

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
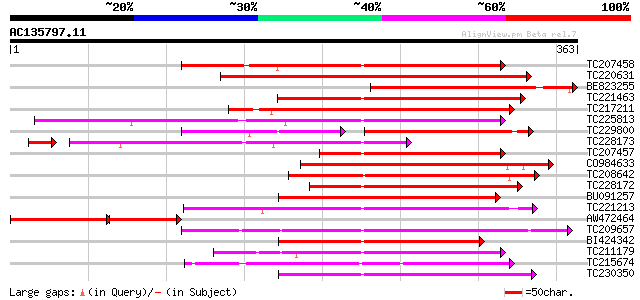
Query= AC135797.11 + phase: 0 /pseudo
(363 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC207458 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, par... 232 2e-61
TC220631 weakly similar to UP|O80638 (O80638) Nodulin-like prote... 188 4e-48
BE823255 similar to PIR|T00561|T005 nodulin-like protein [import... 187 6e-48
TC221463 similar to UP|Q8LRB6 (Q8LRB6) Nodulin-like protein, par... 186 2e-47
TC217211 weakly similar to UP|Q8LA54 (Q8LA54) Nodulin-like prote... 179 2e-45
TC225813 174 7e-44
TC229800 similar to UP|O24091 (O24091) MtN21 protein, partial (53%) 129 9e-44
TC228173 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, par... 164 7e-43
TC207457 weakly similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like prote... 170 8e-43
CO984633 167 5e-42
TC208642 weakly similar to UP|Q8LA54 (Q8LA54) Nodulin-like prote... 165 3e-41
TC228172 weakly similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like prote... 162 3e-40
BU091257 weakly similar to GP|21593603|gb nodulin-like protein {... 157 5e-39
TC221213 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein... 148 3e-36
AW472464 similar to PIR|A84793|A84 nodulin-like protein [importe... 94 2e-35
TC209657 weakly similar to UP|Q94JU2 (Q94JU2) AT3g28050/MMG15_6,... 138 3e-33
BI424342 136 1e-32
TC211179 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein... 132 2e-31
TC215674 weakly similar to UP|Q9SUD5 (Q9SUD5) Medicago nodulin N... 125 4e-29
TC230350 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein... 123 1e-28
>TC207458 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial (47%)
Length = 1006
Score = 232 bits (591), Expect = 2e-61
Identities = 117/211 (55%), Positives = 149/211 (70%), Gaps = 4/211 (1%)
Frame = +2
Query: 111 QMEKIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHA 170
++E + +R +HS AK+VGT TV+GAMVMTL KGP L + A H G + Q +
Sbjct: 29 RLETVNLRKIHSVAKVVGTAVTVSGAMVMTLYKGPALQFI---KGQAATHHESGNSTQPS 199
Query: 171 ----VKGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERG 226
V G++ + C A F ILQ+ TL+ YPAELS+TAWIC LG EG I LI ER
Sbjct: 200 EQNWVLGTVELIASCGGWASFFILQSFTLKMYPAELSVTAWICFLGIFEGAIATLIFER- 376
Query: 227 EPSVWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPF 286
+ SVWS+ D++LLA VYSG+VCSGMAYY+QGVV R RGPVFVT+F+PLCM+I A +
Sbjct: 377 DMSVWSIGMDSRLLACVYSGVVCSGMAYYVQGVVTRERGPVFVTSFSPLCMIITAALGSI 556
Query: 287 ILAEKIYLGRVIGAVVIILGLYLVVWGKSKD 317
