

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135463.1 - phase: 0 /pseudo
(488 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
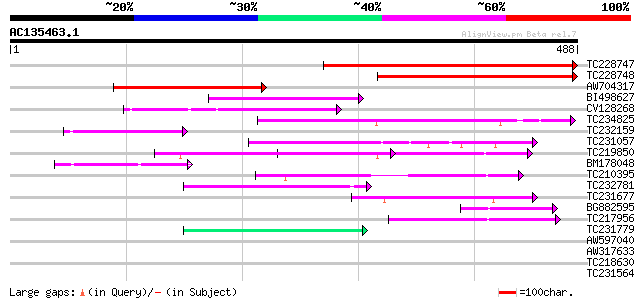
TC228747 weakly similar to UP|Q9XI21 (Q9XI21) F9L1.43, partial (... 348 2e-96
TC228748 weakly similar to UP|Q9LRP6 (Q9LRP6) DNA-binding protei... 248 6e-66
AW704317 weakly similar to GP|6958202|gb| DNA-binding protein {T... 208 4e-54
BI498627 71 1e-12
CV128268 70 3e-12
TC234825 69 5e-12
TC232159 63 3e-10
TC231057 homologue to UP|Q9W1K2 (Q9W1K2) CG12491-PA, partial (8%) 60 3e-09
TC219850 weakly similar to UP|Q9FKI5 (Q9FKI5) Emb|CAB69839.1, pa... 59 6e-09
BM178048 57 2e-08
TC210395 similar to UP|Q9FGR7 (Q9FGR7) Similarity to salt-induci... 57 2e-08
TC232781 similar to UP|Q9ZU27 (Q9ZU27) F5F19.2 protein, partial ... 51 1e-06
TC231677 similar to UP|O23677 (O23677) T7I23.8 protein, partial ... 48 9e-06
BG882595 weakly similar to GP|3107905|dbj| leaf protein {Ipomoea... 48 1e-05
TC217956 similar to UP|O65793 (O65793) Leaf protein, partial (30%) 44 2e-04
TC231779 weakly similar to UP|Q9SGV7 (Q9SGV7) F1N19.15, partial ... 42 5e-04
AW597040 weakly similar to GP|1931651|gb| membrane-associated sa... 39 0.004
AW317633 similar to PIR|T46163|T46 nodulin / glutamate-ammonia l... 37 0.015
TC218630 weakly similar to PIR|D86391|D86391 T1K7.17 protein - A... 37 0.026
TC231564 similar to GB|AAP31962.1|30387593|BT006618 At3g06430 {A... 36 0.034
>TC228747 weakly similar to UP|Q9XI21 (Q9XI21) F9L1.43, partial (35%)
Length = 924
Score = 348 bits (894), Expect = 2e-96
Identities = 165/218 (75%), Positives = 191/218 (86%)
Frame = +1
Query: 271 EAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCESKPRVEDCLAAIEAWGR 330
EA+LKE+EGENLKEN WVC TLLRLYA LG+ADEVERIWKVCESKPRVEDCLAA+EAWG+
Sbjct: 4 EAMLKEMEGENLKENQWVCATLLRLYANLGKADEVERIWKVCESKPRVEDCLAAVEAWGK 183
Query: 331 LKKIEEAEAVFEMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTW 390
L KIEEAEAVFEM+S KWKL ++NY LLKIY +KML KGK+L+K M DSG IGP TW
Sbjct: 184 LNKIEEAEAVFEMVSKKWKLNSKNYSVLLKIYANNKMLTKGKELVKLMADSGVRIGPLTW 363
Query: 391 DALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLR 450
DALV LY+QAGEVEKAD++L KA+QQN+++PMFTT++ I+EQYAKRGDVHN+EKIF ++R
Sbjct: 364 DALVKLYIQAGEVEKADSILHKAIQQNQLQPMFTTYLAILEQYAKRGDVHNSEKIFLKMR 543
Query: 451 QANYISRISPFHALAQAYKNAKLPAYGIRERMKADNLF 488
QA Y SRIS F L QAY NAK+PAYGIRER+KADNLF
Sbjct: 544 QAGYTSRISQFQVLIQAYVNAKVPAYGIRERIKADNLF 657
>TC228748 weakly similar to UP|Q9LRP6 (Q9LRP6) DNA-binding protein
(AT3g15590/MQD17_5), partial (25%)
Length = 763
Score = 248 bits (632), Expect = 6e-66
Identities = 120/172 (69%), Positives = 140/172 (80%)
Frame = +2
