

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
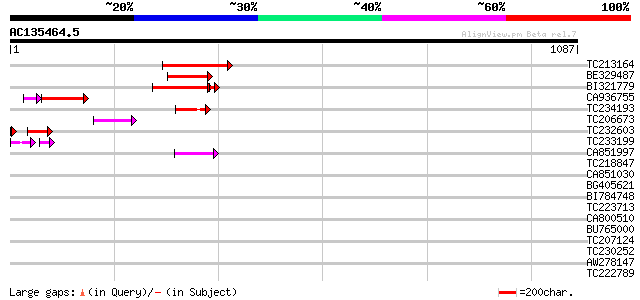
Query= AC135464.5 - phase: 0 /pseudo
(1087 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC213164 137 3e-32
BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase... 124 2e-28
BI321779 105 7e-25
CA936755 94 1e-21
TC234193 60 4e-09
TC206673 58 2e-08
TC232603 weakly similar to UP|Q39482 (Q39482) Legumin (Fragment)... 52 7e-07
TC233199 33 8e-05
CA851997 44 5e-04
TC218847 42 0.001
CA851030 37 0.047
BG405621 36 0.11
BI784748 35 0.18
TC223713 34 0.31
CA800510 32 1.5
BU765000 30 4.4
TC207124 similar to UP|Q7X9B4 (Q7X9B4) Allene oxide synthase pre... 30 4.4
TC230252 similar to UP|Q40793 (Q40793) Tyrosine-rich hydroxyprol... 30 5.8
AW278147 30 7.6
TC222789 29 9.9
>TC213164
Length = 446
Score = 137 bits (345), Expect = 3e-32
Identities = 63/135 (46%), Positives = 94/135 (68%)
Frame = +3
Query: 293 LTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRG 352
L F EV++ +WD + KSPGPDG+NF FIKEFWD LK D++RF+ +F+ NG +
Sbjct: 18 LVARFEKKEVRATMWDRGNAKSPGPDGLNFKFIKEFWDILKTDLLRFLDEFYANGVFLKR 197
Query: 353 INTTFIALISKIDSPQRINDFRPISLVGSLYKILSKVLANRLKLVLGKIISDSQTAFVKD 412
N F+ALI K+ P +N+++PISL+G +YKI++K+L+ R K VL II + QT F++
Sbjct: 198 SNAFFLALIPKVHDP*SLNEYKPISLIGCIYKIVAKLLSGRHKKVLPTIIDERQTVFMEG 377
Query: 413 RQILDGILIANEVVD 427
R L ++IANE+++
Sbjct: 378 RHTLHNVVIANEIME 422
>BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (15%)
Length = 376
Score = 124 bits (311), Expect = 2e-28
Identities = 53/88 (60%), Positives = 70/88 (79%)
Frame = -2
Query: 302 VKSAVWDCDSYKSPGPDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALI 361
+KSAVW C S KSPGPDG+NF FIK FW+ LK D++RF+++FH NG +G N +FIALI
Sbjct: 375 IKSAVWQCGSVKSPGPDGLNFKFIKHFWERLKPDIIRFLNEFHANGIFPKGGNASFIALI 196
Query: 362 SKIDSPQRINDFRPISLVGSLYKILSKV 389
K+ PQ +NDFRPISL+G +YKI++K+
Sbjct: 195 PKVKHPQALNDFRPISLIGCVYKIVAKI 112
>BI321779
Length = 421
Score = 105 bits (263), Expect(2) = 7e-25
Identities = 52/115 (45%), Positives = 74/115 (64%)
Frame = +2
Query: 274 RVGIDNLPFKCLSHTDGSGLTKPFSADEVKSAVWDCDSYKSPGPDGVNFGFIKEFWDELK 333
R ++ + F S+ D + L + F +E+K + DC+S KSPGPDG + IK FW+ +K
Sbjct: 26 RPKLNGVEFLSFSNEDNAMLIQDFEVEEIKKVI*DCESSKSPGPDGYSLLLIKIFWEVIK 205
Query: 334 EDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPISLVGSLYKILSK 388
E+V+ F +FH+ L RG N +FIALI+K D+P I + PISLVG LYKI+ K
Sbjct: 206 EEVINFFKEFHKFDFLPRGTNRSFIALIAKCDNPLDIGQYCPISLVGCLYKIV*K 370
Score = 27.7 bits (60), Expect(2) = 7e-25
Identities = 10/19 (52%), Positives = 16/19 (83%)