+LAE++YLG VIGA++I+ GLY VVWGKSKD
Sbjct: 557 VLAEQVYLGSVIGAIIIVSGLYTVVWGKSKD 649
>TC220631 weakly similar to UP|O80638 (O80638) Nodulin-like protein
(At2g39510), partial (31%)
Length = 797
Score = 188 bits (477), Expect = 4e-48
Identities = 88/200 (44%), Positives = 133/200 (66%), Gaps = 1/200 (0%)
Frame = +1
Query: 136 AMVMTLIKGPILNLFGIHESSA-QIQHNGGVNLQHAVKGSIMITIGCFSCACFTILQAVT 194
A +M L KGP++N+ G S Q ++ + H + G+ + IGC + F ILQA+T
Sbjct: 1 AFLMALYKGPVVNIAGSSASHVGQPENVNDPSGSHWLIGACFLLIGCAGFSAFYILQAIT 180
Query: 195 LETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTKLLAAVYSGIVCSGMAY 254
L YPAE+SL W+C +G ++ V+ MER P VWSL+WD++L+A YSGIV S + +
Sbjct: 181 LRKYPAEMSLATWVCFVGALQSSAVSFFMERNSPDVWSLAWDSRLVAYAYSGIVTSAIQF 360
Query: 255 YIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVIGAVVIILGLYLVVWGK 314
Y+QG+V++ GPVFVT FNPL M+IV ++ +L+EK++LG +IG VV+++GLYLVVWGK
Sbjct: 361 YVQGMVIKTTGPVFVTAFNPLRMIIVTALACIVLSEKLHLGSIIGGVVVVIGLYLVVWGK 540
Query: 315 SKDYDRPSPIIKDEILPAKQ 334
+K+ P +++ +Q
Sbjct: 541 AKEQKHLMPPSPEKVTLQRQ 600
>BE823255 similar to PIR|T00561|T005 nodulin-like protein [imported] -
Arabidopsis thaliana, partial (22%)
Length = 537
Score = 187 bits (475), Expect = 6e-48
Identities = 97/134 (72%), Positives = 108/134 (80%), Gaps = 2/134 (1%)
Frame = -2
Query: 232 SLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEK 291
SL WDTKLLAAVYSGIVCSGMAYYIQG VM+ RGPVFVTTFNPLCMVIVAIM F LAE
Sbjct: 536 SLQWDTKLLAAVYSGIVCSGMAYYIQGAVMKDRGPVFVTTFNPLCMVIVAIMGSFFLAEI 357
Query: 292 IYLGRVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEILPAKQTIENNDKEKFHSHEVIT 351
+YLGRV+GA+VIILGLYLVVWGKS DY+ + I K LP+KQT+E E+ +H+VIT
Sbjct: 356 MYLGRVVGAIVIILGLYLVVWGKSNDYESSNSITKKHTLPSKQTVE----EEHSNHDVIT 189
Query: 352 SSNFGA--IARDEQ 363
SN GA I RDEQ
Sbjct: 188 LSNLGAGNIVRDEQ 147
>TC221463 similar to UP|Q8LRB6 (Q8LRB6) Nodulin-like protein, partial (37%)
Length = 827
Score = 186 bits (471), Expect = 2e-47
Identities = 90/159 (56%), Positives = 118/159 (73%)
Frame = +2
Query: 172 KGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVW 231
KGS+++ + S A F ILQAVTL YPA+LSLTA +C LGT++ V +ME +PSVW
Sbjct: 59 KGSVLLVLATLSWASFFILQAVTLRKYPAQLSLTALVCALGTLQSIAVTFVMEH-QPSVW 235
Query: 232 SLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEK 291
++ WD LLAA Y+GI+ SG+ YY+QG+VM+ +GPVFVT F+PL M+IVAIM FILAEK
Sbjct: 236 TIGWDMNLLAAAYAGIISSGITYYVQGIVMQKKGPVFVTAFSPLMMIIVAIMGTFILAEK 415
Query: 292 IYLGRVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEIL 330
IYLG VIGA++I++GLY V+WGK K+ I E+L