Query: 317 RVEDCLAAIEAWGRLKKIEEAEAVFEMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIK 376
R +DC +EAWG L+ IEEAEAV EM S KWKL + Y LLKIY +KML KGKDLIK
Sbjct: 5 RXDDCXXXVEAWGXLEXIEEAEAVXEMASKKWKLNSXXYSILLKIYANNKMLAKGKDLIK 184
Query: 377 TMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKR 436
M DSG IGP TW+ALV LY+QAGEVEKAD+VLQKA+QQ++++PMFTT++ I+EQYAKR
Sbjct: 185 RMADSGLRIGPLTWNALVKLYIQAGEVEKADSVLQKAIQQSQLQPMFTTYLDILEQYAKR 364
Query: 437 GDVHNAEKIFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNLF 488
GDVHN+EKIF ++RQA Y SRIS F L QAY NAK+PAYGIRERMKADNLF
Sbjct: 365 GDVHNSEKIFLKMRQAGYTSRISQFKVLMQAYVNAKVPAYGIRERMKADNLF 520
>AW704317 weakly similar to GP|6958202|gb| DNA-binding protein {Triticum
aestivum}, partial (21%)
Length = 399
Score = 208 bits (530), Expect = 4e-54
Identities = 101/132 (76%), Positives = 115/132 (86%)
Frame = +2
Query: 90 RRKMYGRALQALDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELL 149
RRKMYGRA Q WLESNKKLEF E +YAS+LDLIAKLRGLP+AEKY+E VP SFRGELL
Sbjct: 2 RRKMYGRAFQLFQWLESNKKLEFMESDYASQLDLIAKLRGLPQAEKYIESVPESFRGELL 181
Query: 150 YRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDKKKIADVLLMMEK 209
YRTLLANCAS NL +E+ FNKM++L P+T FACNQLLL+YKK+DKKKIADVLL+MEK
Sbjct: 182 YRTLLANCASQNNLIASEKIFNKMKDLDLPLTVFACNQLLLLYKKLDKKKIADVLLLMEK 361
Query: 210 ENVKPSSYTYKI 221
ENVKPS +TY+I
Sbjct: 362 ENVKPSLFTYRI 397
>BI498627
Length = 421
Score = 71.2 bits (173), Expect = 1e-12
Identities = 44/135 (32%), Positives = 71/135 (52%), Gaps = 2/135 (1%)
Frame = +3
Query: 172 KMRELGFPVTAFACNQLLLIYKKIDK-KKIADVLLMMEKENVKPSSYTYKILIDVKGLSN 230
KM+EL P+++ N L+ +Y K+ + +KI ++ M+ NV SYTY + + N
Sbjct: 15 KMKELSLPLSSMPYNSLMTLYTKVGQPEKIPSLIQEMKASNVMLDSYTYNVWMRALAAVN 194
Query: 231 DIDGMSQIVETMKAEG-CELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVC 289
DI G+ ++ + MK G D T ++LA + AGL +K E LKE+E N +++
Sbjct: 195 DISGVERVHDEMKRGGQVTGDWTTYSNLASIFVDAGLFDKAEVALKELEKRNAFKDLTAY 374
Query: 290 PTLLRLYAILGRADE 304
L+ LY G E
Sbjct: 375 QFLITLYGRTGNLYE 419
>CV128268
Length = 864
Score = 69.7 bits (169), Expect = 3e-12
Identities = 50/190 (26%), Positives = 100/190 (52%), Gaps = 3/190 (1%)
Frame = +2
Query: 99 QALDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCA 158
Q +W+ S+K L + ++ A +LDLI ++ G+ AE+YL+ + + + ++ LL NC
Sbjct: 269 QVSEWM-SSKGLPISSRDQAVQLDLIGRVHGVESAERYLQSLSDGDKTWKVHGALL-NCY 442
Query: 159 SLENL-RKTEETFNKMRELGFPVTAFACNQLLLIYKKIDK-KKIADVLLMMEKENVKPSS 216
E L K+ KM+++GF V+ N ++ +Y + + +K+ VL M+K+ V P+
Sbjct: 443 VREGLVDKSLSLMQKMKDMGF-VSFLNYNNIMSLYTQTQQYEKVPGVLEQMKKDGVPPNI 619
Query: 217 YTYKILIDVKGLSNDIDGMSQIVETMKAE-GCELDHLTRASLARHYAAAGLTEKTEAILK 275
++Y+I I+ + D+ + +++E M+ E +D +T + + Y A + L
