

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135796.7 - phase: 0
(277 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
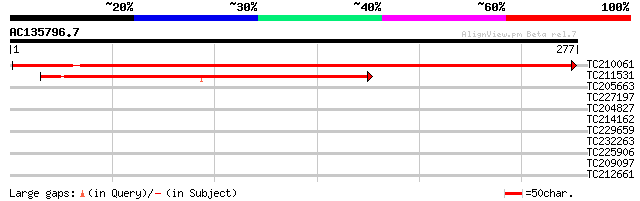
TC210061 similar to UP|Q6NQN0 (Q6NQN0) At1g53760, partial (53%) 405 e-113
TC211531 weakly similar to UP|Q6NQN0 (Q6NQN0) At1g53760, partial... 226 1e-59
TC205663 similar to UP|Q6PT61 (Q6PT61) Induced protein MgI1, par... 29 2.7
TC227197 similar to UP|Q84MB4 (Q84MB4) At1g22800, partial (71%) 29 2.7
TC204827 UP|Q9SLQ6 (Q9SLQ6) Actin isoform B, complete 28 3.5
TC214162 homologue to UP|Q41038 (Q41038) Type II chlorophyll a/b... 28 3.5
TC229659 28 3.5
TC232263 28 4.5
TC225906 similar to UP|Q6L6P7 (Q6L6P7) Cytochrome oxidase subuni... 28 4.5
TC209097 weakly similar to UP|O44936 (O44936) SpHbox7, partial (6%) 28 5.9
TC212661 similar to PIR|T05606|T05606 protein kinase homolog F9D... 27 7.8
>TC210061 similar to UP|Q6NQN0 (Q6NQN0) At1g53760, partial (53%)
Length = 827
Score = 405 bits (1041), Expect = e-113
Identities = 194/276 (70%), Positives = 232/276 (83%)
Frame = +3
Query: 2 KAKLIVFPIRGRNWCFTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAEL 61
+AKL VFPIRGRNWCF+R+IDH+L AS ++ SQSPSTLK LW +IN GDKP N K EL
Sbjct: 3 EAKLFVFPIRGRNWCFSRTIDHSLSASHAS---SQSPSTLKDLWTNINVGDKPLNTKTEL 173
Query: 62 FTDYVANKMNNGWVTLENAPDGSFKKKIHGLGLWLLSRVKPSEIFLKSISKDVTGVEVVY 121
F DY+ANKMNN W+ LE AP+GSFK KIHGLGL LLSRVKPSEIFLKSISK++T VE++Y
Sbjct: 174 FVDYIANKMNNAWIGLEKAPEGSFKNKIHGLGLRLLSRVKPSEIFLKSISKEITSVEIIY 353
Query: 122 PSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVPFFWILFRSYS 181
PSS+NA+LVRRRLRHIA+RG +IHR + YG VSLIPL+SA SILPLPNVPFFW+LFR+YS
Sbjct: 354 PSSLNAQLVRRRLRHIAVRGAVIHRNYLYGLVSLIPLTSALSILPLPNVPFFWVLFRTYS 533
Query: 182 HWRALQGSEKLFQLVSDGSQSSNTYSGKKETEHEDSENESLGLDEPHWVLTPSKELENIV 241
HWRALQGSE+LFQLVSD S++SNT + +K+TEH++S+++ +EP WVL PSKELEN+V
Sbjct: 534 HWRALQGSERLFQLVSDNSKTSNTCTYEKKTEHKESKSQRHSSNEPCWVLRPSKELENLV 713
Query: 242 RQEDGNDGLSRGTIEEICKIYDLNTQDVVKYEKSTF 277
EDG + LS+ I ICKIYDLN DV+KYEKS F
Sbjct: 714 HLEDGQESLSQHAIINICKIYDLNPVDVIKYEKSVF 821
>TC211531 weakly similar to UP|Q6NQN0 (Q6NQN0) At1g53760, partial (51%)
Length = 559
Score = 226 bits (575), Expect = 1e-59
Identities = 117/186 (62%), Positives = 138/186 (73%), Gaps = 24/186 (12%)
Frame = +1
Query: 16 CFTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAKAELFTDYVANKMNNGWV 75
CF+R+IDH+L S+S A SPSTLK LW +IN DKP NAKAELF DY+ANKMN+ W+
Sbjct: 4 CFSRTIDHSL-LSASHASSQSSPSTLKGLWTNINVRDKPLNAKAELFVDYIANKMNSVWI 180
Query: 76 TLENAPDGSFKKKIHGL------------------------GLWLLSRVKPSEIFLKSIS 111
LE AP+GSFK KIHGL GL LLSRVKPSEIFLKSIS
Sbjct: 181 GLEKAPEGSFKNKIHGLFFVFCFVVQW*SCLFLDFEFFCRLGLRLLSRVKPSEIFLKSIS 360
Query: 112 KDVTGVEVVYPSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILPLPNVP 171
K++T VE++YPSS+NA+LVRRRLRHIA+RG IIHRK+FYGSV +IPL++A ILPLPNVP
Sbjct: 361 KEITSVEIIYPSSLNAQLVRRRLRHIAVRGAIIHRKYFYGSVFVIPLTAACGILPLPNVP 540
Query: 172 FFWILF 177
FF +LF
Sbjct: 541 FFLVLF 558
>TC205663 similar to UP|Q6PT61 (Q6PT61) Induced protein MgI1, partial (60%)
Length = 1519
Score = 28.9 bits (63), Expect = 2.7
Identities = 24/81 (29%), Positives = 36/81 (43%), Gaps = 7/81 (8%)
Frame = +1
Query: 152 SVSLIP--LSSAFSILPLPNVPFFWILFRSYSHWRALQGSEKLFQLVSDGSQSSNTYSGK 209
S LIP L+SAFSILP P+ + S + L+G S +++ Y G
Sbjct: 250 SQPLIPPELASAFSILPEPHRTLLDVNRASRNTLSTLRGGGGSVHQAFSSSNNNHNYDGD 429
Query: 210 -----KETEHEDSENESLGLD 225
+E E +D + + G D
