52 2e-07
AW596916 52 2e-07
AW203333 48 3e-06
TC224116 40 0.001
BU965508 39 0.002
TC210377 similar to UP|Q8VAG3 (Q8VAG3) Wsv453 (WSSV513), partial... 38 0.005
TC219982 34 0.068
TC213444 32 0.26
AI736087 homologue to GP|21427473|gb| actin-related protein 8A {... 30 0.75
TC205465 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g5... 30 0.98
TC223647 30 0.98
TC230707 similar to UP|Q12937 (Q12937) BS-84, partial (4%) 30 0.98
TC214926 UP|E13A_SOYBN (Q03773) Glucan endo-1,3-beta-glucosidase... 30 0.98
TC205464 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g5... 30 0.98
TC221836 29 2.2
TC220267 similar to UP|Q6K235 (Q6K235) F-box-like protein, parti... 28 3.7
BM953745 28 4.9
TC231346 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2,... 28 4.9
>AW756593
Length = 370
Score = 65.9 bits (159), Expect = 2e-11
Identities = 29/60 (48%), Positives = 42/60 (69%)
Frame = +3
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
DKIS LPD++L +++ + T+EAV T +LSKRWNNLW L + FN K +S+ +KF+
Sbjct: 63 DKISELPDNILLHMMDFMDTREAVQTCVLSKRWNNLWKRLSTLLFNTSKFESVFKINKFL 242
>TC233295 similar to UP|Q7KVQ0 (Q7KVQ0) CG4038-PA, partial (6%)
Length = 557
Score = 64.7 bits (156), Expect = 4e-11
Identities = 55/171 (32%), Positives = 79/171 (46%), Gaps = 3/171 (1%)
Frame = +3
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIE---SNS 75
D+IS LPD +L ILSL+ TK+AV T ILSKRW LW ++ +DF++ +
Sbjct: 66 DRISHLPDVLLLQILSLLPTKQAVITGILSKRWRPLWPAVSVLDFDDESSPEFHHPGGLT 245
Query: 76 KFIDSVYSVLVSPDTAIGGSHFINRFCLDVQFRNGNPLYGYDNPHLLYKRSCHNVVKWVN 135
F + VYSVL+ D I RF + R NP Y S ++ W+
Sbjct: 246 GFAEFVYSVLLLHDAPA-----IERF----RLRCANPNY-----------SARDIATWLC 365
Query: 136 LVVQRRLKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLH 186
V +RR + + L+L L S LP +F C T+ + L+
Sbjct: 366 HVARRRAERVELSLSL-------------SRYVALPRCLFHCDTVSVMKLN 479
>TC222844 weakly similar to UP|Q9FFW4 (Q9FFW4) Similarity to heat shock
transcription factor HSF30, partial (6%)
Length = 746
Score = 52.4 bits (124), Expect = 2e-07
Identities = 23/47 (48%), Positives = 34/47 (71%)
Frame = +1
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNN 65
D+IS LPD +L ++++ + TK AV T +LSKRWN+L L N+ FN+
Sbjct: 496 DRISALPDSLLFHMINFMDTKSAVQTCVLSKRWNDLSKCLTNLTFNS 636
>AW596916
Length = 455
Score = 52.4 bits (124), Expect = 2e-07
Identities = 26/63 (41%), Positives = 37/63 (58%)
Frame = +3
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKIDSIESNSKFI 78
D++S LPD VL +I+ +S K AV T +LSKRW LW L N+ ++ + SKF+
Sbjct: 264 DRLSDLPDFVLLHIMKFMSMKHAVQTCVLSKRWKELWKRLTNLALHSSDFADLAHFSKFL 443
Query: 79 DSV 81
V
Sbjct: 444 SWV 452
>AW203333
Length = 418
Score = 48.1 bits (113), Expect = 3e-06
Identities = 28/77 (36%), Positives = 39/77 (50%), Gaps = 9/77 (11%)
Frame = +1
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNNIKI---------D 69
D IS LP +LH ILS + ++AV TS+LSK W W + P + F++ I D
Sbjct: 85 DMISTLPKTILHDILSRMPEEDAVRTSVLSKSWAEAWSTYPILYFSDTMIVGTFPRPWED 264
Query: 70 SIESNSKFIDSVYSVLV 86
