

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135463.4 - phase: 2 /pseudo
(1134 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
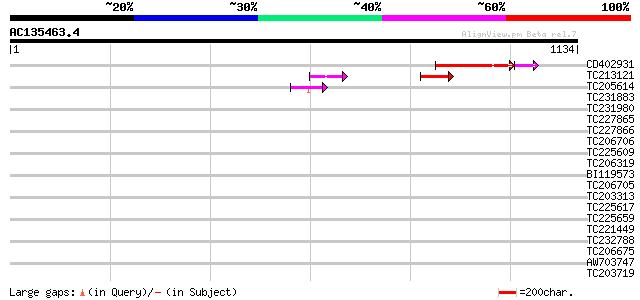
Score E
Sequences producing significant alignments: (bits) Value
CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (ja... 143 3e-38
TC213121 59 9e-09
TC205614 55 2e-07
TC231883 weakly similar to UP|Q7QBQ3 (Q7QBQ3) AgCP4056 (Fragment... 39 0.017
TC231980 similar to UP|Q8S562 (Q8S562) KAP-2, partial (19%) 37 0.038
TC227865 homologue to GB|AAN31116.1|23506209|AY149962 At3g06130/... 35 0.14
TC227866 similar to UP|P2B1_DROME (P48456) Serine/threonine prot... 35 0.19
TC206706 similar to UP|Q9NJZ3 (Q9NJZ3) Fruitless type E, partial... 33 0.93
TC225609 UP|Q84ZV3 (Q84ZV3) R 4 protein, complete 32 1.2
TC206319 similar to UP|Q9Y0C9 (Q9Y0C9) Ras interacting protein R... 32 1.6
BI119573 32 2.1
TC206705 similar to UP|Q8MMZ5 (Q8MMZ5) CGMP-dependent protein ki... 32 2.1
TC203313 PIR|S01425|UQMUM polyubiquitin 5 - Arabidopsis thaliana... 31 2.7
TC225617 UP|Q84ZV8 (Q84ZV8) R 3 protein, complete 31 3.6
TC225659 similar to UP|Q9FUW4 (Q9FUW4) Cold acclimation protein ... 31 3.6
TC221449 UP|Q9FXZ1 (Q9FXZ1) Resistance protein MG23 (Fragment), ... 30 4.6
TC232788 similar to UP|Q9ZVU9 (Q9ZVU9) T5A14.6 protein, partial ... 30 4.6
TC206675 30 4.6
AW703747 30 4.6
TC203719 homologue to UP|Q43437 (Q43437) Photosystem II type I c... 30 6.1
>CD402931 similar to GP|9081793|dbj P0489A01.23 {Oryza sativa (japonica
cultivar-group)}, partial (5%)
Length = 617
Score = 143 bits (361), Expect(2) = 3e-38
Identities = 80/158 (50%), Positives = 106/158 (66%)
Frame = +2
Query: 852 GIGKTFLWNALSYHFRSEGKIVLNVASSSIASLLLPRGRTAHSQFSIPLMLLEESGCTIE 911
G KTFLW LS RS+G ++ + +ASLLLPRG TAHS FSIPL++ ++S C I
Sbjct: 11 GTSKTFLWKTLSAGMRSKG-LMSSSCLKRLASLLLPRG*TAHSTFSIPLVIKDDSTCNIN 187
Query: 912 KEGNKAQLLTMASLIIWDEAPMISRLAFEAFDRTMRDIMSNVVDGASNLPFGGKTVVLGG 971
+ +++LL LIIWDE P++++ FEA DRT++DIM++ + PFGGK VVLGG
Sbjct: 188 QGSARSKLLLHTKLIIWDETPVMNKFCFEALDRTLQDIMASQNKDNATKPFGGK-VVLGG 364
Query: 972 DFRQILPVVPKRGRADIVDACINSSMLWRRCRVLKLTK 1009
DFRQILPV+ K R DIV + IN+S +VLKL K
Sbjct: 365 DFRQILPVIRKGSRQDIVGSAINASK-----KVLKLKK 463
Score = 35.0 bits (79), Expect(2) = 3e-38
Identities = 18/48 (37%), Positives = 28/48 (57%)
