

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135463.3 + phase: 0 /pseudo
(924 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
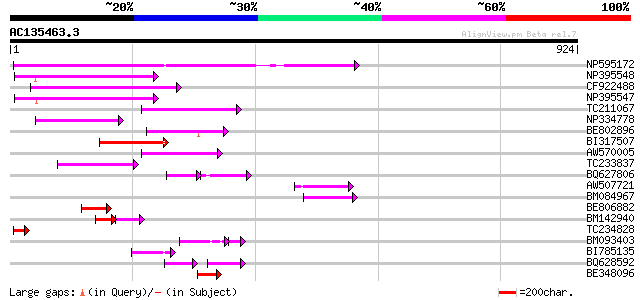
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 430 e-120
NP395548 reverse transcriptase [Glycine max] 157 3e-38
CF922488 152 5e-37
NP395547 reverse transcriptase [Glycine max] 144 1e-34
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 136 4e-32
NP334778 reverse transcriptase [Glycine max] 105 1e-22
BE802896 102 1e-21
BI317507 95 2e-19
AW570005 94 3e-19
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 89 7e-18
BQ627806 58 3e-15
AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine m... 59 1e-08
BM084967 58 2e-08
BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, pa... 53 7e-07
BM142940 37 1e-06
TC234828 47 3e-05
BM093403 44 7e-05
BI785135 45 1e-04
BQ628592 36 4e-04
BE348096 44 4e-04
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 430 bits (1105), Expect = e-120
Identities = 230/563 (40%), Positives = 336/563 (58%)
Frame = +1
Query: 7 ELKKQLEDLLDKKFVRPSASPWGAPVLLVKKKDGTMRLCIDYRQLNKVTIKNRYPLPRID 66
+++K ++++L + ++PS SP+ P+LLVKKKDG+ R C DYR LN +T+K+ +P+P +D
Sbjct: 1834 QIEKMIQEMLVQGIIQPSNSPFSLPILLVKKKDGSWRFCTDYRALNAITVKDSFPMPTVD 2013
Query: 67 DLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFM 126
+L+D+L GA FSK+DLRSGYHQI V+ ED +KTA RT +GHYE+ VMPFG+TNAP F
Sbjct: 2014 ELLDELHGAQYFSKLDLRSGYHQILVQPEDREKTAFRTHHGHYEWLVMPFGLTNAPATFQ 2193
Query: 127 EYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLS 186
MN+IF L KFV+VF DDILIYS + ++H HL+ VL LK+ +L+A+LSKC F +
Sbjct: 2194 CLMNKIFQFALRKFVLVFFDDILIYSASWKDHLKHLESVLQTLKQHQLFARLSKCSFGDT 2373
Query: 187 EVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPL 246
EV + GH +SG G++++ +KV AV W TP +V ++R FLGL GYYRRFI+ ++ +A PL
Sbjct: 2374 EVDYLGHKVSGLGVSMENTKVQAVLDWPTPNNVKQLRGFLGLTGYYRRFIKSYANIAGPL 2553
Query: 247 TQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQD 306
T L K SF+W+ + E++F +LK+ + P+L LP +PF++ DAS +G+G VL Q+
Sbjct: 2554 TDLLQK-DSFLWNNEAEAAFVKLKKAMTEAPVLSLPDFSQPFILETDASGIGVGAVLGQN 2730
Query: 307 GKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQK 366
G +AY S+ L + + EL A+ L +RHYL G++F + +D +SLK L DQ
Sbjct: 2731 GHPIAYFSKKLAPRMQKQSAYTRELLAITEALSKFRHYLLGNKFIIRTDQRSLKSLMDQS 2910
Query: 367 ELNMRQRRWLELLKDYDFGLNYHPGKANVVADALSRKTLHMSALMVKEFDLLEQFRDLSL 426
Q+ WL YDF + Y PGK N ADALSR + +
Sbjct: 2911 LQTPEQQAWLHKFLGYDFKIEYKPGKDNQAADALSR---------------------MFM 3027
Query: 427 VCELTPQSVQLGMLKINNDFLNSIREAQEVDVKLVDLMVAGNGTKDSDFKVDDQGVLRFR 486
+ P S+ FL +R D L LM D+ +G+L ++