+ FID V L+
Sbjct: 265 FLRKRKNFIDHVKRTLL 315
>TC224116
Length = 520
Score = 39.7 bits (91), Expect = 0.001
Identities = 20/59 (33%), Positives = 31/59 (51%)
Frame = +2
Query: 7 RRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNN 65
RR + D IS LP ++ IL + ++AV TSILS +W W S+ + F++
Sbjct: 251 RRSIRMGDVMGPDLISDLPQSIIESILVQLPIRDAVRTSILSSKWRYKWASITQLVFDD 427
>BU965508
Length = 447
Score = 39.3 bits (90), Expect = 0.002
Identities = 18/47 (38%), Positives = 28/47 (59%)
Frame = +2
Query: 19 DKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSLPNIDFNN 65
D IS LP ++ IL + ++AV TSILS +W W S+ + F++
Sbjct: 296 DLISDLPQSIIESILVQLPIRDAVRTSILSSKWRYKWASITRLVFDD 436
>TC210377 similar to UP|Q8VAG3 (Q8VAG3) Wsv453 (WSSV513), partial (10%)
Length = 1077
Score = 37.7 bits (86), Expect = 0.005
Identities = 25/82 (30%), Positives = 45/82 (54%), Gaps = 1/82 (1%)
Frame = +3
Query: 1 MQRQKRRRRSSKASLAVADKISRLPDDVLHYILSLVSTKEAVATSIL-SKRWNNLWLSLP 59
M+ +K+R+R+ K LP+ VL I+ ++TK +V T +L SKRW +L L
Sbjct: 240 MKTKKKRQRTEKRR-----GTGDLPNCVLLQIMEFMNTKYSVQTCVLLSKRWKDL*KYLT 404
Query: 60 NIDFNNIKIDSIESNSKFIDSV 81
++ F+ + D++ FI ++
Sbjct: 405 SLSFSRFQRDNLCGRGYFITNL 470
>TC219982
Length = 590
Score = 33.9 bits (76), Expect = 0.068
Identities = 24/81 (29%), Positives = 35/81 (42%)
Frame = +1
Query: 142 LKYLRLNLRLGYDLHLDVDDYDNSYLPKLPITIFTCRTLVSLDLHHFSVKGVCYRFLLKA 201
LK R R G + L + Y S LP L + + + + +HH S +CY F +
Sbjct: 49 LKVSRNMWRNGIHIQLGLKSYCCSVLPILDTSYWDLMDIYDIYIHHHSCYYLCYYFRARN 228
Query: 202 LSTSNSLRLDTFKLYRSIYQV 222
TSN + RS +QV
Sbjct: 229 QRTSNGRSGEDVGRKRSQFQV 291
>TC213444
Length = 530
Score = 32.0 bits (71), Expect = 0.26
Identities = 11/24 (45%), Positives = 20/24 (82%)
Frame = +2
Query: 19 DKISRLPDDVLHYILSLVSTKEAV 42
D++S +PD ++H+ILS + TK+A+
Sbjct: 458 DRLSDMPDCIIHHILSFMETKDAI 529
>AI736087 homologue to GP|21427473|gb| actin-related protein 8A {Arabidopsis
thaliana}, partial (13%)
Length = 427
Score = 30.4 bits (67), Expect = 0.75
Identities = 19/55 (34%), Positives = 29/55 (52%), Gaps = 8/55 (14%)
Frame = +1
Query: 10 SSKASLAVAD--KISRLPDDVLHYILSLVSTKEAVATSILSKRW------NNLWL 56
SS +A +D ++ LP D+L +IL L+ KEA S++ K N LW+
Sbjct: 55 SSPPEVACSDMGELELLPSDILMHILRLLGPKEAAKLSVVCKALRPLVSDNRLWI 219
>TC205465 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g57790
{Arabidopsis thaliana;} , partial (15%)
Length = 490
Score = 30.0 bits (66), Expect = 0.98
Identities = 14/37 (37%), Positives = 24/37 (64%)
Frame = +3
Query: 22 SRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSL 58
S LP ++L ILS +S + V S++ KRW+++ S+
Sbjct: 180 SDLPTELLELILSRLSLDDNVRASVVCKRWHSVATSV 290
>TC223647
Length = 448
Score = 30.0 bits (66), Expect = 0.98
Identities = 18/51 (35%), Positives = 28/51 (54%), Gaps = 3/51 (5%)
Frame = +3
Query: 7 RRRSS---KASLAVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNL 54
RR+SS KAS +S L +D+L IL+ + + + +SK WN+L
Sbjct: 63 RRKSSTKVKASTDANTNLSMLNEDLLQNILARLPALHFASAACVSKSWNSL 215
>TC230707 similar to UP|Q12937 (Q12937) BS-84, partial (4%)
