

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135566.1 + phase: 0 /pseudo
(437 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
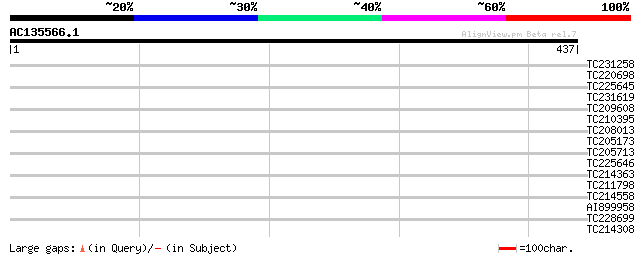
Score E
Sequences producing significant alignments: (bits) Value
TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kin... 35 0.087
TC220698 similar to GB|AAM67427.1|21689611|AY122894 At1g27650/T2... 31 0.96
TC225645 similar to UP|Q9LMB4 (Q9LMB4) F14D16.29, partial (64%) 30 1.6
TC231619 similar to UP|BRD3_HUMAN (Q15059) Bromodomain-containin... 30 2.8
TC209608 homologue to UP|Q6ILS4 (Q6ILS4) HDC08496, partial (10%) 29 3.7
TC210395 similar to UP|Q9FGR7 (Q9FGR7) Similarity to salt-induci... 29 3.7
TC208013 weakly similar to UP|Q8VYJ6 (Q8VYJ6) At2g30880/F7F1.9, ... 28 6.2
TC205173 homologue to UP|Q6K8Y4 (Q6K8Y4) Basic helix-loop-helix ... 28 6.2
TC205713 weakly similar to UP|Q6WCQ5 (Q6WCQ5) Block of prolifera... 28 6.2
TC225646 similar to UP|Q9LMB4 (Q9LMB4) F14D16.29, partial (88%) 28 6.2
TC214363 homologue to UP|Q8GTE3 (Q8GTE3) Ribosomal protein S3a, ... 28 8.1
TC211798 28 8.1
TC214558 homologue to UP|Q6FWH5 (Q6FWH5) Similarities with tr|Q0... 28 8.1
AI899958 28 8.1
TC228699 similar to GB|BAB72177.2|27808144|AB074573 phytosulfoki... 28 8.1
TC214308 homologue to UP|Q8GTE3 (Q8GTE3) Ribosomal protein S3a, ... 28 8.1
>TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kinase/response
regulator, partial (3%)
Length = 1374
Score = 34.7 bits (78), Expect = 0.087
Identities = 29/109 (26%), Positives = 43/109 (38%), Gaps = 6/109 (5%)
Frame = +2
Query: 64 LPTFDGR--GDPSEHVTVFNTRMSVYGVADSLKCKLLAGTFADA----ALRWYMSLPCFS 117
LP F G DP +H+ F+ S D + + F + A W L S
Sbjct: 545 LPKFHGLVGEDPHKHLKEFHIVCSTMKPPDVQEDHIFLKAFPHSLEGVAKDWLYYLAPRS 724
Query: 118 IMGYQDMIKKFTQQFSGTRHRKVLSTSLFNVRQGPSESLREYLARFNDS 166
I + D+ + F ++F + + +RQ ESL EY RF S
Sbjct: 725 IFNWDDLKRVFLEKFFPASRTTTIRKDISGIRQLSRESLYEYWERFKKS 871
>TC220698 similar to GB|AAM67427.1|21689611|AY122894 At1g27650/T22C5_2
{Arabidopsis thaliana;} , partial (70%)
Length = 995
Score = 31.2 bits (69), Expect = 0.96
Identities = 15/50 (30%), Positives = 23/50 (46%)
Frame = +2
Query: 213 YVKGEESNAEKKAREAKERGNSGGERRNHYVPPNRDRGTFKKPYERNQNR 262
Y + + + R ++ER SGG RR H PP R+ ++ NR
Sbjct: 728 YRRERRESDSRGGRRSRERDGSGGRRRQHGSPPAREGSEERRARIEQWNR 877
>TC225645 similar to UP|Q9LMB4 (Q9LMB4) F14D16.29, partial (64%)
Length = 1635
Score = 30.4 bits (67), Expect = 1.6
Identities = 13/46 (28%), Positives = 23/46 (49%)
Frame = -1
Query: 322 HLKREIEKLLLSGKLRGYAKERRHNERQEDKPNPEQKHTLHTISGG 367
H + +++ G GYA E RH R+ +K + + +H + GG
Sbjct: 279 HYGERVADVMMPGFGAGYAMEARHTPRRNEKIDGGRYRAIHFVFGG 142