Frame = +3
Query: 384 KILSKVLANRLKLVLGKII 402
K+ K++ANR+ +VLGK+I
Sbjct: 357 KLYKKIIANRMSIVLGKLI 413
>CA936755
Length = 424
Score = 94.0 bits (232), Expect(2) = 1e-21
Identities = 37/90 (41%), Positives = 58/90 (64%)
Frame = -2
Query: 61 FLLSEEWCLQWPNSFQVALLRGLSDHCPLQLSVDEENWGPRPSRMLKCWQDLPGYNQFVK 120
FL+SE+W L WP+S Q L R SDHCP+ + + +WGP+P R++ W GY + V+
Sbjct: 294 FLVSEQWLLSWPDSSQYTLPRDFSDHCPIIMQTKKVDWGPKPFRVVDWWLHQKGYQRLVR 115
Query: 121 DKLKSFQIEGWGGFVLREKLKFIKTALKEW 150
+ + Q GWGG +L+ KL+ +K ++K+W
Sbjct: 114 ETWSAEQQPGWGGILLKNKLRMLKLSIKQW 25
Score = 28.9 bits (63), Expect(2) = 1e-21
Identities = 11/36 (30%), Positives = 20/36 (55%)
Frame = -1
Query: 26 FSDFIVDNSLIDLSLCGRRFTWFKGDGNAMSRIDRF 61
F+ +I + L ++ G +TW + + SR+DRF
Sbjct: 400 FNTWISEMELQEIKCVGSTYTWIRPNARVKSRLDRF 293
>TC234193
Length = 697
Score = 60.5 bits (145), Expect = 4e-09
Identities = 31/67 (46%), Positives = 44/67 (65%)
Frame = -3
Query: 318 DGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPIS 377
+G NF FIK+ + +KEDV+R I +FH +G L+RG N +F+ P + DFRP++
Sbjct: 194 NGYNFNFIKKC*EVMKEDVVRAIQEFHSHGCLSRGTNASFLT-----HPP*GLGDFRPVT 30
Query: 378 LVGSLYK 384
LVGSL K
Sbjct: 29 LVGSLCK 9
>TC206673
Length = 742
Score = 58.2 bits (139), Expect = 2e-08
Identities = 31/82 (37%), Positives = 46/82 (55%)
Frame = +3
Query: 162 IDSLKKRQADFDDKGDVGDLSADEILELRGITHDLHSLSRVNSSILWQQSRLLWLKDGDA 221
I ++KK+ + +D LS DEI + + L S S+L Q+SR WLK+GD
Sbjct: 21 IHNIKKKLNEVEDLASNRSLSVDEIKAKKELQQQLWEASTAYESLLRQKSRDKWLKEGDN 200
Query: 222 NSKYFHSVLSGRRRRNSIISLL 243
NS YFH+V++ RR N + +L
Sbjct: 201 NSAYFHTVINFRRHYNGLQGIL 266
>TC232603 weakly similar to UP|Q39482 (Q39482) Legumin (Fragment), partial
(4%)
Length = 1123
Score = 52.4 bits (124), Expect(2) = 7e-07
Identities = 22/47 (46%), Positives = 32/47 (67%)
Frame = +2
Query: 35 LIDLSLCGRRFTWFKGDGNAMSRIDRFLLSEEWCLQWPNSFQVALLR 81
+++L+L GR++ W+K D MSR+DRFL+ EE WPN + LLR
Sbjct: 524 VVNLALVGRKYNWYKDDKTTMSRLDRFLVYEE*LTTWPNMTLILLLR 664
Score = 20.0 bits (40), Expect(2) = 7e-07
Identities = 9/12 (75%), Positives = 9/12 (75%)
Frame = +1
Query: 1 DFNAVRYAEERK 12
DFNAV EERK
Sbjct: 445 DFNAVCSEEERK 480
>TC233199
Length = 1105
Score = 33.5 bits (75), Expect(2) = 8e-05
Identities = 20/51 (39%), Positives = 28/51 (54%), Gaps = 2/51 (3%)
Frame = +2
Query: 1 DFNAVRYAEER--KSSRGISGVIDYHHFSDFIVDNSLIDLSLCGRRFTWFK 49
DFN VR ER S R G I F+D+IVD + D+ G ++TW++
Sbjct: 305 DFNCVRRQSERVGASQREQGGGI-MTEFNDWIVDLEVEDMPSVGHKYTWYR 454
Score = 32.0 bits (71), Expect(2) = 8e-05
Identities = 14/29 (48%), Positives = 17/29 (58%)
