

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
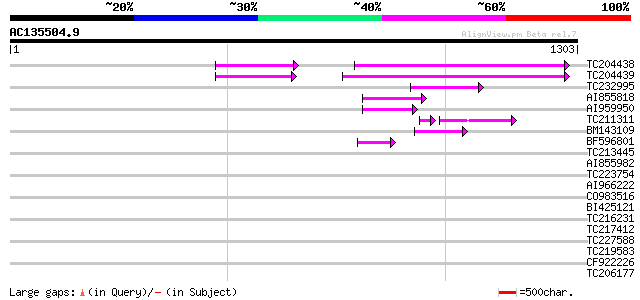
Query= AC135504.9 + phase: 0 /pseudo
(1303 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 131 2e-30
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 122 1e-27
TC232995 77 5e-14
AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza ... 72 1e-12
AI959950 64 4e-10
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 56 1e-08
BM143109 54 3e-07
BF596801 weakly similar to GP|29423270|g gag-pol polyprotein {Gl... 44 5e-04
TC213445 41 0.003
AI855982 40 0.007
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 39 0.020
AI966222 37 0.074
CO983516 35 0.17
BI425121 35 0.28
TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (G... 34 0.48
TC217412 similar to UP|Q6L724 (Q6L724) ATP-dependent RNA helicas... 33 0.63
TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L1... 33 0.63
TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich p... 33 0.63
CF922226 33 0.63
TC206177 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (G... 33 1.1
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 131 bits (330), Expect = 2e-30
Identities = 141/493 (28%), Positives = 237/493 (47%)
Frame = +3
Query: 793 N*A*NC*RSSLR*WMDISYARRAKSVPKE*CVRSDTQT*SEEHY*NKMGIQKQAE*TRRG 852
N*A C R + * +D YARR ++ KE* + + + T +++ +Q+Q +* R
Sbjct: 3198 N*AQECERGTD**VLDQCYARRIGAIQKE*SLGASS*TRGN*CDWHQVDLQEQNQ*RRCY 3377
Query: 853 NQKQS*TCCTRIQSTRRH*LH*NICSSCKIGSNQVTSILRS*SWHNIISNGCQECFS*WC 912
NQKQ TCC+R+ S R L *N C +Q+ + + +GC+E S W
Sbjct: 3378 NQKQGQTCCSRLHSD*RCRL**NFRPCC*T*VHQIVTWCSLHPQIQAVPDGCEERVSEWI 3557
Query: 913 H*RRSVCKTTSWV*GS*AS*PCL*T*EITIWLETSS*SLV**TK*FLN*K*F*KRIS*RN 972
RS+C + S +S C+ E ++W+E SS SLV* *+ +*++
Sbjct: 3558 PE*RSLCGAAKGICRSNSSRSCIQAQEGSLWIEASSKSLV*KANRVPYSARV*EGRN*QD 3737
Query: 973 TLQKDS*ERHSDCANIC**YNIWFY*CISLQGIF*VNAG*I*NEHDGRIEVLSWNSNQPK 1032
+L + + D +IC**+ +W +A *I*+E R ++ S ++
Sbjct: 3738 SLCQTRC*KLDDSTDIC**HCVWRDVE*DASTFCPTDAI*I*DESCWRADLFSGTPSEAD 3917
Query: 1033 QRRSICSSNKIYEGASEEVQARRL*SDEHSNASNLHLKQRRYWNYSRPEAIQRYDWLSVI 1092
R I + ++ + +EV + +++ +L +R W+ +++Q++DW I
Sbjct: 3918 GRLHIPLTKQVCKEHCQEVWDGKCQP*KNTCTYSLEAVKR*SWHQC*SKSVQKHDWELTI 4097
Query: 1093 PHCI*T*YFI*CMLVCKISIRS*RISFNCS*ENPQASERNN*SWTPV*EIPRL*VDWIL* 1152
+ T*+ + +CKIS +S* S S EN + + + * W V + R W+L*