>TC231619 similar to UP|BRD3_HUMAN (Q15059) Bromodomain-containing protein 3
(RING3-like protein), partial (3%)
Length = 421
Score = 29.6 bits (65), Expect = 2.8
Identities = 18/59 (30%), Positives = 27/59 (45%)
Frame = -1
Query: 219 SNAEKKAREAKERGNSGGERRNHYVPPNRDRGTFKKPYERNQNRYVPEHFTPLNTRPER 277
SN + K + KE+ E+ R RG +KP +R NR++P L PE+
Sbjct: 229 SNKKLKQKN*KEKKKKKKEKEMKEEGLKRKRGKKEKPRDRAVNRWIPRK*GYLRDSPEK 53
>TC209608 homologue to UP|Q6ILS4 (Q6ILS4) HDC08496, partial (10%)
Length = 808
Score = 29.3 bits (64), Expect = 3.7
Identities = 16/47 (34%), Positives = 20/47 (42%), Gaps = 4/47 (8%)
Frame = -2
Query: 34 HHGDLDEPEPQPFSAKIWNAPVPD----NFKPPHLPTFDGRGDPSEH 76
HH + P PQ + W+ P D + PP PT G G P H
Sbjct: 444 HHPHVRVPNPQHHLLRRWDFPDSDLGCESDPPPPEPTLPGSGAPKLH 304
>TC210395 similar to UP|Q9FGR7 (Q9FGR7) Similarity to salt-inducible protein,
partial (33%)
Length = 954
Score = 29.3 bits (64), Expect = 3.7
Identities = 9/26 (34%), Positives = 18/26 (68%)
Frame = +3
Query: 296 ARVMGQDKDAWCKYHLVQGHNTDDCV 321
+R + +D+++WCK H ++TD C+
Sbjct: 90 SRCILEDEESWCKTHFTVLYSTDPCI 167
>TC208013 weakly similar to UP|Q8VYJ6 (Q8VYJ6) At2g30880/F7F1.9, partial
(45%)
Length = 1065
Score = 28.5 bits (62), Expect = 6.2
Identities = 22/75 (29%), Positives = 36/75 (47%)
Frame = +2
Query: 157 REYLARFNDSTIKVSNPNQEVFVGAFQNGLWAGQFNESIAQKPADSMEEIIARAECYVKG 216
RE L R D+ I+ +N +W + + A++ +E + RAE V+
Sbjct: 209 REQLIRQRDAAIQEAN-------------MWRSELAK--AREHDVILEAAVVRAEEKVRV 343
Query: 217 EESNAEKKAREAKER 231
E+NAE + REA +R
Sbjct: 344 AEANAETRIREAVQR 388
>TC205173 homologue to UP|Q6K8Y4 (Q6K8Y4) Basic helix-loop-helix (BHLH)-like,
partial (29%)
Length = 1490
Score = 28.5 bits (62), Expect = 6.2
Identities = 17/53 (32%), Positives = 25/53 (47%), Gaps = 4/53 (7%)
Frame = +2
Query: 351 DKPNPEQKHTLHTISGGFAGG----GESSNSMKKYARQVMLLGDGPSGAYFPM 399
+ PNP K ++ T+ GFAG G+SSN + + P G+ PM
Sbjct: 236 ESPNPSGKASVQTLYNGFAGSLHGVGQSSNQTQHFQH--------PQGSSNPM 370
>TC205713 weakly similar to UP|Q6WCQ5 (Q6WCQ5) Block of proliferation
protein, partial (16%)
Length = 751
Score = 28.5 bits (62), Expect = 6.2
Identities = 10/45 (22%), Positives = 26/45 (57%), Gaps = 2/45 (4%)
Frame = +1
Query: 242 YVPPNRDRGTFKKPYERNQNRYVPEHFTPLNTRP--ERILKEVFE 284
Y+P + +++ +E ++ +++P+ F + + P E +K+ FE
Sbjct: 73 YIPTQEEINSYQLMFEEDRPKFIPKRFASMRSIPAYENAMKDCFE 207
>TC225646 similar to UP|Q9LMB4 (Q9LMB4) F14D16.29, partial (88%)
Length = 873
Score = 28.5 bits (62), Expect = 6.2
Identities = 12/41 (29%), Positives = 22/41 (53%)
Frame = -2
Query: 327 IEKLLLSGKLRGYAKERRHNERQEDKPNPEQKHTLHTISGG 367
+ +++ G GYA E RH R+ +K + + +H + GG
