

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135465.2 + phase: 0
(194 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
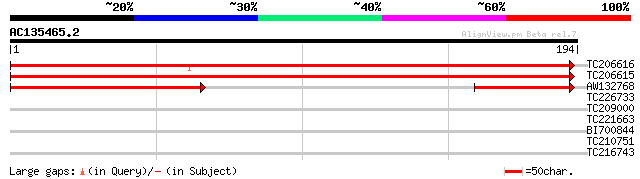
TC206616 homologue to UP|Q9ARI4 (Q9ARI4) PII protein, partial (68%) 292 8e-80
TC206615 similar to UP|Q9ARI4 (Q9ARI4) PII protein, partial (84%) 290 2e-79
AW132768 similar to GP|13277515|gb| PII protein {Medicago sativa... 72 1e-13
TC226733 similar to UP|O22165 (O22165) 60S ribosomal protein L30... 29 1.2
TC209000 similar to UP|RU17_ARATH (Q42404) U1 small nuclear ribo... 28 2.0
TC221663 28 3.5
BI700844 similar to SP|Q42404|RU17_ U1 small nuclear ribonucleop... 27 4.5
TC210751 27 5.9
TC216743 similar to UP|Q9LJE0 (Q9LJE0) Gb|AAC16072.1 (AT3g13510/... 27 5.9
>TC206616 homologue to UP|Q9ARI4 (Q9ARI4) PII protein, partial (68%)
Length = 1301
Score = 292 bits (747), Expect = 8e-80
Identities = 149/194 (76%), Positives = 168/194 (85%), Gaps = 1/194 (0%)
Frame = +3
Query: 1 MALIAKPNVFNGLNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN 60
MA IA +VF ++F + + + PF+ S+IRK GDS RNV L+ N +ILP+IRAQN
Sbjct: 45 MAAIAGAHVFGVVSFQLKQAEMPFACSSLIRKHIGDSPQRNVALRRRANGTILPQIRAQN 224
Query: 61 -LPDYVPESKFYKVEAILRPWRIPQVSSGLLKMGIRGVTVSDVKGFGAQGGSKERQGGSE 119
LPDYVP+S+FYKVEAILRPWR+PQVSS +LKMGIRGVTVSDVKGFG+QGGSKERQGGSE
Sbjct: 225 VLPDYVPKSEFYKVEAILRPWRVPQVSSAMLKMGIRGVTVSDVKGFGSQGGSKERQGGSE 404
Query: 120 FSEDNFVAKVKMEIVVRKDQVEAVINKIMETARTGEIGDGKIFLIPVSDVIRIRTGERGE 179
FSEDNFVAKVKMEIVVRKDQVEAVI+KI+E ARTGEIGDGKIFLIPVSDVIRIRTGERGE
Sbjct: 405 FSEDNFVAKVKMEIVVRKDQVEAVIDKIIEEARTGEIGDGKIFLIPVSDVIRIRTGERGE 584
Query: 180 QAERMAGGLTDALS 193
QAERM GG +D LS
Sbjct: 585 QAERMTGGRSDMLS 626
>TC206615 similar to UP|Q9ARI4 (Q9ARI4) PII protein, partial (84%)
Length = 837
Score = 290 bits (743), Expect = 2e-79
Identities = 145/193 (75%), Positives = 166/193 (85%)
Frame = +1
Query: 1 MALIAKPNVFNGLNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN 60
M IA +VF ++F + + + PF+ +IRKR GDS RNV L+ N +ILP+IRAQN
Sbjct: 28 MTAIAGTHVFGVVSFQLKQAEMPFACSCLIRKRIGDSPQRNVALRRRVNGTILPQIRAQN 207
Query: 61 LPDYVPESKFYKVEAILRPWRIPQVSSGLLKMGIRGVTVSDVKGFGAQGGSKERQGGSEF 120
LPDYVP+S+FYKVEAILRPWR+PQVS+ LLKMGIRGVTVSDV+GFGAQGGSKERQGGSEF
Sbjct: 208 LPDYVPKSEFYKVEAILRPWRVPQVSAALLKMGIRGVTVSDVRGFGAQGGSKERQGGSEF 387
Query: 121 SEDNFVAKVKMEIVVRKDQVEAVINKIMETARTGEIGDGKIFLIPVSDVIRIRTGERGEQ 180
SEDNFVAKVKME+VVRKDQVEAVI+KI+E ARTGEIGDGKIFLIP+SDVIRIRTGERGEQ
Sbjct: 388 SEDNFVAKVKMEVVVRKDQVEAVIDKIIEEARTGEIGDGKIFLIPISDVIRIRTGERGEQ 567
Query: 181 AERMAGGLTDALS 193
A RM GG +D LS
Sbjct: 568 AARMTGGRSDMLS 606
>AW132768 similar to GP|13277515|gb| PII protein {Medicago sativa}, partial
(36%)
Length = 426
Score = 72.4 bits (176), Expect = 1e-13
Identities = 34/67 (50%), Positives = 46/67 (67%)
Frame = +1
Query: 1 MALIAKPNVFNGLNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN 60
M IA +VF ++F + + + PF+ +IRKR GDS RNV L+ N +ILP+IRAQN
Sbjct: 22 MTAIAGTHVFGVVSFQLKQAEMPFACSCLIRKRIGDSPQRNVALRRRVNGTILPQIRAQN 201
Query: 61 LPDYVPE 67
LPDYVP+
Sbjct: 202LPDYVPK 222