Sbjct: 4098 FNS*QT*HHLCSRCLCKISSQS*DKSLESSKENSEICKWHQ*LWDYVLSLFRFNAGWVL* 4277
Query: 1153 C*LCW**D*KKINQWKLSIPGRESDILGKQKTSNYCYVYSRSRIHFSCKLLYTTTLDETS 1212
C*L W *+K + W + + G +S + +Q+ +Y SR++ S K L+TT+LDE
Sbjct: 4278 C*LGWKCR*QKKHFWWMFLFGNQSYFMVQQEAELCVPIYC*SRVYCSRKQLFTTSLDEAD 4457
Query: 1213 VGRLSD*C*QYSHFL**YYCYLFVKESNSTF*SQAY*DQTPLYQRLCSKRNFRYTIH*Y* 1272
+ + L* + CY + +S ST +QA+* T LY R C ++ *+*
Sbjct: 4458 AEGVQCRTRCHDIVL*QHECY*YF*KSCSTQQNQAH*H*TSLY*RSC***SYHTGAC*H* 4637
Query: 1273 ASMG*YIYKAFIC 1285
+ Y +K C
Sbjct: 4638 GTNSRYFHKGIGC 4676
Score = 52.0 bits (123), Expect = 2e-06
Identities = 55/190 (28%), Positives = 93/190 (48%)
Frame = +3
Query: 473 C**LQQMDLGKIH*K*RLCM*SV*QLLHSNTI*KRIENFESQK*SWWRI*K*AI*IFL*K 532
C * Q+ LG+++ + +*S+ + + KR+ + E+Q+* W R+*K + L
Sbjct: 2334 CG*FLQIYLGQLYQREIRHL*SIQGVESKTSKRKRLCHQENQE*PWQRV*KQQVY*ILHI 2513
Query: 533 TWDSP*VFFS*NSTTKWGCREKKQNFTRNGQNHDP*KQFS*TFLGRSS*YFMLYSK*DLY 592
*V S +TTKW ++KQ+F R+ H ++ S LG S + ML+ +
Sbjct: 2514 *RHHS*VLCSHYTTTKWHS*KEKQDFARSC*GHASCQRTSL*SLG*SHEHSMLHPQQSHT 2693
Query: 593 QTYVGENCI*TL*RKKTQYLLLSSVWMYLLHFKH*RLSEEI*CQGSKRNLFRLL*KVKGV 652
+ +* L R++ L +W +LHF *R E+ Q RN+ +L K + +
Sbjct: 2694 *KRDSNHTV*NLEREEANCQALPHLWKSMLHFGR*RAKEKDGSQE*CRNILGILYKQQSI 2873
Query: 653 QSV*FRNTLC 662
S+ F+N C
Sbjct: 2874 *SIQFQNQNC 2903
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 122 bits (306), Expect = 1e-27
Identities = 144/521 (27%), Positives = 245/521 (46%)
Frame = +3
Query: 765 ANHWKQRQS*KNKISLQTRRIFDRIAFNN*A*NC*RSSLR*WMDISYARRAKSVPKE*CV 824
A++ + +Q KI R+ N*A C R + R* +D YARR ++ KE* +
Sbjct: 3111 ADYRRSKQRGHYKIKGG*DRLKLMFCLQN*AQECERGTDR*VLDQCYARRIGAIQKE*SL 3290
Query: 825 RSDTQT*SEEHY*NKMGIQKQAE*TRRGNQKQS*TCCTRIQSTRRH*LH*NICSSCKIGS 884
+ + * +++ +Q+Q +* R NQKQ T C+R+ S R L *+ C SC
Sbjct: 3291 GASS*A*GN*CDWHQVDLQEQNQ*RRCHNQKQGQTGCSRLHSD*RCRL**DFCPSC*T*V 3470
Query: 885 NQVTSILRS*SWHNIISNGCQECFS*WCH*RRSVCKTTSWV*GS*AS*PCL*T*EITIWL 944
+Q+ + + +GC+E S W RS+C + +S C+ E ++W+
Sbjct: 3471 HQIITWCSLYPQIQAVPDGCEERISEWIPE*RSLCGAAKGICRPDSSRSCIQAQEGSLWI 3650
Query: 945 ETSS*SLV**TK*FLN*K*F*KRIS*RNTLQKDS*ERHSDCANIC**YNIWFY*CISLQG 1004
E SS SLV* *+ +*++ L + + DC +IC**+ +W
Sbjct: 3651 EASSKSLV*KANRVPYSARV*EGRN*QDPLCQTRC*KLDDCTDIC**HCVWRDVE*DAST 3830
Query: 1005 IF*VNAG*I*NEHDGRIEVLSWNSNQPKQRRSICSSNKIYEGASEEVQARRL*SDEHSNA 1064