Sbjct: 347 VADVMMPGFGAGYAMEARHTPRRNEKIDGGRYRAIHFVFGG 225
>TC214363 homologue to UP|Q8GTE3 (Q8GTE3) Ribosomal protein S3a, complete
Length = 1070
Score = 28.1 bits (61), Expect = 8.1
Identities = 13/36 (36%), Positives = 23/36 (63%)
Frame = +3
Query: 326 EIEKLLLSGKLRGYAKERRHNERQEDKPNPEQKHTL 361
++E+L LS ++RG+ +ERR R+ P P + T+
Sbjct: 72 DLEELWLSERIRGFPRERRVARRKPLIPLPRKTGTI 179
>TC211798
Length = 728
Score = 28.1 bits (61), Expect = 8.1
Identities = 16/54 (29%), Positives = 27/54 (49%)
Frame = -1
Query: 93 LKCKLLAGTFADAALRWYMSLPCFSIMGYQDMIKKFTQQFSGTRHRKVLSTSLF 146
L C+ LA +L + SL CF+++ F+ FSG +H+K ++ F
Sbjct: 236 LVCEDLAAIPNSLSLSFLDSLSCFALIHLISC*TSFSYSFSGPQHKKRIAEK*F 75
>TC214558 homologue to UP|Q6FWH5 (Q6FWH5) Similarities with tr|Q07738
Saccharomyces cerevisiae YDL241w, partial (18%)
Length = 1101
Score = 28.1 bits (61), Expect = 8.1
Identities = 12/28 (42%), Positives = 18/28 (63%)
Frame = -1
Query: 218 ESNAEKKAREAKERGNSGGERRNHYVPP 245
++ KKAR+ K + ++ GERRNH P
Sbjct: 219 KNRKRKKARKIKSQKHTKGERRNHQKNP 136
>AI899958
Length = 416
Score = 28.1 bits (61), Expect = 8.1
Identities = 13/46 (28%), Positives = 21/46 (45%)
Frame = +2
Query: 244 PPNRDRGTFKKPYERNQNRYVPEHFTPLNTRPERILKEVFESKIIP 289
PP F ++ + ++Y P+ P P R +EV E + IP
Sbjct: 161 PPRSSTWDFLNFFDNSDDKYYPQTHYPATATPSRDSREVREEEGIP 298
>TC228699 similar to GB|BAB72177.2|27808144|AB074573 phytosulfokine precursor
3_2 {Arabidopsis thaliana;} , partial (42%)
Length = 596
Score = 28.1 bits (61), Expect = 8.1
Identities = 12/28 (42%), Positives = 17/28 (59%)
Frame = +2
Query: 201 DSMEEIIARAECYVKGEESNAEKKAREA 228
D MEE++ ECY+K EE + + EA
Sbjct: 197 DDMEELMGSEECYMKDEECISRRMMVEA 280
>TC214308 homologue to UP|Q8GTE3 (Q8GTE3) Ribosomal protein S3a, complete
Length = 1112
Score = 28.1 bits (61), Expect = 8.1
Identities = 13/36 (36%), Positives = 23/36 (63%)
Frame = +2
Query: 326 EIEKLLLSGKLRGYAKERRHNERQEDKPNPEQKHTL 361
++E+L LS ++RG+ +ERR R+ P P + T+
Sbjct: 140 DLEELWLSERIRGFPRERRVARRKPLIPLPRRTGTI 247
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.134 0.407
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,570,382
Number of Sequences: 63676
Number of extensions: 308401
Number of successful extensions: 1725
Number of sequences better than 10.0: 33
Number of HSP's better than 10.0 without gapping: 1685
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1725
length of query: 437
length of database: 12,639,632
effective HSP length: 100
effective length of query: 337
effective length of database: 6,272,032
effective search space: 2113674784
effective search space used: 2113674784
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135566.1