Sbjct: 3028 LAWSEPHSI----------FLEELRARLISDPHLKQLMETYKQGADASHYTVREGLLYWK 3177
Query: 487 GRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQFVYACLTC 546
R+ IP +E+ IL+E H S + T+ LK F+W ++ DV ++ CL C
Sbjct: 3178 DRVVIPAEEEIVNKILQEYHSSPIGGHAGITRTLARLKAQFYWPKMQEDVKAYIQKCLIC 3357
Query: 547 LKSKVEHQKPAELLTPLDVPAGV 569
++K + PA LL PL +P V
Sbjct: 3358 QQAKSNNTLPAGLLQPLPIPQQV 3426
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 157 bits (396), Expect = 3e-38
Identities = 92/254 (36%), Positives = 141/254 (55%), Gaps = 19/254 (7%)
Frame = +1
Query: 8 LKKQLEDLLDKKFVRP-SASPWGAPVLLVKKKDG------------------TMRLCIDY 48
++K++ LL+ + P S S W +PVL+V KK+G + +LCIDY
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 49 RQLNKVTIKNRYPLPRIDDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGH 108
R+LN+ T K+ +PLP +D ++++L G + +D GY+QI V +D +K A +G
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 109 YEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*V 168
+ Y+ +PFG+ NAP F M IF ++K + VF+DD ++ + E L++VL
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQR 540
Query: 169 LKEKKLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGL 228
E L KC F + E GH IS GI VD +K+D + + P +V IRSFLG
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQ 720
Query: 229 AGYYRRFIEGFSKL 242
A +YRRFI+ F+K+
Sbjct: 721 ARFYRRFIKDFTKV 762
>CF922488
Length = 741
Score = 152 bits (385), Expect = 5e-37
Identities = 87/246 (35%), Positives = 134/246 (54%)
Frame = +3
Query: 35 VKKKDGTMRLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGANVFSKIDLRSGYHQIKVKD 94
V K+DG + +C+DYR LN + K+++PLP I+ L+D + FS +D SGY+QIK+
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 95 EDMQKTASRTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRT 154
EDM+KT T +G + YK M FG+ N + M +F + K + V++DD+++ SRT
Sbjct: 183 EDMEKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRT 362
Query: 155 EEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSKVDAVSQWE 214
EEEH +L+ + L++ +L +KC F + I S GI VD +KV + +
Sbjct: 363 EEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMA 542
Query: 215 TPKSVTEIRSFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRLP 274
P + +++ FLG Y RFI PL L CK + WD C +F +KQ L
Sbjct: 543 KPHTEKQVQGFLGRLNYIVRFIS*LIATCEPLFILLCKNQFVKWDHDC*VAFERIKQCLI 722
Query: 275 TNPILI 280
+L+
Sbjct: 723 NPHVLV 740
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 144 bits (364), Expect = 1e-34
Identities = 88/254 (34%), Positives = 133/254 (51%), Gaps = 19/254 (7%)
Frame = +1
Query: 8 LKKQLEDLLDKKFVRP-SASPWGAPVLLVKKKDGTM------------------RLCIDY 48
++K++ LL+ + P S S W +PV +V KK G R+CIDY