Score = 58.2 bits (139), Expect = 2e-09
Identities = 28/34 (82%), Positives = 30/34 (87%)
Frame = +3
Query: 160 KIFLIPVSDVIRIRTGERGEQAERMAGGLTDALS 193
KIFLIP+SDVIRIRTGERGEQA RM GG +D LS
Sbjct: 219 KIFLIPISDVIRIRTGERGEQAARMTGGRSDMLS 320
>TC226733 similar to UP|O22165 (O22165) 60S ribosomal protein L30
(At2g44860/T13E15.13), partial (86%)
Length = 822
Score = 29.3 bits (64), Expect = 1.2
Identities = 19/61 (31%), Positives = 28/61 (45%)
Frame = +2
Query: 103 KGFGAQGGSKERQGGSEFSEDNFVAKVKMEIVVRKDQVEAVINKIMETARTGEIGDGKIF 162
KG A+GGS+ G +F + +F A + KDQ + TA GE G++
Sbjct: 452 KGKAAEGGSEGVGAGHQFGQSSFCASTGSISHITKDQSQG-----FPTAIRGESCHGRVI 616
Query: 163 L 163
L
Sbjct: 617 L 619
>TC209000 similar to UP|RU17_ARATH (Q42404) U1 small nuclear
ribonucleoprotein 70 kDa (U1 snRNP 70 kDa) (snRNP70)
(U1-70K), partial (42%)
Length = 689
Score = 28.5 bits (62), Expect = 2.0
Identities = 14/48 (29%), Positives = 25/48 (51%)
Frame = +3
Query: 13 LNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN 60
LNF + + F+++R GD++ N N NA++ + +AQN
Sbjct: 72 LNFFL----YALCDFTLLRHPMGDNNSNNDAFMRNQNAAVQARTKAQN 203
>TC221663
Length = 781
Score = 27.7 bits (60), Expect = 3.5
Identities = 11/32 (34%), Positives = 20/32 (62%)
Frame = +1
Query: 53 LPKIRAQNLPDYVPESKFYKVEAILRPWRIPQ 84
L K+RA LP + +S+ ++ L+PW+I +
Sbjct: 196 LKKVRATGLPTILEDSESPRIMEDLKPWKIDE 291
>BI700844 similar to SP|Q42404|RU17_ U1 small nuclear ribonucleoprotein 70
kDa (U1 snRNP 70 kDa) (snRNP70) (U1-70K). [Mouse-ear
cress], partial (14%)
Length = 421
Score = 27.3 bits (59), Expect = 4.5
Identities = 14/48 (29%), Positives = 24/48 (49%)
Frame = +2
Query: 13 LNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN 60
LNF + + F+++R GD++ N N NA++ +AQN
Sbjct: 77 LNFFL----YALCDFTLLRHPMGDNNSNNDAFMRNQNAAVQASSKAQN 208
>TC210751
Length = 713
Score = 26.9 bits (58), Expect = 5.9
Identities = 14/48 (29%), Positives = 27/48 (56%)
Frame = +3
Query: 13 LNFHINETQFPFSSFSVIRKRFGDSSHRNVVLKSNGNASILPKIRAQN 60
L+ + + +FPF SFS+ F R V+ +++ + ++ PK R+ N
Sbjct: 45 LHSKLKQARFPFPSFSLQNPTF-----RPVICRASESPTVPPKKRSSN 173
>TC216743 similar to UP|Q9LJE0 (Q9LJE0) Gb|AAC16072.1 (AT3g13510/MRP15_15),
partial (90%)
Length = 1869
Score = 26.9 bits (58), Expect = 5.9
Identities = 12/49 (24%), Positives = 22/49 (44%)
Frame = +3
Query: 32 KRFGDSSHRNVVLKSNGNASILPKIRAQNLPDYVPESKFYKVEAILRPW 80
KR+G HR + + ++ + Q+ YV K+Y +A + W
Sbjct: 612 KRYGRKKHRAIPKPRSAEPDLINQSGHQHAIAYVEGDKYYGAKATINVW 758
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.137 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,234,032
Number of Sequences: 63676
Number of extensions: 68091
Number of successful extensions: 425
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 423
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 424
length of query: 194
length of database: 12,639,632
effective HSP length: 92
effective length of query: 102
effective length of database: 6,781,440
effective search space: 691706880
effective search space used: 691706880
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC135465.2