Sbjct: 416 IYLGGVIGAILIVMGLYSVLWGKHKENKEKEAEITIEVL 532
>TC217211 weakly similar to UP|Q8LA54 (Q8LA54) Nodulin-like protein, partial
(40%)
Length = 909
Score = 179 bits (454), Expect = 2e-45
Identities = 85/188 (45%), Positives = 129/188 (68%), Gaps = 5/188 (2%)
Frame = +2
Query: 141 LIKGPILNLFGIHESSAQIQHNGGVN----LQHAVKGSIMITIGCFSCACFTILQAVTLE 196
L KGP + LF +++ H G + ++H + G++ + +GC + + F ILQ++TL+
Sbjct: 2 LYKGPQIKLFFSPDTT---HHQDGSHSPQVIKHWLSGTLFLLLGCVAWSSFFILQSITLK 172
Query: 197 TYPAELSLTAWICLLGTVEGGIVALIMERGEPSV-WSLSWDTKLLAAVYSGIVCSGMAYY 255
YPAELSL++ +CL G ++ +VA++ R V W+L WD +L +Y+GIV SG+ YY
Sbjct: 173 RYPAELSLSSLVCLSGALQASVVAIVATRHSGLVAWALGWDFRLYGPLYTGIVTSGITYY 352
Query: 256 IQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVIGAVVIILGLYLVVWGKS 315
QG++++ RGPVF+T FNPLCMVI + + F+ AE+++LG +IGAV+I LGLY VVWGK
Sbjct: 353 AQGLILQTRGPVFLTAFNPLCMVITSALGSFLFAEQLHLGSIIGAVIIALGLYSVVWGKG 532
Query: 316 KDYDRPSP 323
KDY P+P
Sbjct: 533 KDYSNPTP 556
>TC225813
Length = 1551
Score = 174 bits (440), Expect = 7e-44
Identities = 108/329 (32%), Positives = 174/329 (52%), Gaps = 28/329 (8%)
Frame = +1
Query: 17 AVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPFAVILENGSQCARACY* 76
A++ LQFGYAG ++S++ALN G+S V VYR+ +AF++++PFA LE + A
Sbjct: 157 AMLALQFGYAGFHVVSRAALNMGISKLVFPVYRNIIAFLLLLPFAYFLEKKERPAITLNF 336
Query: 77 ---------------SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKMRSVH 121
F G + + F S + ++E++++
Sbjct: 337 LLQFFLLALVGITANQGFYLLGLDNTSPTFASAIQNSVPAITFLMAVILRIEQVRLNRKD 516
Query: 122 SQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAVKGSI------ 175
AK+ GTI VAGA V+TL KGP I+ S +Q V ++ S+
Sbjct: 517 GIAKVAGTIFCVAGATVITLYKGPT-----IYSPSPPLQSESSVVVEFGTLSSLGDAKGK 681
Query: 176 ------MITIG-CFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEP 228
+ IG C S + + +LQA L+ YPA LS+T++ C G ++ ++ALI+ER +
Sbjct: 682 NWTLGCLYLIGHCLSWSAWLVLQAPVLKKYPARLSVTSYTCFFGLIQFLVIALIVER-DA 858
Query: 229 SVWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFIL 288
W ++ +Y+G+V SG+A+ +Q + GPVFV + P+ ++VAIM+ L
Sbjct: 859 QAWIFQSGGEVFTILYAGVVASGIAFAVQIWCIDRGGPVFVAVYQPVQTLVVAIMASLAL 1038
Query: 289 AEKIYLGRVIGAVVIILGLYLVVWGKSKD 317
E+ YLG +IGAV+I++GLY V+WGKS++
Sbjct: 1039GEEFYLGGIIGAVLIVVGLYFVLWGKSEE 1125
>TC229800 similar to UP|O24091 (O24091) MtN21 protein, partial (53%)
Length = 1303
Score = 129 bits (323), Expect(2) = 9e-44
Identities = 62/109 (56%), Positives = 82/109 (74%), Gaps = 1/109 (0%)
Frame = +2
Query: 228 PSVWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFI 287