Frame = +1
Query: 58 IDRFLLSEEWCLQWPNSFQVALLRGLSDH 86
+DR L+S EW +W S Q L R SDH
Sbjct: 466 LDRVLVSVEWFSKWSGSIQFILDRNYSDH 552
>CA851997
Length = 636
Score = 43.5 bits (101), Expect = 5e-04
Identities = 33/86 (38%), Positives = 40/86 (46%), Gaps = 2/86 (2%)
Frame = +1
Query: 317 PDGVNFGFIKEFWDELKEDVMRFISDFHRNGKLTRGINTTFIALISKIDSPQRINDFRPI 376
PDG N GF FW EDV + + G +N T IA+I +I R
Sbjct: 316 PDGFNLGFYHRFWGMCGEDVFQACCMWLAEGAFPSSVNDTTIAIILEI**S*RYERS*TN 495
Query: 377 SLVGSLYKI--LSKVLANRLKLVLGK 400
L +KI LS+VLA RLK VL K
Sbjct: 496 LLCNGGFKILFLSEVLAKRLKNVLDK 573
>TC218847
Length = 521
Score = 42.4 bits (98), Expect = 0.001
Identities = 22/86 (25%), Positives = 47/86 (54%)
Frame = -3
Query: 200 SRVNSSILWQQSRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLADSNLVEGVQPIRNAV 259
++ + S+L Q++R W+++ D NS YFH +++ R N++ ++ + VE ++ AV
Sbjct: 273 AQAHESLLRQKARTKWIRERDCNSSYFHKLINYIHRTNAVNGVMIGGSWVEEPNRVK-AV 97
Query: 260 LSHFRDHFAARHTSRVGIDNLPFKCL 285
+ F+ F+ R I+ F+ +
Sbjct: 96 RNFFQIRFSESDYDRP*INGANFRSI 19
>CA851030
Length = 358
Score = 37.0 bits (84), Expect = 0.047
Identities = 19/64 (29%), Positives = 30/64 (46%)
Frame = +1
Query: 186 ILELRGITHDLHSLSRVNSSILWQQSRLLWLKDGDANSKYFHSVLSGRRRRNSIISLLAD 245
I + + L SL + Q+SR +W K+ D N+++FH R+ NSI + D
Sbjct: 121 IASTKVVEESLDSLLKQEEEYWQQRSRAIWFKERDKNTEFFHRKTLQRKASNSITKIQDD 300
Query: 246 SNLV 249
V
Sbjct: 301 EGNV 312
>BG405621
Length = 402
Score = 35.8 bits (81), Expect = 0.11
Identities = 15/22 (68%), Positives = 18/22 (81%), Gaps = 1/22 (4%)
Frame = +2
Query: 208 WQQ-SRLLWLKDGDANSKYFHS 228
WQQ S++ WLK GD NSK+FHS
Sbjct: 335 WQQRSKIFWLKYGDLNSKFFHS 400
>BI784748
Length = 259
Score = 35.0 bits (79), Expect = 0.18
Identities = 13/29 (44%), Positives = 24/29 (81%)
Frame = +2
Query: 406 QTAFVKDRQILDGILIANEVVDEARKNKR 434
Q+AF++ R +L ++IANEV+DEA++ ++
Sbjct: 14 QSAFIQGRHMLHSMMIANEVIDEAKRGQK 100
>TC223713
Length = 420
Score = 34.3 bits (77), Expect = 0.31
Identities = 22/61 (36%), Positives = 30/61 (49%)
Frame = +2
Query: 923 RCRLRFPFLLGDYCVIGCRLRIIFLGVA*STVRFVCVSQGAVWKRLHLIFLFIAPLLVLC 982
RC+ +LLG + IGC L+ I+ G * + + V A WKR I+ IA L
Sbjct: 122 RCQSNLQYLLGGFSEIGCLLKQIYTGDK*RS-QIGVVRSAATWKRKQGIYSSIAAKLSQS 298
Query: 983 G 983
G
Sbjct: 299 G 301
>CA800510
Length = 409
Score = 32.0 bits (71), Expect = 1.5
Identities = 13/23 (56%), Positives = 16/23 (69%)
Frame = -1
Query: 220 DANSKYFHSVLSGRRRRNSIISL 242
D NSKYFH + R+RRN I+ L
Sbjct: 97 DYNSKYFHICIKSRQRRNQILEL 29
>BU765000
Length = 425
Score = 30.4 bits (67), Expect = 4.4