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 49 RQLNKVTIKNRYPLPRIDDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTASRTRYGH 108
R+LN+ T K+ YPLP +D ++ +L + + +D SGY+QI V +D +KTA +
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 109 YEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIVL*V 168
+ Y+ MPFG+ NA F M IF ++K + VF+DD + + +L+ VL
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQR 540
Query: 169 LKEKKLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGL 228
++ L KC F + E GH IS GI V K+D + + P +V I SFLG
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGH 720
Query: 229 AGYYRRFIEGFSKL 242
G+YRRFI+ F+K+
Sbjct: 721 VGFYRRFIKDFTKV 762
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 136 bits (343), Expect = 4e-32
Identities = 70/163 (42%), Positives = 99/163 (59%)
Frame = +1
Query: 215 TPKSVTEIRSFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRLP 274
T KSV +IRSF GLA +YRRF+ FS +A PL +L K +F W + E +F LK++L
Sbjct: 100 TLKSVGDIRSFHGLASFYRRFVPNFSTVASPLNELVKKNMAFTWGEKQEQAFALLKEKLT 279
Query: 275 TNPILILPKPEEPFVVYFDASKLGLGVVLMQDGKVVAYASRLLRIHEKNYPTHDLELAAV 334
P+L LP + F + DAS +G+ VL+Q G +AY S L NYPT+D EL A+
Sbjct: 280 KAPVLALPDFSKTFELECDASGVGVRAVLLQGGHPIAYFSEKLHSATLNYPTYDKELYAL 459
Query: 335 VLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQRRWLE 377
+ + W H+L F + SDH+SLKY+ + +LN R +W+E
Sbjct: 460 IRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKWVE 588
Score = 39.7 bits (91), Expect = 0.006
Identities = 22/76 (28%), Positives = 38/76 (49%)
Frame = +2
Query: 183 FWLSEVSFPGHIISGSGIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKL 242
F + + F G ++ +G+ +DP K+ A+ +W P V EI LG + + IEG +
Sbjct: 2 FCVDNIFFSGFVVGRNGVQMDPEKIKAIQEWPPP*KVWEI---LGASMG*QASIEGSFLI 172
Query: 243 ALPLTQLTCKGKSFVW 258
+L L L+ +W
Sbjct: 173 SLQLHHLSMSW*RRIW 220
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 105 bits (261), Expect = 1e-22
Identities = 55/143 (38%), Positives = 85/143 (58%)
Frame = +3
Query: 43 RLCIDYRQLNKVTIKNRYPLPRIDDLMDQLVGANVFSKIDLRSGYHQIKVKDEDMQKTAS 102
R+C+DYR LN+ + K+ +PLP ID LM + +FS +D SGY+QIK+ EDM+KT
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 103 RTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HL 162
T +G + YKVM FG+ N + M +F + K + ++D+++ SR EEEH +L
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNL 362
Query: 163 KIVL*VLKEKKLYAKLSKCEFWL 185
+ + L++ +L KC F L
Sbjct: 363 QNLFGQLRKYRLRLNPRKCVFGL 431
>BE802896
Length = 416
Score = 102 bits (253), Expect = 1e-21
Identities = 60/137 (43%), Positives = 82/137 (59%), Gaps = 4/137 (2%)
Frame = -2
Query: 224 SFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRLPTNPILILPK 283