PSVW + WD LLAA Y+GIV S ++YY+QG+V++ +GPVF T F+PL M+IVAIM FI
Sbjct: 416 PSVWRIGWDVSLLAAAYAGIVTSSISYYVQGLVIKMKGPVFATAFSPLMMIIVAIMGSFI 595
Query: 288 LAEKIYLGRVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEI-LPAKQT 335
LAE+IYLG VIGA++I++GLY V+WGK K+ + DEI LP K +
Sbjct: 596 LAEQIYLGGVIGAILIVIGLYSVLWGKHKEQIESK--VADEIPLPVKDS 736
Score = 66.2 bits (160), Expect(2) = 9e-44
Identities = 40/109 (36%), Positives = 63/109 (57%), Gaps = 4/109 (3%)
Frame = +1
Query: 111 QMEKIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGI---HESSAQIQHNGGVNL 167
+MEKI ++ V AKIVGT+ TVAGAM+MTL +GPI+ + H + G ++
Sbjct: 58 RMEKIDIKKVRCIAKIVGTLVTVAGAMLMTLYRGPIVEMVWAKHPHNKTNATTTTGSLDK 237
Query: 168 QHAVKGSIMITIGCFSCACFTILQAVTLETYP-AELSLTAWICLLGTVE 215
+ G + I + A +LQA ++TY +LSLT+ +C +GT++
Sbjct: 238 DWFL-GCTFLIIATLAWASLFVLQAKAIQTYKNHQLSLTSLVCFIGTLQ 381
>TC228173 similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial (54%)
Length = 904
Score = 164 bits (414), Expect(2) = 7e-43
Identities = 94/241 (39%), Positives = 134/241 (55%), Gaps = 22/241 (9%)
Frame = +2
Query: 39 GMSCYVLVVYRHAVAFVVIVPFAVILENGSQ---------------CARACY*SKFIFFG 83
GM ++L RHA A ++IVP A++LE + G
Sbjct: 191 GMGFWILSRXRHAAAAIIIVPLALVLERKIRPLMTLPIFLRIVALGFLEPVLDQNLYHMG 370
Query: 84 NEVYNSYFCSCHVQCPSCYYLCRGLDSQMEKIKMRSVHSQAKIVGTIATVAGAMVMTLIK 143
++ ++ F S V L ++EK+ +R HS AK++GT+ TV+GAMVMTL K
Sbjct: 371 MKMTSTTFASATVNVLPAITFVMALIFRLEKVNLRKFHSVAKVIGTLITVSGAMVMTLYK 550
Query: 144 GPILNLFGIHESSAQIQHNGGVNL-------QHAVKGSIMITIGCFSCACFTILQAVTLE 196
GP + I A H+ + QH + G++ + C S A F ILQ+ TL+
Sbjct: 551 GPAFQI--IKGGGAISNHSNSSSTSTTEPSDQHWIVGTVYLISSCASWAGFFILQSFTLK 724
Query: 197 TYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTKLLAAVYSGIVCSGMAYYI 256
YPAELSLTAWIC++G +EG I +LI ER + SVW++ WD++LLA VYSG++CSGMAYY+
Sbjct: 725 KYPAELSLTAWICVMGIIEGSIASLIFER-DFSVWAIGWDSRLLACVYSGVICSGMAYYV 901
Query: 257 Q 257
Q
Sbjct: 902 Q 904
Score = 28.1 bits (61), Expect(2) = 7e-43
Identities = 10/18 (55%), Positives = 15/18 (82%)
Frame = +3
Query: 13 KPFIAVVLLQFGYAGMDI 30
KP++A++ L GY+GMDI
Sbjct: 114 KPYLAILXLXXGYSGMDI 167
>TC207457 weakly similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial
(32%)
Length = 757
Score = 170 bits (431), Expect = 8e-43
Identities = 79/119 (66%), Positives = 97/119 (81%)
Frame = +1
Query: 199 PAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTKLLAAVYSGIVCSGMAYYIQG 258
PAELS+TAWIC LG EG I LI ER + SVWS+ D++LLA VYSG+VCSGMAYY+QG
Sbjct: 1 PAELSVTAWICFLGIFEGAIATLIFER-DMSVWSIGMDSRLLACVYSGVVCSGMAYYVQG 177