+A *I*+E R ++ S S++ I + ++ + +EV S + +
Sbjct: 3831 FCSTDAI*I*DESCWRADLFSGTSSEADGGLHIPLTKQVCKEHCQEVWDGECQS*KDTCT 4010
Query: 1065 SNLHLKQRRYWNYSRPEAIQRYDWLSVIPHCI*T*YFI*CMLVCKISIRS*RISFNCS*E 1124
+L + + +++Q++D I + T + + +CKIS +S S + S E
Sbjct: 4011 YSLEAVKG*SRHQC*SKSVQKHDRELTIFNS*QTRHHLCSRCLCKISSQSQDKSLDSSKE 4190
Query: 1125 NPQASERNN*SWTPV*EIPRL*VDWIL*C*LCW**D*KKINQWKLSIPGRESDILGKQKT 1184
N + + + * W V + + W+L*C*L W *+K + W + + G++ + +Q+
Sbjct: 4191 NSEICKWH**LWDYVLSLFKSNAGWVL*C*LGWKCR*QKKHFWWMLLFGKQPYFMVQQEA 4370
Query: 1185 SNYCYVYSRSRIHFSCKLLYTTTLDETSVGRLSD*C*QYSHFL**YYCYLFVKESNSTF* 1244
+YSRSR++ S K L+T +LDE + + L* + CY + +S ST
Sbjct: 4371 ELCVPIYSRSRVYCSRKQLFTASLDEADAEGVQCRTRCHDIVL*QHECY*YF*KSCSTQQ 4550
Query: 1245 SQAY*DQTPLYQRLCSKRNFRYTIH*Y*ASMG*YIYKAFIC 1285
+QA+* T LYQR C + *+* + Y +K F C
Sbjct: 4551 NQAH*H*TSLYQRSC***SDHTEAC*H*GTNSRYFHKGFGC 4673
Score = 48.1 bits (113), Expect = 2e-05
Identities = 53/187 (28%), Positives = 92/187 (48%)
Frame = +3
Query: 473 C**LQQMDLGKIH*K*RLCM*SV*QLLHSNTI*KRIENFESQK*SWWRI*K*AI*IFL*K 532
C * Q+ LGK++ + +*S+ ++ + +R+ + E+Q+* W RI*K + L
Sbjct: 2331 CG*FLQIYLGKLYQREIRNL*SIQRVESKTSKRERLCHQENQE*PWQRI*KQQVH*ILHI 2510
Query: 533 TWDSP*VFFS*NSTTKWGCREKKQNFTRNGQNHDP*KQFS*TFLGRSS*YFMLYSK*DLY 592
*V S +TT+W E+KQ+F R H ++ S LG S + ML+ +
Sbjct: 2511 *RHHS*VLCSHYTTTEWDS*EEKQDFARGCSGHASCQRTSL*SLG*SHEHSMLHPQQSHT 2690
Query: 593 QTYVGENCI*TL*RKKTQYLLLSSVWMYLLHFKH*RLSEEI*CQGSKRNLFRLL*KVKGV 652
+ + +* L R++ L +W +LH *R ++ Q RN+ +L K + +
Sbjct: 2691 EKRDSNHPV*NLEREEAICQALPHLWKSMLHLGR*RAKKKDGSQE*CRNIPGILYKQQSI 2870
Query: 653 QSV*FRN 659
S+ F+N
Sbjct: 2871 *SIQFQN 2891
>TC232995
Length = 1009
Score = 77.0 bits (188), Expect = 5e-14
Identities = 62/168 (36%), Positives = 92/168 (53%)
Frame = +1
Query: 921 TTSWV*GS*AS*PCL*T*EITIWLETSS*SLV**TK*FLN*K*F*KRIS*RNTLQKDS*E 980
TT W * * + PCL* + ++W ETS * +V* K*F +*K +R S + + K+
Sbjct: 7 TTPWF*NF**TKPCL*ITKGSLWFETSP*GMV*TIK*FSS*KRILQR*SGYHIIHKEKA* 186
Query: 981 RHSDCANIC**YNIWFY*CISLQGIF*VNAG*I*NEHDGRIEVLSWNSNQPKQRRSICSS 1040
+ +NIC**YN W +* +QG+F A *I*N +DGR +VLS +NQ R I S
Sbjct: 187 *YFVGSNIC**YNFWIH**FIVQGVFP*YAK*I*NVNDGRTKVLSGITNQANSIRYIHQS 366
Query: 1041 NKIYEGASEEVQARRL*SDEHSNASNLHLKQRRYWNYSRPEAIQRYDW 1088
+I +G +++ + +++ L L+ R W+ R + I R W
Sbjct: 367 IQILQGIDQKIWDG*CKTHVYTDEH*LLLR*R*IWSVYRHKTISRCYW 510
>AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 463
Score = 72.4 bits (176), Expect = 1e-12
Identities = 58/148 (39%), Positives = 81/148 (54%)
Frame = -2
Query: 811 YARRAKSVPKE*CVRSDTQT*SEEHY*NKMGIQKQAE*TRRGNQKQS*TCCTRIQSTRRH 870
+ARR +S+ K+*CV + +T* + NKMG K *T K * R+ S R +
Sbjct: 456 HARRTESI*KK*CVETSRKT*KLSCHRNKMGF*K*IR*TWHNY*K*G*ISSKRV*SRRGN 277
Query: 871 *LH*NICSSCKIGSNQVTSILRS*SWHNIISNGCQECFS*WCH*RRSVCKTTSWV*GS*A 930
L NICSSCKI S+ + + +SNGC +CFS W + RRS+C TT +*
Sbjct: 276 RL*RNICSSCKIRSH*NAFSICIHNEF*TLSNGC*KCFSKWFNSRRSIC*TTPRL*NPG* 97
Query: 931 S*PCL*T*EITIWLETSS*SLV**TK*F 958
+ CL* + ++W +TS * +V* K F
Sbjct: 96 TNSCL*IAKGSLWFKTSP*GVV*TYKQF 13
>AI959950
Length = 466
Score = 63.9 bits (154), Expect = 4e-10
Identities = 45/128 (35%), Positives = 69/128 (53%)
Frame = -3
Query: 810 SYARRAKSVPKE*CVRSDTQT*SEEHY*NKMGIQKQAE*TRRGNQKQS*TCCTRIQSTRR 869
S ARR SV KE*C+ + T +E ++M I Q G + QS C R+ +T R
Sbjct: 392 SDARRT*SVSKE*CLEAR*ITKKKEGSWSEMDIL*QTRRGW*GCEIQSKISC*RLLTTGR 213
Query: 870 H*LH*NICSSCKIGSNQVTSILRS*SWHNIISNGCQECFS*WCH*RRSVCKTTSWV*GS* 929
+ L N+C+ C SN + + + + ++SNGC++C S W + + S+C TT+W+*
Sbjct: 212 YRLPKNLCTCCTFRSNMHLTFICNL**YEVVSNGCKKCISKWLNPKGSLC*TTAWI*K*N 33
Query: 930 AS*PCL*T 937
S C *T
Sbjct: 32 PSSTCF*T 9
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 55.8 bits (133), Expect(2) = 1e-08
Identities = 62/179 (34%), Positives = 94/179 (51%), Gaps = 3/179 (1%)
Frame = +2
Query: 988 IC**YNIWFY*CISLQGIF*VNAG*I*NEHDGRIEVLSWNSNQPKQRRSICSSNKIYEGA 1047
+C**+N+W +QG+F*VN G I*NE++ +V SN + S +IY+
Sbjct: 503 LC**HNLWCNLKKDVQGVF*VNEGWI*NEYER*AKVPPRTSNHSESLWDFYPSREIYKVP 682
Query: 1048 SEEVQARRL*SDEHSN---ASNLHLKQRRYWNYSRPEAIQRYDWLSVIPHCI*T*YFI*C 1104
S++VQ *S + N + + H + +Y + + YD +I + *T Y +
Sbjct: 683 SKKVQ--NG*SQTYGNPYASFHNH*QG*ER*SYF-IKGV*WYD*FFIIFNF**TRYCVCR 853
Query: 1105 MLVCKISIRS*RISFNCS*ENPQASERNN*SWTPV*EIPRL*VDWIL*C*LCW**D*KK 1163
+ +CKIS+ S S CS*++ + S N *S + V*E +* IL*C CW** K+
Sbjct: 854 LPLCKISVLSKNFSCYCS*KDLKISCWNY*SLSMV*EKV*V*SFRIL*CLFCW**SRKE 1030
Score = 22.7 bits (47), Expect(2) = 1e-08
Identities = 16/37 (43%), Positives = 21/37 (56%)
Frame = +1
Query: 942 IWLETSS*SLV**TK*FLN*K*F*KRIS*RNTLQKDS 978
+W ETSS SLV* K + K +R + T+QK S
Sbjct: 364 LWFETSSKSLV*KAKFISSFKWIHQRNNGPRTIQKGS 474
>BM143109
Length = 415
Score = 54.3 bits (129), Expect = 3e-07