Sbjct: 430 GDGGVEEEEDDDDDRDGSGPD 492
>TC227197 similar to UP|Q84MB4 (Q84MB4) At1g22800, partial (71%)
Length = 1482
Score = 28.9 bits (63), Expect = 2.7
Identities = 9/27 (33%), Positives = 19/27 (70%)
Frame = +2
Query: 174 WILFRSYSHWRALQGSEKLFQLVSDGS 200
W++F SYS WR ++G++ + ++G+
Sbjct: 794 WLIFSSYSWWRNIKGAQNSLYVGTNGA 874
>TC204827 UP|Q9SLQ6 (Q9SLQ6) Actin isoform B, complete
Length = 1733
Score = 28.5 bits (62), Expect = 3.5
Identities = 18/64 (28%), Positives = 28/64 (43%), Gaps = 1/64 (1%)
Frame = -1
Query: 17 FTRSIDHTLPASSSTADFSQSPSTLKQLWKSINTGDKPFNAK-AELFTDYVANKMNNGWV 75
FT H L + D S +P+T+ LW S N +K ++ F V+N +
Sbjct: 1301 FTLRYPHLLEGAEGCKD*SSNPNTILSLWWSHNLNLHAARSKCSDFFAHTVSNTREHS*A 1122
Query: 76 TLEN 79
T +N
Sbjct: 1121 TTKN 1110
>TC214162 homologue to UP|Q41038 (Q41038) Type II chlorophyll a/b binding
protein from photosystem I precursor, partial (97%)
Length = 1285
Score = 28.5 bits (62), Expect = 3.5
Identities = 19/53 (35%), Positives = 25/53 (46%), Gaps = 8/53 (15%)
Frame = -2
Query: 154 SLIPLSSAFSILPLPNVPFFWIL--------FRSYSHWRALQGSEKLFQLVSD 198
S +PL +LP + PFFW L +H A+ G KLF LVS+
Sbjct: 234 SFLPLRKEAFVLPKSD-PFFWELGVEMATAAMAEEAHEEAMAGLMKLFSLVSE 79
>TC229659
Length = 882
Score = 28.5 bits (62), Expect = 3.5
Identities = 14/44 (31%), Positives = 25/44 (56%)
Frame = +2
Query: 224 LDEPHWVLTPSKELENIVRQEDGNDGLSRGTIEEICKIYDLNTQ 267
+DE W L N+V +DGND L +G + + C++ +L ++
Sbjct: 50 VDEHVWNL-------NVVTNDDGNDILIKGMVCDCCRLDELQSK 160
>TC232263
Length = 897
Score = 28.1 bits (61), Expect = 4.5
Identities = 12/30 (40%), Positives = 20/30 (66%)
Frame = +3
Query: 151 GSVSLIPLSSAFSILPLPNVPFFWILFRSY 180
G + L+PLS +++ P + FF++LFR Y
Sbjct: 729 GLILLLPLSDDDALVHNPKIFFFFLLFRIY 818
>TC225906 similar to UP|Q6L6P7 (Q6L6P7) Cytochrome oxidase subunit I
(Fragment), partial (10%)
Length = 851
Score = 28.1 bits (61), Expect = 4.5
Identities = 17/45 (37%), Positives = 23/45 (50%)
Frame = -3
Query: 122 PSSMNARLVRRRLRHIAMRGTIIHRKFFYGSVSLIPLSSAFSILP 166
P+S++ RL R RH T R+ S S PLS +F +LP
Sbjct: 435 PTSLSFRLDEERSRHGVCPWTARTRRLQASSRSTSPLSPSFFLLP 301
>TC209097 weakly similar to UP|O44936 (O44936) SpHbox7, partial (6%)
Length = 983
Score = 27.7 bits (60), Expect = 5.9
Identities = 16/53 (30%), Positives = 25/53 (46%)
Frame = -2
Query: 196 VSDGSQSSNTYSGKKETEHEDSENESLGLDEPHWVLTPSKELENIVRQEDGND 248
+SD S+ + G + ED ENES+G E K++ + R E+ D
Sbjct: 892 ISDSSEVDDEEYGLNVSSDEDCENESVGDSE------EKKDVAGLFRAEESRD 752
>TC212661 similar to PIR|T05606|T05606 protein kinase homolog F9D16.210 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(8%)
Length = 497
Score = 27.3 bits (59), Expect = 7.8
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = -2
Query: 136 HIAMRGTIIHRKFFYGSVSLIPLSSAFSIL 165
+I +RG IIH K+ GSV PL S ++
Sbjct: 277 NITLRGFIIHLKWLKGSVLNFPLGSQMILM 188
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.134 0.400
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,033,241
Number of Sequences: 63676
Number of extensions: 182432
Number of successful extensions: 1125
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 1118
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1122
length of query: 277
length of database: 12,639,632
effective HSP length: 96
effective length of query: 181
effective length of database: 6,526,736
effective search space: 1181339216
effective search space used: 1181339216
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135796.7