Query: 259 VVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVIGAVVIILGLYLVVWGKSKD 317
VV R RGPVFVT+F+PLCM+I A + +LAE++Y+G VIGA++I+ GLY VVWGKSKD
Sbjct: 178 VVTRERGPVFVTSFSPLCMIITAALGSIVLAEQVYMGSVIGAIIIVSGLYTVVWGKSKD 354
>CO984633
Length = 705
Score = 167 bits (424), Expect = 5e-42
Identities = 84/175 (48%), Positives = 113/175 (64%), Gaps = 13/175 (7%)
Frame = -3
Query: 187 FTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTKLLAAVYSG 246
F ILQA+TL YPAE+SL W+C +G ++ IVA+ ER P WSL WDT+L A Y+G
Sbjct: 700 FYILQAITLRKYPAEMSLATWVCFVGALQSSIVAIFAERHHPHAWSLGWDTRLFAPAYAG 521
Query: 247 IVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVIGAVVIILG 306
IV SG+ YYIQG+V + GPV VT FNPL M+IV ++ IL+E+++LG +IGA+V++LG
Sbjct: 520 IVTSGVQYYIQGMVSKIMGPVIVTAFNPLRMIIVTALACIILSEQLFLGSIIGAIVVVLG 341
Query: 307 LYLVVWGKSKD---YDRPSPIIKD---------EILPAKQTIENNDKE-KFHSHE 348
LYLVVWGK+K+ PSP + P +I NN+K H H+
Sbjct: 340 LYLVVWGKAKERRGLMTPSPAENNFPEDQRQLPVTAPRNDSINNNNKA*LVHRHD 176
>TC208642 weakly similar to UP|Q8LA54 (Q8LA54) Nodulin-like protein, partial
(37%)
Length = 779
Score = 165 bits (417), Expect = 3e-41
Identities = 78/165 (47%), Positives = 113/165 (68%), Gaps = 4/165 (2%)
Frame = +2
Query: 179 IGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTK 238
+GC + + F ILQ++T++ YPAELSL++ IC G ++ +VALI + P W++ +D
Sbjct: 2 LGCLAWSSFYILQSITVKRYPAELSLSSLICFAGALQSAVVALIADHN-PRAWAIGFDYS 178
Query: 239 LLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVI 298
L +Y+GI+ SG+AYYIQG+VM+ RGPVFVT+FNPLCM+IV + F+L E +YLG +I
Sbjct: 179 LYGPLYTGIMSSGIAYYIQGLVMQSRGPVFVTSFNPLCMIIVTALGSFLLGEHLYLGSII 358
Query: 299 GAVVIILGLYLVVWGKSKDY----DRPSPIIKDEILPAKQTIENN 339
G ++I +GLY VVWGK KDY P+ + E + T+ NN
Sbjct: 359 GGIIIAVGLYSVVWGKGKDYKDDTSSPATTKETETMQLPITLPNN 493
>TC228172 weakly similar to UP|Q9SUF0 (Q9SUF0) Nodulin-like protein, partial
(36%)
Length = 764
Score = 162 bits (409), Expect = 3e-40
Identities = 75/136 (55%), Positives = 102/136 (74%)
Frame = +2
Query: 193 VTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTKLLAAVYSGIVCSGM 252
+ L+ YPAELSLTAW ++G + I +LI + SVW++ WD++LLA VYSG++CSGM
Sbjct: 14 IHLKKYPAELSLTAWXXVMGXMRVSIASLIFXX-DFSVWAIGWDSRLLACVYSGVICSGM 190
Query: 253 AYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVIGAVVIILGLYLVVW 312
AYY+QGVV R RGPVFVT+F+PLCM+I A + +LAE+++LG + GA++I+ GLY VVW
Sbjct: 191 AYYVQGVVTRERGPVFVTSFSPLCMIITAALGSLVLAEQVHLGSIFGAILIVCGLYTVVW 370
Query: 313 GKSKDYDRPSPIIKDE 328
GKSKD + I K E
Sbjct: 371 GKSKDRKSTTEIEKGE 418