Identities = 46/121 (38%), Positives = 65/121 (53%)
Frame = +3
Query: 931 S*PCL*T*EITIWLETSS*SLV**TK*FLN*K*F*KRIS*RNTLQKDS*ERHSDCANIC* 990
+* CL*T + IW++TS * LV* +* + F KR * E ++ +IC*
Sbjct: 33 A*SCL*TEKGFIWIKTSP*GLV*TFE*ISFRQGFFKR*G*Y*PFYLKEIE*YTLSTDIC* 212
Query: 991 *YNIWFY*CISLQGIF*VNAG*I*NEHDGRIEVLSWNSNQPKQRRSICSSNKIYEGASEE 1050
*Y WF * SLQ IF A *I*N +D +++ SW +NQ + +I S KI + +
Sbjct: 213 *YYFWFN**FSLQKIFSRYAK*I*NVNDA*VKLFSWTTNQANKEWNIYQSIKILQRPDSQ 392
Query: 1051 V 1051
+
Sbjct: 393 I 395
>BF596801 weakly similar to GP|29423270|g gag-pol polyprotein {Glycine max},
partial (7%)
Length = 336
Score = 43.9 bits (102), Expect = 5e-04
Identities = 36/86 (41%), Positives = 44/86 (50%)
Frame = +2
Query: 800 RSSLR*WMDISYARRAKSVPKE*CVRSDTQT*SEEHY*NKMGIQKQAE*TRRGNQKQS*T 859
RS R* +D +ARR K + K+ C+ +T* Y NKMG K *T K S
Sbjct: 20 RSHSR**LDHCHARRTKPI*KKQCMEISRKT*KLSCYWNKMGF*K*IR*TWYNY*K*SQV 199
Query: 860 CCTRIQSTRRH*LH*NICSSCKIGSN 885
RI S R + L NICS CKI S+
Sbjct: 200 SSERI*SRRGNRL*RNICSCCKIRSH 277
>TC213445
Length = 705
Score = 41.2 bits (95), Expect = 0.003
Identities = 30/79 (37%), Positives = 44/79 (54%)
Frame = +3
Query: 1176 SDILGKQKTSNYCYVYSRSRIHFSCKLLYTTTLDETSVGRLSD*C*QYSHFL**YYCYLF 1235
S I+ K C + RSRI+F KLL T LDET+ L * Y++ +* Y C
Sbjct: 450 SSIMA**KAK*CCLINCRSRIYFC*KLLCTNLLDETTTF*LWFET*SYTYPM*QYKCN*S 629
Query: 1236 VKESNSTF*SQAY*DQTPL 1254
+++S S *++AY*++ L
Sbjct: 630 IQKSYSVL*NKAY*NKASL 686
Score = 31.6 bits (70), Expect = 2.4
Identities = 16/28 (57%), Positives = 20/28 (71%)
Frame = +1
Query: 1098 T*YFI*CMLVCKISIRS*RISFNCS*EN 1125
T Y +*C+ VCKIS +S RIS C *+N
Sbjct: 232 TSYNV*CLYVCKISSKSQRISPKCH*KN 315
>AI855982
Length = 484
Score = 40.0 bits (92), Expect = 0.007
Identities = 34/86 (39%), Positives = 46/86 (52%)
Frame = +1
Query: 800 RSSLR*WMDISYARRAKSVPKE*CVRSDTQT*SEEHY*NKMGIQKQAE*TRRGNQKQS*T 859
RS R* +D +ARR +S+ K+*CV + +T* + NKMG+ K *T *T
Sbjct: 115 RSHSR**LDNCHARRTESI*KK*CVETSRKT**LSCHMNKMGL*K*IR*TSHNYYT*G*T 294
Query: 860 CCTRIQSTRRH*LH*NICSSCKIGSN 885
R+ S+RR L IC C I S+
Sbjct: 295 SSRRL*SSRRTRL*TYICFYC*IISH 372
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 38.5 bits (88), Expect = 0.020
Identities = 19/39 (48%), Positives = 22/39 (55%)
Frame = +2
Query: 185 GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADC 223
GGRF G G GG K++ CFNC KPGHF +C