SFLG AG+YRRFI F K+ALPL+ L K F ++ +C+ +F+ LK+ L T PI+ P
Sbjct: 415 SFLGHAGFYRRFIRDFRKVALPLSNLLQKEVEFDFNDKCK*AFDCLKRALITTPIIQAPD 236
Query: 284 PEEPFVVYFDASKLGLGVVLMQD----GKVVAYASRLLRIHEKNYPTHDLELAAVVLVLK 339
PF + DAS LGVVL Q +V+ Y+SR L + NY T + EL A+V L+
Sbjct: 235 WTAPFELMCDASNYALGVVLAQKIDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVFALE 56
Query: 340 IWRHYLYGSRFEVFSDH 356
+ YL G+R V+ DH
Sbjct: 55 KFHSYLLGTRIIVYIDH 5
>BI317507
Length = 359
Score = 94.7 bits (234), Expect = 2e-19
Identities = 52/113 (46%), Positives = 71/113 (62%), Gaps = 1/113 (0%)
Frame = -1
Query: 147 DILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSK 206
+ILIYS + H HL VL VLK+++L A KC F + + + GH+IS +A+D +K
Sbjct: 353 NILIYSPDWKSHIMHLTAVLDVLKKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDSNK 174
Query: 207 VDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLA-LPLTQLTCKGKSFVW 258
V +V +W PK+V + SFL L GYYR+FI+ + KLA PLT LT K F W
Sbjct: 173 VKSVIEWPVPKNVKRVCSFLRLTGYYRKFIKDYGKLAPRPLTDLT-KNDGFKW 18
>AW570005
Length = 413
Score = 94.0 bits (232), Expect = 3e-19
Identities = 52/132 (39%), Positives = 75/132 (56%)
Frame = -2
Query: 216 PKSVTEIRSFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVWDAQCESSFNELKQRLPT 275
P++ +R FL L G+YRRFI+G++ +A PL+ L K SFVW + + +F LK +
Sbjct: 406 PRTARSLRGFLRLTGFYRRFIKGYAAMAAPLSHLLTKD-SFVWSPEADVAFQALKNVVTN 230
Query: 276 NPILILPKPEEPFVVYFDASKLGLGVVLMQDGKVVAYASRLLRIHEKNYPTHDLELAAVV 335
+L LP +PF V DAS +G VL Q+G +A+ S+ T+ ELAA+
Sbjct: 229 TLVLALPDFTKPFTVETDASGSDMGAVLSQEGHPIAFFSKEFCPKLVRSSTYVHELAAIT 50
Query: 336 LVLKIWRHYLYG 347
V+K WR YL G
Sbjct: 49 NVVKKWRQYLLG 14
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 89.4 bits (220), Expect = 7e-18
Identities = 49/133 (36%), Positives = 76/133 (56%)
Frame = +2
Query: 78 FSKIDLRSGYHQIKVKDEDMQKTASRTRYGHYEYKVMPFGVTNAPGVFMEYMNRIFHAYL 137
FS +D SGY+QI + ED++KT T +G + Y+VM FG+ N + M +FH +
Sbjct: 2 FSFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMM 181
Query: 138 DKFVVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKLYAKLSKCEFWLSEVSFPGHIISG 197
K + V++DD++ SRTE EH +L + L++ +L +KC F + G I+S
Sbjct: 182 HKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVSQ 361
Query: 198 SGIAVDPSKVDAV 210
GI +DP KV A+
Sbjct: 362 KGIEIDPEKVKAL 400
>BQ627806
Length = 435
Score = 58.2 bits (139), Expect(2) = 3e-15
Identities = 31/82 (37%), Positives = 45/82 (54%)
Frame = +1
Query: 312 YASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMR 371
+ S+LLR T+ ELAA+ + +K WR YL G F + +DH+SLK L Q
Sbjct: 205 FCSKLLRAS-----TYVRELAAITVAVKKWRQYLLGHHFVILTDHRSLKELMSQAVQTPE 369
Query: 372 QRRWLELLKDYDFGLNYHPGKA 393