>BU091257 weakly similar to GP|21593603|gb nodulin-like protein {Arabidopsis
thaliana}, partial (35%)
Length = 429
Score = 157 bits (398), Expect = 5e-39
Identities = 68/143 (47%), Positives = 105/143 (72%), Gaps = 1/143 (0%)
Frame = +1
Query: 173 GSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIM-ERGEPSVW 231
G++ + +GC + + F ILQ++TL+ YPAELSL++ +CL G ++ G+V L+ + W
Sbjct: 1 GTLFLLLGCVAWSSFIILQSITLKRYPAELSLSSLVCLSGALQAGVVTLVATHQSGLGPW 180
Query: 232 SLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEK 291
+L WD +L +Y+G+V SG+ YY+QG+V++ +GPVF T FNPLCM+I + + FI AE+
Sbjct: 181 ALGWDFRLYGPLYTGVVTSGITYYVQGLVLQSKGPVFFTAFNPLCMIITSALGSFIFAEQ 360
Query: 292 IYLGRVIGAVVIILGLYLVVWGK 314
++LG +IGA++I LGLY VVWGK
Sbjct: 361 LHLGSIIGAIIIALGLYSVVWGK 429
>TC221213 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein;
91922-89607, partial (37%)
Length = 941
Score = 148 bits (374), Expect = 3e-36
Identities = 87/230 (37%), Positives = 133/230 (57%), Gaps = 3/230 (1%)
Frame = +1
Query: 112 MEKIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQ---HNGGVNLQ 168
ME++ +R+ +AKIVGT+ + GAMV+T +KG + H + Q H
Sbjct: 139 MERLNLRTAAGKAKIVGTLIGIGGAMVLTFVKGVHIEFGSFHLNLLHPQNGTHAHSATGA 318
Query: 169 HAVKGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEP 228
H + GS+ S A + I+QA E+YP S TA + L G++ ++AL +ER +
Sbjct: 319 HTLLGSLCALASGISYALWLIIQAKMSESYPRPYSSTALMSLWGSLLSIVLALCVER-DW 495
Query: 229 SVWSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFIL 288
S W L W+ KLL A Y+GIV SG + + RGP+F + F+PL +V A+ IL
Sbjct: 496 SQWRLGWNIKLLTAAYTGIVVSGEMVDVISWCVHMRGPLFASVFSPLMLVTEALAGSTIL 675
Query: 289 AEKIYLGRVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEILPAKQTIEN 338
EK++LG VIGAV+I+ GLY+V+WGKSK+ + K++++PA+ +N
Sbjct: 676 NEKLHLGCVIGAVLIVCGLYVVLWGKSKEMKK-----KNQLVPAQSPHDN 810
>AW472464 similar to PIR|A84793|A84 nodulin-like protein [imported] -
Arabidopsis thaliana, partial (32%)
Length = 444
Score = 93.6 bits (231), Expect(2) = 2e-35
Identities = 48/65 (73%), Positives = 51/65 (77%)
Frame = +1
Query: 1 MAFCNQKSFQNWKPFIAVVLLQFGYAGMDILSKSALNKGMSCYVLVVYRHAVAFVVIVPF 60
MA QK F KPFI VV LQFGYAGMDILSK+ALNKGMS YV VVYRH AFVV+ PF
Sbjct: 34 MAGDKQKLFNRLKPFIGVVFLQFGYAGMDILSKAALNKGMSNYVFVVYRHVFAFVVMAPF 213
Query: 61 AVILE 65
A+ILE
Sbjct: 214ALILE 228
Score = 73.9 bits (180), Expect(2) = 2e-35
Identities = 31/46 (67%), Positives = 37/46 (80%)
Frame = +3
Query: 65 ENGSQCARACY*SKFIFFGNEVYNSYFCSCHVQCPSCYYLCRGLDS 110
+N SQ ARACY* KF+FFG+EV++S C HVQCP C+YLC GLDS
Sbjct: 270 DNDSQLARACY*PKFVFFGHEVHHSNLCCFHVQCPPCHYLCHGLDS 407
