

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135565.5 + phase: 0 /pseudo
(93 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
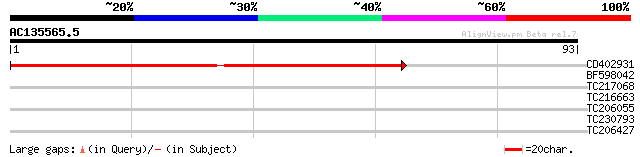
CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (ja... 95 7e-21
BF598042 27 2.2
TC217068 similar to UP|Q93Z30 (Q93Z30) AT4g18950/F13C5_120 (Prot... 26 3.8
TC216663 UP|C823_SOYBN (O49858) Cytochrome P450 82A3 (P450 CP6)... 26 3.8
TC206055 similar to PRF|1604369A.0|226743|1604369A sulfated surf... 25 8.4
TC230793 UP|O06921 (O06921) MadZ, partial (12%) 25 8.4
TC206427 similar to UP|Q8S9I6 (Q8S9I6) AT5g55180/MCO15_13, parti... 25 8.4
>CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 617
Score = 94.7 bits (234), Expect = 7e-21
Identities = 45/65 (69%), Positives = 56/65 (85%)
Frame = +2
Query: 1 MMNRFCFVALDKTLRNVMGSENKDNKTKPFGGKVFVLVGDFRQVLPVVRVATRQQIVSSA 60
+MN+FCF ALD+TL+++M S+NKDN TKPFGGKV VL GDFRQ+LPV+R +RQ IV SA
Sbjct: 251 VMNKFCFEALDRTLQDIMASQNKDNATKPFGGKV-VLGGDFRQILPVIRKGSRQDIVGSA 427
Query: 61 VNASK 65
+NASK
Sbjct: 428 INASK 442
>BF598042
Length = 292
Score = 26.6 bits (57), Expect = 2.2
Identities = 11/28 (39%), Positives = 17/28 (60%)
Frame = -3
Query: 6 CFVALDKTLRNVMGSENKDNKTKPFGGK 33
C++ D +R V G E +D + K FGG+
Sbjct: 149 CYIRGDDNIRPVSGREWQDYRVK*FGGR 66
>TC217068 similar to UP|Q93Z30 (Q93Z30) AT4g18950/F13C5_120 (Protein
kinase-like protein), partial (88%)
Length = 1650
Score = 25.8 bits (55), Expect = 3.8
Identities = 11/32 (34%), Positives = 21/32 (65%), Gaps = 2/32 (6%)
Frame = +3
Query: 62 NASKVWRRC--EVLRLTMNMGLSTANNNADQE 91
NASK+WR C E++++ + + + A + A+ E
Sbjct: 1269 NASKIWRPC*REIIQMEVVVAVHLAPHQAEYE 1364
>TC216663 UP|C823_SOYBN (O49858) Cytochrome P450 82A3 (P450 CP6) , complete
Length = 1814
Score = 25.8 bits (55), Expect = 3.8
Identities = 8/36 (22%), Positives = 19/36 (52%)
Frame = -1
Query: 52 TRQQIVSSAVNASKVWRRCEVLRLTMNMGLSTANNN 87
+ ++ + +++W RC +LRL N ++ N+
Sbjct: 548 SNSSLIEVRTSETRMWFRCSILRLERNSNVTILRNS 441
>TC206055 similar to PRF|1604369A.0|226743|1604369A sulfated surface
glycoprotein SSG185. {Volvox carteri;} , partial (6%)
Length = 824
Score = 24.6 bits (52), Expect = 8.4
Identities = 13/36 (36%), Positives = 16/36 (44%)
Frame = -1
Query: 14 LRNVMGSENKDNKTKPFGGKVFVLVGDFRQVLPVVR 49
LRN + K KPF +F L F+Q P R
Sbjct: 602 LRNKKNETKEGKKKKPFCSIIFPLEAPFQQCRPPPR 495
>TC230793 UP|O06921 (O06921) MadZ, partial (12%)
Length = 1100
Score = 24.6 bits (52), Expect = 8.4
Identities = 9/18 (50%), Positives = 13/18 (72%)
Frame = -3
Query: 63 ASKVWRRCEVLRLTMNMG 80
A KVWR C++ RL ++G
Sbjct: 462 ALKVWRNCKI*RLIKDLG 409
>TC206427 similar to UP|Q8S9I6 (Q8S9I6) AT5g55180/MCO15_13, partial (89%)
Length = 1660
Score = 24.6 bits (52), Expect = 8.4
Identities = 7/16 (43%), Positives = 12/16 (74%)
Frame = +2
Query: 56 IVSSAVNASKVWRRCE 71
I+ S +N +K+W+ CE
Sbjct: 71 IIRSGINRNKLWKNCE 118
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.133 0.380
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,336,049
Number of Sequences: 63676
Number of extensions: 27268
Number of successful extensions: 127
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 126
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 126
length of query: 93
length of database: 12,639,632
effective HSP length: 69
effective length of query: 24
effective length of database: 8,245,988
effective search space: 197903712
effective search space used: 197903712
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC135565.5