Frame = +1
Query: 1009 KNIRLQFPTNGQDDASLRLFARWILDVGDGKLGIDNDGEAVIEIPDDI 1056
KN+RL N +D +R F WIL + DG +++GE I+IP ++
Sbjct: 460 KNMRLGTTNNNEDRDDIREFVDWILMIRDGNRDEEDEGE--IDIPKNL 597
>TC213121
Length = 713
Score = 59.3 bits (142), Expect = 9e-09
Identities = 28/66 (42%), Positives = 41/66 (61%)
Frame = +2
Query: 822 LNSDQLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFLWNALSYHFRSEGKIVLNVASSSI 881
L + S + KI++++ + ++F+ GYGG GKTF+W LS S IVL +ASS I
Sbjct: 410 LTDEHASIFNKIMHMIATQSGGVYFLYGYGGTGKTFVWKTLSSTIHSNSGIVLTMASSGI 589
Query: 882 ASLLLP 887
S+LLP
Sbjct: 590 LSMLLP 607
Score = 47.4 bits (111), Expect = 4e-05
Identities = 27/77 (35%), Positives = 46/77 (59%)
Frame = +3
Query: 599 CRYLSPSESIWRIFGFDVHCRYPAVKRLTFHLHGEQRIMFRDSSRPDNVLTRNRHKRTMF 658
C Y+SP E+ +I F +H R P V+ L FHL E + ++S + ++L+++ K ++F
Sbjct: 30 CIYVSPPETC*KISAFPMHGRAPTVECLYFHL--ENQ*P*KESQQIGSMLSKSTIKESIF 203
Query: 659 LAWMDANRNYSQSWNLT 675
A MD+N+ Y +LT
Sbjct: 204 TACMDSNKMYHHGRDLT 254
>TC205614
Length = 742
Score = 54.7 bits (130), Expect = 2e-07
Identities = 31/81 (38%), Positives = 45/81 (55%), Gaps = 7/81 (8%)
Frame = +2
Query: 561 SNSIKYLFKYITKGVDRVTAT-LETSEEPSVDEIQ*------YYDCRYLSPSESIWRIFG 613
SN ++F YI+ + T L+ + S++ Q Y DC+Y+SP E+ WRIF
Sbjct: 497 SNLFSFIFHYISMVENHN*RTSLKLKDLASIEASQVSFPHMYYLDCQYISPCEACWRIFS 676
Query: 614 FDVHCRYPAVKRLTFHLHGEQ 634
F ++ R P V+RL FHL EQ
Sbjct: 677 FQIYGRIPIVERLYFHLPNEQ 739
>TC231883 weakly similar to UP|Q7QBQ3 (Q7QBQ3) AgCP4056 (Fragment), partial
(17%)
Length = 1263
Score = 38.5 bits (88), Expect = 0.017
Identities = 16/31 (51%), Positives = 24/31 (76%)
Frame = +1
Query: 516 LHLRRDTGVTVLKNGIQLDNRSVVPYNPKLI 546
++ RR+ G T+LKN + +DNR +VPYN KL+
Sbjct: 1168 VYRRRNDGRTILKNSVIVDNRYLVPYNAKLL 1260
>TC231980 similar to UP|Q8S562 (Q8S562) KAP-2, partial (19%)
Length = 590
Score = 37.4 bits (85), Expect = 0.038
Identities = 15/36 (41%), Positives = 24/36 (66%)
Frame = -3
Query: 783 LLLTSCMSDPLNVWNQTWHILSEGILYERRRTLNSP 818
LLLT+ S VW+QTWH +++ I+Y +++ SP
Sbjct: 582 LLLTATXSKSEQVWDQTWHWMADDIVYNYKKSSTSP 475
>TC227865 homologue to GB|AAN31116.1|23506209|AY149962 At3g06130/F28L1_7
{Arabidopsis thaliana;} , partial (6%)
Length = 1290
Score = 35.4 bits (80), Expect = 0.14
Identities = 13/33 (39%), Positives = 19/33 (57%), Gaps = 1/33 (3%)