Identities = 20/58 (34%), Positives = 28/58 (47%)
Frame = +3
Query: 919 YGISRCRLRFPFLLGDYCVIGCRLRIIFLGVA*STVRFVCVSQGAVWKRLHLIFLFIA 976
YG R L++ L G Y IG + I G S + +CV W+R+H+ FIA
Sbjct: 60 YGSLRSHLKWLCLHGGYSKIGYQPEIT*EGSKLSYMN-ICVLSAEQWRRVHVTCFFIA 230
>TC207124 similar to UP|Q7X9B4 (Q7X9B4) Allene oxide synthase precursor ,
partial (30%)
Length = 486
Score = 30.4 bits (67), Expect = 4.4
Identities = 17/43 (39%), Positives = 25/43 (57%), Gaps = 1/43 (2%)
Frame = +3
Query: 153 GHASNIPGKIDSLKK-RQADFDDKGDVGDLSADEILELRGITH 194
G A PG DSL++ R+ D G G+ +ADE+ +RG+ H
Sbjct: 330 GEAPRAPGGGDSLRRERRRRGDHHGGDGEHAADEVGGVRGVPH 458
>TC230252 similar to UP|Q40793 (Q40793) Tyrosine-rich hydroxyproline-rich
glycoprotein (Fragment), partial (42%)
Length = 506
Score = 30.0 bits (66), Expect = 5.8
Identities = 20/45 (44%), Positives = 26/45 (57%), Gaps = 6/45 (13%)
Frame = -2
Query: 590 VSILRIRGYMRRLLF*IVRWVRFRLCI------WAYRLVVILGG* 628
V ++ RG+ RR F IVRW R RL I W +RLV++ G*
Sbjct: 136 VIVVGSRGWRRRR-FVIVRWWRRRLVIVRSWGWWRWRLVIVRSG* 5
>AW278147
Length = 427
Score = 29.6 bits (65), Expect = 7.6
Identities = 20/67 (29%), Positives = 32/67 (46%)
Frame = +1
Query: 138 EKLKFIKTALKEWHIGHASNIPGKIDSLKKRQADFDDKGDVGDLSADEILELRGITHDLH 197
+K+K + A H HA+ I + SL KRQ F+D+ + +A + + + H
Sbjct: 43 QKVKLLLNATVLSHAWHAATILRGV-SLSKRQISFNDRPGLMMCTASDETDSEDLVSPAH 219
Query: 198 SLSRVNS 204
SL R S
Sbjct: 220 SLQRTIS 240
>TC222789
Length = 954
Score = 29.3 bits (64), Expect = 9.9
Identities = 19/54 (35%), Positives = 26/54 (47%)
Frame = +3
Query: 923 RCRLRFPFLLGDYCVIGCRLRIIFLGVA*STVRFVCVSQGAVWKRLHLIFLFIA 976
RC+ +LLG Y IGC L+ I +G + V +WKR+ FIA
Sbjct: 375 RCQSNMKYLLGGYLGIGCLLK*ICIGDKYRS-WIGAVHFAEMWKRMQGTCFFIA 533
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.344 0.155 0.538
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 57,465,891
Number of Sequences: 63676
Number of extensions: 967598
Number of successful extensions: 9168
Number of sequences better than 10.0: 43
Number of HSP's better than 10.0 without gapping: 6682
Number of HSP's successfully gapped in prelim test: 285
Number of HSP's that attempted gapping in prelim test: 2277
Number of HSP's gapped (non-prelim): 7289
length of query: 1087
length of database: 12,639,632
effective HSP length: 107
effective length of query: 980
effective length of database: 5,826,300
effective search space: 5709774000
effective search space used: 5709774000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.5 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135464.5