Length = 990
Score = 30.0 bits (66), Expect = 0.98
Identities = 21/56 (37%), Positives = 31/56 (54%)
Frame = +3
Query: 164 NSYLPKLPITIFTCRTLVSLDLHHFSVKGVCYRFLLKALSTSNSLRLDTFKLYRSI 219
NS+ +P +I T ++L+SLDL H S+ G + +L+T L L KL SI
Sbjct: 117 NSFSGTIPPSITTLKSLLSLDLAHNSLSGYLPN-SMNSLTTLRRLDLSFNKLTGSI 281
>TC214926 UP|E13A_SOYBN (Q03773) Glucan endo-1,3-beta-glucosidase precursor
((1->3)-beta-glucan endohydrolase)
((1->3)-beta-glucanase) (Beta-1,3-endoglucanase) ,
partial (8%)
Length = 425
Score = 30.0 bits (66), Expect = 0.98
Identities = 20/52 (38%), Positives = 26/52 (49%), Gaps = 5/52 (9%)
Frame = +2
Query: 194 CYRFLLKALSTSNSLRLDTFKLYRSIYQVRFIMNTMHVI-----YFLFLSEI 240
C + ST SLR TF L+ +IY +IMN MHV Y +F S +
Sbjct: 107 CVSLCINMFSTLPSLRALTF-LHVTIYLYEYIMNRMHVYSYESRYIVFWSRL 259
>TC205464 weakly similar to GB|AAP42752.1|30984578|BT008739 At1g57790
{Arabidopsis thaliana;} , partial (17%)
Length = 756
Score = 30.0 bits (66), Expect = 0.98
Identities = 14/37 (37%), Positives = 24/37 (64%)
Frame = +1
Query: 22 SRLPDDVLHYILSLVSTKEAVATSILSKRWNNLWLSL 58
S LP ++L ILS +S + V S++ KRW+++ S+
Sbjct: 184 SDLPTELLELILSRLSLDDNVRASVVCKRWHSVATSV 294
>TC221836
Length = 641
Score = 28.9 bits (63), Expect = 2.2
Identities = 16/48 (33%), Positives = 23/48 (47%), Gaps = 8/48 (16%)
Frame = -2
Query: 35 LVSTKEAVATSILSKRWNNLWLS--------LPNIDFNNIKIDSIESN 74
+++ V +I S RW N WLS L +ID NI ++ SN
Sbjct: 271 IITDLHKVHVAIFSWRWRNFWLSFLVR*LCFLSSIDIFNILLEMCHSN 128
>TC220267 similar to UP|Q6K235 (Q6K235) F-box-like protein, partial (7%)
Length = 586
Score = 28.1 bits (61), Expect = 3.7
Identities = 13/39 (33%), Positives = 20/39 (50%)
Frame = +2
Query: 16 AVADKISRLPDDVLHYILSLVSTKEAVATSILSKRWNNL 54
A A + LPDD+L IL+ + +SKRW+ +
Sbjct: 314 ATAVSLDVLPDDLLERILAYLPIASIFRAGCVSKRWHEI 430
>BM953745
Length = 420
Score = 27.7 bits (60), Expect = 4.9
Identities = 11/27 (40%), Positives = 17/27 (62%)
Frame = -2
Query: 31 YILSLVSTKEAVATSILSKRWNNLWLS 57
Y S S+ ++A+ + RWNNLWL+
Sbjct: 215 YYFSFHSSILSLASKVCFTRWNNLWLA 135
>TC231346 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2, partial
(14%)
Length = 1790
Score = 27.7 bits (60), Expect = 4.9
Identities = 18/56 (32%), Positives = 26/56 (46%)
Frame = +1
Query: 164 NSYLPKLPITIFTCRTLVSLDLHHFSVKGVCYRFLLKALSTSNSLRLDTFKLYRSI 219
NS +P+T+ T LV+LDL ++G L T LRL L+ S+
Sbjct: 301 NSLTGDVPVTLGTLSNLVTLDLSSNLLEGSIKESNFVKLFTLKELRLSWTNLFLSV 468
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,287,290
Number of Sequences: 63676
Number of extensions: 184751
Number of successful extensions: 1384
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 1378
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1384
length of query: 242
length of database: 12,639,632
effective HSP length: 95
effective length of query: 147
effective length of database: 6,590,412
effective search space: 968790564
effective search space used: 968790564
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135799.3