Sbjct: 368 GGRFGGGGGSGGGGGKST-------CFNCGKPGHFAREC 463
>AI966222
Length = 430
Score = 36.6 bits (83), Expect = 0.074
Identities = 23/66 (34%), Positives = 40/66 (59%)
Frame = +3
Query: 561 NGQNHDP*KQFS*TFLGRSS*YFMLYSK*DLYQTYVGENCI*TL*RKKTQYLLLSSVWMY 620
+G NH * * LG S+ Y ML S+ +LY+T++ ++ +* + KTQ+++ S +
Sbjct: 3 DG*NHAK**LNP*ALLG*SNEYCMLSSEQNLYKTHLEKDSL*IMEGTKTQHIIFLSF*V* 182
Query: 621 LLHFKH 626
+ H+KH
Sbjct: 183 VFHYKH 200
>CO983516
Length = 724
Score = 35.4 bits (80), Expect = 0.17
Identities = 29/96 (30%), Positives = 49/96 (50%)
Frame = +1
Query: 900 ISNGCQECFS*WCH*RRSVCKTTSWV*GS*AS*PCL*T*EITIWLETSS*SLV**TK*FL 959
+ +GC+E S W RS+C + S +S C+ E ++W+E SS SLV*
Sbjct: 436 VPDGCEERVSEWIPE*RSLCGAAKGIYRSNSSRSCIQAQEGSLWIEASSKSLV*KANRVT 615
Query: 960 N*K*F*KRIS*RNTLQKDS*ERHSDCANIC**YNIW 995
*+ +*+++L + + D +IC**+ +W
Sbjct: 616 YSARV*EGRN*QDSLCQTRC*KLDDSTDIC**HCVW 723
>BI425121
Length = 412
Score = 34.7 bits (78), Expect = 0.28
Identities = 28/86 (32%), Positives = 44/86 (50%)
Frame = +2
Query: 529 FL*KTWDSP*VFFS*NSTTKWGCREKKQNFTRNGQNHDP*KQFS*TFLGRSS*YFMLYSK 588
FL*K W *+ + N + K G ++K NF R G +H S + +L SK
Sbjct: 197 FL*KEWYFS*LVLTENPSRK*GS*KEK*NFARKG*DH--------------SFHNLLCSK 334
Query: 589 *DLYQTYVGENCI*TL*RKKTQYLLL 614
++ +T ++ +*T+ RKKT Y +L
Sbjct: 335 QNIDKTNDKKDSL*TMERKKTHYFIL 412
>TC216231 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (GRP2)
(AT4g38680/F20M13_240), partial (42%)
Length = 891
Score = 33.9 bits (76), Expect = 0.48
Identities = 17/40 (42%), Positives = 19/40 (47%)
Frame = +3
Query: 185 GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCP 224
GG RYG G GG C+NC + GHF DCP
Sbjct: 285 GGGGRYGGGGGGGG----------SCYNCGESGHFARDCP 374
Score = 31.2 bits (69), Expect = 3.1
Identities = 14/39 (35%), Positives = 17/39 (42%)
Frame = +3
Query: 185 GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADC 223
GG + G G GGY GC+ C + GH DC
Sbjct: 156 GGGYGGGGYGGGGGYGGGGGGGGGGCYKCGETGHIARDC 272
>TC217412 similar to UP|Q6L724 (Q6L724) ATP-dependent RNA helicase, partial
(3%)
Length = 868
Score = 33.5 bits (75), Expect = 0.63
Identities = 16/39 (41%), Positives = 24/39 (61%), Gaps = 4/39 (10%)
Frame = +3
Query: 197 GGYKNSR--KEDQKG--CFNCKKPGHFIADCPDLQKEKS 231
GG ++SR D+ G CFNC + GH +DCP++ +S
Sbjct: 261 GGRRSSRPSSSDRFGGTCFNCGESGHRASDCPNVSNRRS 377
>TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L16.11 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(10%)
Length = 1300
Score = 33.5 bits (75), Expect = 0.63
Identities = 17/40 (42%), Positives = 21/40 (52%)
Frame = +1
Query: 194 LGRGGYKNSRKEDQKGCFNCKKPGHFIADCPDLQKEKSKS 233
LG + S+ +D CF CKK GH DCP+ SKS
Sbjct: 145 LGHNARQCSKVQD---CFICKKGGHRAKDCPEKHTSTSKS 255
>TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich protein 2,
partial (45%)
Length = 995
Score = 33.5 bits (75), Expect = 0.63
Identities = 11/20 (55%), Positives = 15/20 (75%)
Frame = +2
Query: 209 GCFNCKKPGHFIADCPDLQK 228
GCFNC + GHF +CP++ K
Sbjct: 542 GCFNCGEEGHFARECPNVGK 601
Score = 30.4 bits (67), Expect = 5.3
Identities = 15/39 (38%), Positives = 20/39 (50%)
Frame = +2
Query: 185 GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADC 223
GGR+ G+ GRG R+ C+NC + GH DC
Sbjct: 359 GGRYGGGEGRGRGF---GRRGGGPECYNCGRIGHLARDC 466
>CF922226
Length = 667
Score = 33.5 bits (75), Expect = 0.63
Identities = 17/49 (34%), Positives = 23/49 (46%), Gaps = 9/49 (18%)
Frame = -3
Query: 200 KNSRKEDQKG---------CFNCKKPGHFIADCPDLQKEKSKSRPKKQS 239
K + E+QK C++CKK GH CP+ QK + KK S
Sbjct: 299 KKQKPENQKNGEGNIFKIRCYHCKKEGHTRKVCPERQKNGGSNNRKKDS 153
>TC206177 similar to UP|Q41188 (Q41188) Glycine-rich protein 2 (GRP2)
(AT4g38680/F20M13_240), partial (22%)
Length = 406
Score = 32.7 bits (73), Expect = 1.1
Identities = 17/40 (42%), Positives = 20/40 (49%)
Frame = +1
Query: 185 GGRFRYGKVLGRGGYKNSRKEDQKGCFNCKKPGHFIADCP 224
GG RYG G GG S C++C + GHF DCP
Sbjct: 52 GGGGRYGGGGGGGGGGGS-------CYSCGESGHFARDCP 150
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.382 0.170 0.717
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 59,602,807
Number of Sequences: 63676
Number of extensions: 895109
Number of successful extensions: 14660
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 4453
Number of HSP's successfully gapped in prelim test: 621
Number of HSP's that attempted gapping in prelim test: 9569
Number of HSP's gapped (non-prelim): 6002
length of query: 1303
length of database: 12,639,632
effective HSP length: 108
effective length of query: 1195
effective length of database: 5,762,624
effective search space: 6886335680
effective search space used: 6886335680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.8 bits)
S2: 65 (29.6 bits)
Medicago: description of AC135504.9