Q+ +L L +D+ + Y GKA
Sbjct: 370 QQIYLARLMGFDYTIQYRAGKA 435
Score = 42.7 bits (99), Expect(2) = 3e-15
Identities = 21/57 (36%), Positives = 34/57 (58%)
Frame = +3
Query: 256 FVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQDGKVVAY 312
F W+ + + +F++LK L P+L LP FVV DAS +G+G +L Q+ +A+
Sbjct: 21 FHWNEEADRAFSQLKLALCQAPVLGLPDFNSSFVVETDASGIGMGAILSQNHHPLAF 191
>AW507721 weakly similar to GP|27764548|gb polyprotein {Glycine max}, partial
(3%)
Length = 464
Score = 58.5 bits (140), Expect = 1e-08
Identities = 31/98 (31%), Positives = 54/98 (54%), Gaps = 1/98 (1%)
Frame = +1
Query: 464 MVAGNGTKDSDFKVDDQGVLRFRGRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDL 523
+ + N K D+ V V+ ++GRI +P++ +L K+I+ ESH S + T+ +
Sbjct: 172 LCSNNAGKSGDY-VLHHDVIIWKGRIMLPNDSQLLKMIMTESHASKVGGHAGTTRTIVRI 348
Query: 524 KKLFWWSGLKRDVAQFVYACLTCLKSKVEHQ-KPAELL 560
F+W ++ D+ +FV C+ C ++KV H PA LL
Sbjct: 349 NAQFYWPKMREDIMKFVQECVICQQAKVTHSLLPAGLL 462
Score = 33.1 bits (74), Expect = 0.58
Identities = 14/21 (66%), Positives = 16/21 (75%)
Frame = +2
Query: 382 YDFGLNYHPGKANVVADALSR 402
YDF + Y PGK N+ ADALSR
Sbjct: 20 YDFIIQYSPGKENIPADALSR 82
>BM084967
Length = 426
Score = 57.8 bits (138), Expect = 2e-08
Identities = 34/87 (39%), Positives = 43/87 (49%)
Frame = -2
Query: 480 QGVLRFRGRICIPDNDELKKLILEESHKSSLSIRPEATKMYHDLKKLFWWSGLKRDVAQF 539
Q ++ G I +P +L E H S TK L + F W G+++DV QF
Sbjct: 356 QDLILKNGCIWLPSGFSFIPTLLLEYHSSPTDAHIGVTKTMARLSENFTWIGIRKDVEQF 177
Query: 540 VYACLTCLKSKVEHQKPAELLTPLDVP 566
V ACL C +K E QK A LL PL VP
Sbjct: 176 VAACLDCQYTKYEAQKMAGLLCPLPVP 96
>BE806882 similar to GP|27764548|gb polyprotein {Glycine max}, partial (3%)
Length = 153
Score = 52.8 bits (125), Expect = 7e-07
Identities = 26/48 (54%), Positives = 32/48 (66%)
Frame = -1
Query: 118 VTNAPGVFMEYMNRIFHAYLDKFVVVFIDDILIYSRTEEEHA*HLKIV 165
+TNAP F MN IF L K+V+VF DDIL+YS T EH HL++V
Sbjct: 153 LTNAPTSFQCLMNHIFQHALRKYVLVFFDDILVYSSTWHEHLCHLEVV 10
>BM142940
Length = 422
Score = 37.0 bits (84), Expect(2) = 1e-06
Identities = 15/47 (31%), Positives = 25/47 (52%)
Frame = -3
Query: 173 KLYAKLSKCEFWLSEVSFPGHIISGSGIAVDPSKVDAVSQWETPKSV 219
K+ A KC F + + H+IS G++ DP K++ + W PK +
Sbjct: 165 KVGANRKKCSFGCKKFEYLVHMISVKGVSADPKKLEVMVAWPIPKEM 25
Score = 34.7 bits (78), Expect(2) = 1e-06
Identities = 19/34 (55%), Positives = 21/34 (60%)
Frame = -1
Query: 141 VVVFIDDILIYSRTEEEHA*HLKIVL*VLKEKKL 174
+ F DILIYS E +H HLKIVL KE KL
Sbjct: 260 ICFFSYDILIYSINEGKHEKHLKIVLQTFKEAKL 159
>TC234828
Length = 857
Score = 47.4 bits (111), Expect = 3e-05
Identities = 20/26 (76%), Positives = 24/26 (91%)
Frame = +3
Query: 6 AELKKQLEDLLDKKFVRPSASPWGAP 31