Frame = -1
Query: 411 MMMYMMCCLCCYKMMFMMCCLCCYKMM-FMMCC 442
M + CC CC+ +M + CC CC M+ + CC
Sbjct: 747 MFACICCCCCCFIIMAISCCCCCCCML*GLGCC 649
Score = 30.8 bits (68), Expect = 3.6
Identities = 10/29 (34%), Positives = 16/29 (54%)
Frame = -1
Query: 414 YMMCCLCCYKMMFMMCCLCCYKMMFMMCC 442
Y+MC + + CC CC+ +M + CC
Sbjct: 774 YIMCGGYTFMFACICCCCCCFIIMAISCC 688
>TC227866 similar to UP|P2B1_DROME (P48456) Serine/threonine protein
phosphatase 2B catalytic subunit 1
(Calmodulin-dependent calcineurin A1 subunit) , partial
(3%)
Length = 1013
Score = 35.0 bits (79), Expect = 0.19
Identities = 14/44 (31%), Positives = 21/44 (46%), Gaps = 4/44 (9%)
Frame = -2
Query: 403 PHVVNEVLMMMYMMCCLCCYKMMFM----MCCLCCYKMMFMMCC 442
P+++ M M+ CC CC + M CC CC + + CC
Sbjct: 646 PYIMCGGYMFMFACCCCCCCWFIIMAISCCCCCCCCML*GLGCC 515
>TC206706 similar to UP|Q9NJZ3 (Q9NJZ3) Fruitless type E, partial (4%)
Length = 884
Score = 32.7 bits (73), Expect = 0.93
Identities = 9/25 (36%), Positives = 20/25 (80%), Gaps = 2/25 (8%)
Frame = -3
Query: 409 VLMMMYMMCCLCCY--KMMFMMCCL 431
+L++++++CC CC+ K + ++CCL
Sbjct: 513 LLLLLFVLCCCCCFFLKFVVLLCCL 439
Score = 32.3 bits (72), Expect = 1.2
Identities = 10/31 (32%), Positives = 21/31 (67%), Gaps = 4/31 (12%)
Frame = -3
Query: 420 CCYKMM--FMMCCLCCY--KMMFMMCCYKMM 446
CC+ ++ F++CC CC+ K + ++CC ++
Sbjct: 522 CCFLLLLLFVLCCCCCFFLKFVVLLCCLVLL 430
>TC225609 UP|Q84ZV3 (Q84ZV3) R 4 protein, complete
Length = 2688
Score = 32.3 bits (72), Expect = 1.2
Identities = 19/59 (32%), Positives = 33/59 (55%), Gaps = 4/59 (6%)
Frame = +1
Query: 826 QLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFL----WNALSYHFRSEGKIVLNVASSS 880
Q++ K+++V ++L H+ + G GG+GKT L +N ++ HF E + NV S
Sbjct: 580 QVTEVMKLLDVGSDDLVHIIGIHGMGGLGKTTLALEVYNLIALHF-DESCFLQNVREES 753
>TC206319 similar to UP|Q9Y0C9 (Q9Y0C9) Ras interacting protein RIPA, partial
(5%)
Length = 1428
Score = 32.0 bits (71), Expect = 1.6
Identities = 10/26 (38%), Positives = 12/26 (45%)
Frame = -3
Query: 417 CCLCCYKMMFMMCCLCCYKMMFMMCC 442
CC+CC+ CC CC CC
Sbjct: 766 CCICCWSCNCCWCCCCCCWGC*CCCC 689
Score = 30.8 bits (68), Expect = 3.6
Identities = 10/26 (38%), Positives = 12/26 (45%)
Frame = -3
Query: 417 CCLCCYKMMFMMCCLCCYKMMFMMCC 442
CC CC CC+CC+ CC
Sbjct: 790 CCCCCCNC----CCICCWSCNCCWCC 725
>BI119573
Length = 605
