

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135504.16 - phase: 0 /pseudo
(343 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
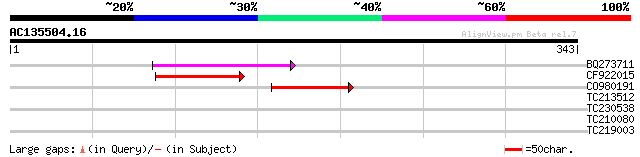
BQ273711 89 4e-18
CF922015 84 1e-16
CO980191 47 1e-05
TC213512 29 2.7
TC230538 29 3.5
TC210080 weakly similar to UP|Q8VZH8 (Q8VZH8) AT4g30550/F17I23_1... 28 4.6
TC219003 homologue to UP|Q43455 (Q43455) Heat shock transcriptio... 28 4.6
>BQ273711
Length = 409
Score = 88.6 bits (218), Expect = 4e-18
Identities = 39/87 (44%), Positives = 50/87 (56%)
Frame = -2
Query: 87 RMLETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEADDWW 146
R L F +NHPP F G YDP+G + WL E E+IF M C E KV + T ML EA++WW
Sbjct: 261 RGLMAFRKNHPPKFSGDYDPEGARLWLAETEKIFEAMGCLEEHKVLYATFMLQGEAENWW 82
Query: 147 VSLLPVLEQDGAVVTWAVFRREFLNRY 173
+ P G V+ W F+ +FL Y
Sbjct: 81 KFVKPSFVAPGGVIPWNAFKDKFLENY 1
>CF922015
Length = 172
Score = 83.6 bits (205), Expect = 1e-16
Identities = 35/54 (64%), Positives = 45/54 (82%)
Frame = -3
Query: 89 LETFLRNHPPTFKGRYDPDGTQKWLKEVERIFRVMQCSEVQKVRFGTHMLAEEA 142
L+ F RN+PPTFKG YDP+G + WL+E+E+IFRVM+C + QKV F THMLA+EA
Sbjct: 164 LDRFQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADEA 3
>CO980191
Length = 802
Score = 47.0 bits (110), Expect = 1e-05
Identities = 23/50 (46%), Positives = 30/50 (60%)
Frame = +1
Query: 159 VVTWAVFRREFLNRYFPEDVRGKNEIEFLELKHGDMSVTKYAAKFVELAK 208
V+T VF+ FL ++ ED+ + EFLELKHG+M V Y F EL K
Sbjct: 361 VIT*EVFKEIFLEKHLHEDI*NHKKTEFLELKHGNMIVANYLVNFDELPK 510
>TC213512
Length = 560
Score = 29.3 bits (64), Expect = 2.7
Identities = 11/27 (40%), Positives = 19/27 (69%)
Frame = +3
Query: 195 SVTKYAAKFVELAKFYPYYTAEIVEFS 221
S T+Y+ +F+EL ++YP +T + FS
Sbjct: 18 SQTRYSQRFIELIRYYPNFTNPNLAFS 98
>TC230538
Length = 1261
Score = 28.9 bits (63), Expect = 3.5
Identities = 24/73 (32%), Positives = 40/73 (53%), Gaps = 3/73 (4%)
Frame = +2
Query: 101 KGRYDPDGTQKWLKE-VERIFRVMQCSEVQKVRFGTHMLA--EEADDWWVSLLPVLEQDG 157
+GRYD DG++K L V ++F +M + V R G ++A E + ++ L + EQ
Sbjct: 17 EGRYDSDGSEKSLDSFVIQLFELM-LTIVGNPRLGKVVVANIRELVYYTIAFLQMTEQQ- 190
Query: 158 AVVTWAVFRREFL 170
V TW+V +F+
Sbjct: 191 -VHTWSVDANQFI 226
>TC210080 weakly similar to UP|Q8VZH8 (Q8VZH8) AT4g30550/F17I23_110, partial
(45%)
Length = 693
Score = 28.5 bits (62), Expect = 4.6
Identities = 17/55 (30%), Positives = 27/55 (48%), Gaps = 2/55 (3%)
Frame = -1
Query: 80 AAGNDGVRMLETFLRNHPPTFKGRYDPDGTQK--WLKEVERIFRVMQCSEVQKVR 132
A G DG ++ R+HP + + P+ W K++ RIF M+ S K+R
Sbjct: 516 ANGKDGDSVMVLTPRSHPVGERVTFPPNARPSIWWPKQIPRIFLFMESSLRSKLR 352
>TC219003 homologue to UP|Q43455 (Q43455) Heat shock transcription factor 29
(Fragment), partial (76%)
Length = 1188
Score = 28.5 bits (62), Expect = 4.6
Identities = 16/45 (35%), Positives = 27/45 (59%), Gaps = 2/45 (4%)
Frame = -3
Query: 130 KVRFGTHMLAEEADDWWVSLLPV--LEQDGAVVTWAVFRREFLNR 172
+ +F +L + +W+ LLPV L+ + AV++ AV R E L+R
Sbjct: 301 ETQFSPPVLPQPPSPYWL*LLPVPRLQAEPAVISGAVCREEELSR 167
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.355 0.158 0.577
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,640,731
Number of Sequences: 63676
Number of extensions: 206894
Number of successful extensions: 4511
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 3611
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4262
length of query: 343
length of database: 12,639,632
effective HSP length: 98
effective length of query: 245
effective length of database: 6,399,384
effective search space: 1567849080
effective search space used: 1567849080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC135504.16