>TC209657 weakly similar to UP|Q94JU2 (Q94JU2) AT3g28050/MMG15_6, partial
(31%)
Length = 1014
Score = 138 bits (348), Expect = 3e-33
Identities = 81/250 (32%), Positives = 133/250 (52%)
Frame = +1
Query: 111 QMEKIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHA 170
+MEK+ R + S AK++GTI ++AGA ++TL KGP L L G+ ++ Q +
Sbjct: 88 RMEKLDWRKLSSLAKLLGTIVSIAGAFIVTLYKGPAL-LMGVSSANTSQQPLLSEDSNWI 264
Query: 171 VKGSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSV 230
+ G + + C + + I+QA L+ YPAEL + + C ++ + L++ER + S
Sbjct: 265 LAG-LFLAADCVMASAYIIVQASILKKYPAELIVVFFYCFFVAIQSAVTCLVVER-DISA 438
Query: 231 WSLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAE 290
WSL +LLA +YSG+ S I + GPVFV+ F PL ++I ++ L +
Sbjct: 439 WSLEPKLRLLAVLYSGVFGSAFQVGIICWCLHQTGPVFVSMFKPLGILISVVLGVLFLGD 618
Query: 291 KIYLGRVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEILPAKQTIENNDKEKFHSHEVI 350
YLG +IGA VI++G Y V+WGK+KD + ++ + A +E N E H I
Sbjct: 619 AFYLGSLIGATVIVVGFYSVLWGKAKDIEDAGLSLESKGKQA-PLLEENSHEDIQGH*FI 795
Query: 351 TSSNFGAIAR 360
+ +++
Sbjct: 796 LKGCYNIVSK 825
>BI424342
Length = 426
Score = 136 bits (343), Expect = 1e-32
Identities = 66/133 (49%), Positives = 97/133 (72%), Gaps = 1/133 (0%)
Frame = +3
Query: 173 GSIMITIGCFSCACFTILQAVTLETYPA-ELSLTAWICLLGTVEGGIVALIMERGEPSVW 231
GSI++ I + A +LQA +ETY +LSLT+ IC +GT++ V +ME +PSVW
Sbjct: 30 GSILLIIATLAWASLFVLQAKAIETYKNHQLSLTSLICFIGTLQAIAVTFVMEH-KPSVW 206
Query: 232 SLSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEK 291
++ WD LLAA Y+GIV S + YY+QG+V++ +GPVF T F+PL M+IVAIM FIL+E+
Sbjct: 207 TIGWDMNLLAAAYAGIVTSSITYYVQGLVIKKKGPVFATAFSPLMMIIVAIMGSFILSEQ 386
Query: 292 IYLGRVIGAVVII 304
++LG V+GA++I+
Sbjct: 387 LFLGGVLGAILIV 425
>TC211179 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein;
91922-89607, partial (30%)
Length = 771
Score = 132 bits (332), Expect = 2e-31
Identities = 79/194 (40%), Positives = 112/194 (57%), Gaps = 7/194 (3%)
Frame = +2
Query: 131 ATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAVKGSIMITIGCF-------S 183
A ++GAM++T IKGP + + H + NG V HA G +M G S
Sbjct: 5 AGISGAMLLTFIKGPEVKMLSFHVNLFN-HRNGHVVHPHATSG-LMTIFGALASVASNVS 178
Query: 184 CACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWSLSWDTKLLAAV 243
A + I+QA E YP S TA + L+G V A +ER + S W L W+ +LL
Sbjct: 179 YAMWLIIQAKMSERYPCPYSSTALMSLMGAVLSISFAFCVER-DLSQWRLGWNIRLLTVA 355
Query: 244 YSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKIYLGRVIGAVVI 303
Y+GIV SG+ + +R RGP+FV+ F+PL +V+VA IL EK+YLG +IG+++I