Score = 31.6 bits (70), Expect = 2.1
Identities = 15/41 (36%), Positives = 22/41 (53%), Gaps = 2/41 (4%)
Frame = +3
Query: 410 LMMMYMMCCLCCYKMMFMMCCLCCYKM--MFMMCCYKMMFM 448
L+ Y CC CC M + C CYK+ +F +C +MF+
Sbjct: 258 LLNQYYCCCNCCSCSM-LSCSNLCYKLIILFYVCVLVIMFV 377
>TC206705 similar to UP|Q8MMZ5 (Q8MMZ5) CGMP-dependent protein kinase,
partial (3%)
Length = 1324
Score = 31.6 bits (70), Expect = 2.1
Identities = 9/25 (36%), Positives = 15/25 (60%)
Frame = -1
Query: 409 VLMMMYMMCCLCCYKMMFMMCCLCC 433
VL+++ + CC CC ++ C CC
Sbjct: 493 VLLLLEVCCCCCCSSLLSCSSCCCC 419
Score = 29.6 bits (65), Expect = 7.9
Identities = 10/42 (23%), Positives = 21/42 (49%), Gaps = 3/42 (7%)
Frame = -1
Query: 407 NEVLMMMYMMCCLCCYKMMFMMCCLCCYKMMF---MMCCYKM 445
N+ L +++ + LC ++ +CC CC + CC ++
Sbjct: 538 NKNLFLLFFVVALCVVLLLLEVCCCCCCSSLLSCSSCCCCRL 413
>TC203313 PIR|S01425|UQMUM polyubiquitin 5 - Arabidopsis thaliana
{Arabidopsis thaliana;} , complete
Length = 1377
Score = 31.2 bits (69), Expect = 2.7
Identities = 27/92 (29%), Positives = 39/92 (42%), Gaps = 1/92 (1%)
Frame = -1
Query: 768 VVVGSGSSIRKVFTSLLLTSCMSDPLNVWNQTWHILSEGI-LYERRRTLNSPG*LLNSDQ 826
+V+ GSSI K+FTS PL +W T +L G+ + R + G L S
Sbjct: 468 IVISKGSSILKLFTS------KYQPLLIWRDTLLVLDLGLDIINGVRGFDLEGDGLASQS 307
Query: 827 LSAYEKIVNVVDNELDHMFFVDGYGGIGKTFL 858
L +E++ F +D G G T L
Sbjct: 306 LDKDLHATPEAKHEVESGFLLDVVVGKGSTIL 211
>TC225617 UP|Q84ZV8 (Q84ZV8) R 3 protein, complete
Length = 2697
Score = 30.8 bits (68), Expect = 3.6
Identities = 18/59 (30%), Positives = 32/59 (53%), Gaps = 4/59 (6%)
Frame = +1
Query: 826 QLSAYEKIVNVVDNELDHMFFVDGYGGIGKTFL----WNALSYHFRSEGKIVLNVASSS 880
Q+ K+++V +++ H+ + G GG+GKT L +N ++ HF E + NV S
Sbjct: 580 QVIEVRKLLDVGSHDVVHIIGIHGMGGLGKTTLALAVYNLIALHF-DESCFLQNVREES 753
>TC225659 similar to UP|Q9FUW4 (Q9FUW4) Cold acclimation protein WCOR413-like
protein, partial (48%)
Length = 619
Score = 30.8 bits (68), Expect = 3.6
Identities = 10/24 (41%), Positives = 15/24 (61%), Gaps = 1/24 (4%)
Frame = +1
Query: 411 MMMYMMCCLCCYKMMFMM-CCLCC 433
+ + ++CC CCY + M CC CC
Sbjct: 283 LFLRILCCHCCYGRAWKMDCCSCC 354
>TC221449 UP|Q9FXZ1 (Q9FXZ1) Resistance protein MG23 (Fragment), complete
Length = 1307
Score = 30.4 bits (67), Expect = 4.6
Identities = 14/44 (31%), Positives = 28/44 (62%), Gaps = 4/44 (9%)
Frame = +1