AE+K Q++DLL K+FVRPSASPWGAP
Sbjct: 777 AEVKAQVQDLLSKQFVRPSASPWGAP 854
>BM093403
Length = 419
Score = 43.5 bits (101), Expect(2) = 7e-05
Identities = 26/81 (32%), Positives = 45/81 (55%)
Frame = -1
Query: 278 ILILPKPEEPFVVYFDASKLGLGVVLMQDGKVVAYASRLLRIHEKNYPTHDLELAAVVLV 337
+L+ P +PF+V DA G+G ++MQ+ + +AY S+ L I N EL A+V+
Sbjct: 404 VLVTPNFSKPFIVEADALGSGIGAIVMQESRPIAYYSKALLICPYNIVMQ--ELLALVMP 231
Query: 338 LKIWRHYLYGSRFEVFSDHKS 358
++ W ++ + VF+ KS
Sbjct: 230 IQYWCP-IFWAENSVFTTKKS 171
Score = 21.9 bits (45), Expect(2) = 7e-05
Identities = 10/28 (35%), Positives = 14/28 (49%)
Frame = -2
Query: 357 KSLKYLFDQKELNMRQRRWLELLKDYDF 384
KSL +L +Q Q+ W + YDF
Sbjct: 178 KSLHHLLEQ*ITTQNQQDWFVKILRYDF 95
>BI785135
Length = 420
Score = 45.1 bits (105), Expect = 1e-04
Identities = 22/71 (30%), Positives = 36/71 (49%)
Frame = +1
Query: 199 GIAVDPSKVDAVSQWETPKSVTEIRSFLGLAGYYRRFIEGFSKLALPLTQLTCKGKSFVW 258
G+ ++P ++ + +W TP S+ I F L +Y+RF+ FS L PL +L + W
Sbjct: 208 GVPMNPKRIKVIPEWPTPPSIR*IWGFHNLTNFYKRFVP*FSILVAPLIELV-RNHVPSW 384
Query: 259 DAQCESSFNEL 269
+ E F L
Sbjct: 385 EEAQEMGFQTL 417
>BQ628592
Length = 423
Score = 36.2 bits (82), Expect(2) = 4e-04
Identities = 15/54 (27%), Positives = 30/54 (54%)
Frame = -1
Query: 252 KGKSFVWDAQCESSFNELKQRLPTNPILILPKPEEPFVVYFDASKLGLGVVLMQ 305
K ++ +W++ + +F ++KQ L +L+ P PF++Y +G VL+Q
Sbjct: 417 KNQAILWNSNYQEAFEKIKQSLANPSVLMPPVTGRPFLLYMTMLDESMGCVLVQ 256
Score = 26.6 bits (57), Expect(2) = 4e-04
Identities = 15/61 (24%), Positives = 26/61 (42%)
Frame = -3
Query: 323 NYPTHDLELAAVVLVLKIWRHYLYGSRFEVFSDHKSLKYLFDQKELNMRQRRWLELLKDY 382
NY + +V R Y+ + S +KY+F++ L R RW LL ++
Sbjct: 187 NYSMLERTCCTLVWASHRLRQYMLSHTTWLISKMDPVKYIFEKPALTGRIARWQVLLSEF 8
Query: 383 D 383
+
Sbjct: 7 N 5
>BE348096
Length = 445
Score = 43.5 bits (101), Expect = 4e-04
Identities = 18/39 (46%), Positives = 24/39 (61%)
Frame = -2
Query: 307 GKVVAYASRLLRIHEKNYPTHDLELAAVVLVLKIWRHYL 345
G+V Y ++ H + YPTHDLEL A+V WRHY+
Sbjct: 183 GQVATYVLP*VKTH*RIYPTHDLELVAMVCAFTFWRHYI 67
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.341 0.151 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 41,350,923
Number of Sequences: 63676
Number of extensions: 584101
Number of successful extensions: 3991
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 3955
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3985
length of query: 924
length of database: 12,639,632
effective HSP length: 106
effective length of query: 818
effective length of database: 5,889,976
effective search space: 4818000368
effective search space used: 4818000368
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC135463.3