Sbjct: 356 YAGIVVSGVMVAVISWCVRTRGPLFVSIFSPLMLVVVAFAGSTILDEKLYLGSIIGSMLI 535
Query: 304 ILGLYLVVWGKSKD 317
I GLY+V+WGKSK+
Sbjct: 536 ICGLYVVLWGKSKE 577
>TC215674 weakly similar to UP|Q9SUD5 (Q9SUD5) Medicago nodulin N21-like
protein, partial (28%)
Length = 1577
Score = 125 bits (313), Expect = 4e-29
Identities = 71/211 (33%), Positives = 117/211 (54%)
Frame = +3
Query: 113 EKIKMRSVHSQAKIVGTIATVAGAMVMTLIKGPILNLFGIHESSAQIQHNGGVNLQHAVK 172
EK+ + S+ S AKI+GT+ VAGA+ M L+KG L +H H G +
Sbjct: 588 EKVDI-SLRSTAKILGTVCCVAGALTMALVKGQKL----LHTEFLPSIHLTGSQGDDWLL 752
Query: 173 GSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWS 232
G +++ +C+ ILQ P L T W+CL T++ + AL+ E + W
Sbjct: 753 GCLLLLASSVFWSCWMILQVPITSCCPDHLLSTFWMCLFSTIQAALFALLSE-SDLQAWI 929
Query: 233 LSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKI 292
L ++ ++Y+GI + ++++IQ + RGP++ FNPL VI A++S L E++
Sbjct: 930 LQSPLQISCSLYAGIGIA-VSFFIQSWCISERGPLYCAMFNPLATVITALISATFLEEEV 1106
Query: 293 YLGRVIGAVVIILGLYLVVWGKSKDYDRPSP 323
Y+G ++GAV +I GLY+V+WGK+K++ P
Sbjct: 1107YVGSLVGAVGVIAGLYVVLWGKAKEFAEIKP 1199
>TC230350 weakly similar to UP|Q9C9F2 (Q9C9F2) MtN21-like protein;
91922-89607, partial (29%)
Length = 1057
Score = 123 bits (309), Expect = 1e-28
Identities = 63/165 (38%), Positives = 95/165 (57%)
Frame = +3
Query: 173 GSIMITIGCFSCACFTILQAVTLETYPAELSLTAWICLLGTVEGGIVALIMERGEPSVWS 232
G+I CFS A + +QA + YP S TA + G ++ ER + + W
Sbjct: 78 GAICSLASCFSFALWLTIQAKMSKEYPCHYSSTALMSTAGAIQATAFGFCFER-DLTQWK 254
Query: 233 LSWDTKLLAAVYSGIVCSGMAYYIQGVVMRYRGPVFVTTFNPLCMVIVAIMSPFILAEKI 292
L W+ +LLA YSGIV SG+ I ++ RGP+F + FNPL +V+VAI S +L E +
Sbjct: 255 LGWNIRLLAVAYSGIVASGIVVIITAWCIQMRGPLFASVFNPLMLVLVAIASSLMLNENL 434
Query: 293 YLGRVIGAVVIILGLYLVVWGKSKDYDRPSPIIKDEILPAKQTIE 337
Y+ V+GAV+I+ GLY+V+WGKSK+ + ++ E + + IE
Sbjct: 435 YVRSVVGAVLIVCGLYMVLWGKSKEMKNITQLVPSETIREAEAIE 569
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.328 0.140 0.437
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,385,751
Number of Sequences: 63676
Number of extensions: 284648
Number of successful extensions: 2198
Number of sequences better than 10.0: 111
Number of HSP's better than 10.0 without gapping: 2142
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2165
length of query: 363
length of database: 12,639,632
effective HSP length: 98
effective length of query: 265
effective length of database: 6,399,384
effective search space: 1695836760
effective search space used: 1695836760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135797.11