Query: 829 AYEKIVNVVDNELDHMFFVDGYGGIGKTFL----WNALSYHFRS 868
A + +++V +++ HM + G GG+GKT L +N+++ HF +
Sbjct: 589 AVKSLLDVGADDVVHMVGIHGLGGVGKTTLAVAVYNSIACHFEA 720
>TC232788 similar to UP|Q9ZVU9 (Q9ZVU9) T5A14.6 protein, partial (39%)
Length = 1069
Score = 30.4 bits (67), Expect = 4.6
Identities = 20/61 (32%), Positives = 28/61 (45%), Gaps = 1/61 (1%)
Frame = +3
Query: 218 SKYIPLQYPVLFP-LGEDRYEEKIKLNHRTTTGSVKKRVRVSLREFVAYRLQKRDNENSV 276
S Y P Y LF E++Y +++ NH TT VSL+ + Q D EN +
Sbjct: 435 SIYDPSYYSSLFEDAQENKYSSEME-NHETTQNESTSSAEVSLKSTMGRSFQAADEENGL 611
Query: 277 I 277
I
Sbjct: 612 I 614
>TC206675
Length = 957
Score = 30.4 bits (67), Expect = 4.6
Identities = 17/59 (28%), Positives = 27/59 (44%)
Frame = -2
Query: 514 FPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSIKYLFKYIT 572
F LH R ++ + R ++ + P+ M HAHINI+ N K KY++
Sbjct: 581 FHLHGERSLTQISIQRQVTQHKRRIIHHIPRKKMNIHAHINIDHNNLIMKPKKKSKYLS 405
>AW703747
Length = 414
Score = 30.4 bits (67), Expect = 4.6
Identities = 23/69 (33%), Positives = 28/69 (40%)
Frame = -3
Query: 505 CQHDVEFKCFPLHLRRDTGVTVLKNGIQLDNRSVVPYNPKLIMKYHAHINIEFCNKSNSI 564
C + F C V VLK QLD R+VV I YH N +K+N
Sbjct: 232 CSSSLGFNCGKCIATHAPMVLVLKCNFQLDERTVVVKMKNPIPLYHKDCNTS--SKTNFF 59
Query: 565 KYLFKYITK 573
K L Y +K
Sbjct: 58 KNLSCYTSK 32
>TC203719 homologue to UP|Q43437 (Q43437) Photosystem II type I chlorophyll
a/b-binding protein precursor, partial (69%)
Length = 908
Score = 30.0 bits (66), Expect = 6.1
Identities = 13/32 (40%), Positives = 17/32 (52%), Gaps = 1/32 (3%)
Frame = -3
Query: 420 CCYKMMFMMCCLCCYKMMF-MMCCYKMMFMLL 450
C + F CCL C+KMM+ M+FM L
Sbjct: 99 CALHLWFKCCCLLCFKMMY*TRFSESMLFMFL 4
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.337 0.148 0.482
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 52,860,052
Number of Sequences: 63676
Number of extensions: 789287
Number of successful extensions: 8632
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 7207
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 8289
length of query: 1134
length of database: 12,639,632
effective HSP length: 107
effective length of query: 1027
effective length of database: 5,826,300
effective search space: 5983610100
effective search space used: 5983610100
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135463.4