Sbjct: 620 FSYRICINSYCVRGDLANVEKLLEEMEREPHIGIDWITYSMVTNFYIKADMRXXXLVCLM 799
Query: 276 EIEGENLKEN 285
+ E + N
Sbjct: 800 KCEXXTHRGN 829
>TC234825
Length = 874
Score = 68.9 bits (167), Expect = 5e-12
Identities = 66/283 (23%), Positives = 118/283 (41%), Gaps = 9/283 (3%)
Frame = +1
Query: 214 PSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAI 273
P T+ + + ND++ +++ +K + D +T ++L Y EK A
Sbjct: 7 PDIVTFNLWLAACASQNDVETAERVLLELKKAKIDPDWVTYSTLTNLYIKNASLEKAGAT 186
Query: 274 LKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCES---KPRVEDCLAAIEAWGR 330
+KE+E ++ +LL L+ +G D+V RIW+ ++ K + + I + +
Sbjct: 187 VKEMENRTSRKTRVAYSSLLSLHTNMGNKDDVNRIWEKMKASFRKMNDNEYICMISSLLK 366
Query: 331 LKKIEEAEAVF-EMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTT 389
L AE ++ E S R LL YI + +D + G TT
Sbjct: 367 LGDFAGAEDLYREWESVSGTNDVRVSNILLGSYINQDQMEMAEDFCNQIVQKGVIPCYTT 546
Query: 390 WDALVSLYVQAGEVEKADTVLQKALQQ-NKMKP----MFTTFMTIMEQYAKRGDVHNAEK 444
W+ Y++ +VEK KA+ K P + F I EQ +G AE+
Sbjct: 547 WELFTWGYLKRKDVEKFLDYFSKAISSVTKWSPDQRLVQEAFKIIEEQAHTKG----AEQ 714
Query: 445 IFYRLRQANYISRISPFHALAQAYKNAKLPAYGIRERMKADNL 487
+ LR A +++ + ++ + Y A + ERM+ DN+
Sbjct: 715 LLVILRNAGHVN-TNIYNLFLKTYATAGKMPMIVAERMRKDNV 840
>TC232159
Length = 483
Score = 63.2 bits (152), Expect = 3e-10
Identities = 32/107 (29%), Positives = 60/107 (55%)
Frame = +1
Query: 47 RSELFKAIVSVSGLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRALQALDWLES 106
RS +F+ + + S + L KWV +G ++ ++ LRR + Y AL+ +W+ S
Sbjct: 169 RSRIFR--LRLPKRSATNVLQKWVLQGNPITLSQLRDISKELRRSQRYKHALEISEWMVS 342
Query: 107 NKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTL 153
N++ E ++ +YA+++DL K+ G+ AE+Y E +P + + Y L
Sbjct: 343 NEEYELSDSDYAARIDLTTKVFGIDAAERYFEGLPLATKTTETYTAL 483
>TC231057 homologue to UP|Q9W1K2 (Q9W1K2) CG12491-PA, partial (8%)
Length = 1050
Score = 59.7 bits (143), Expect = 3e-09
Identities = 58/260 (22%), Positives = 118/260 (45%), Gaps = 11/260 (4%)
Frame = +3
Query: 206 MMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEG-CELDHLTRASLARHYAAA 264
+M+K + S +TY I ++ SND+ G ++ E MK E ++ T ++LA Y
Sbjct: 9 LMKKRTIPMSPFTYHIWMNSCASSNDLGGAERVYEEMKTENEGQIGWHTYSNLASIYVKF 188
Query: 265 GLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCESKPRVEDCLAA 324
EK E +LK +E + + LL LYA G EV R+W +S V + +
Sbjct: 189 KDFEKAEMMLKMLEEQVKPKQRDAYHCLLGLYAGTGNLGEVHRVWDSLKSVSPVTN-FSY 365
Query: 325 IEAWGRLKKIEEAEAVFEMMSNKWKLTARNYESLL-----KIYIRHKMLNKGKDLIKTMG 379
+ L+++ + E + + +W+ + +Y++ L ++ ML + + + +
Sbjct: 366 LVMLSTLRRLNDMEGLTKCF-KEWEASCVSYDARLVSVCVSAHLNQNMLEEAELVFEEA- 539
Query: 380 DSGCTIGP--TTWDALVSLYVQAGEVEKADTVLQKALQQ---NKMKPMFTTFMTIMEQYA 434
S + GP + + +++ E++ A L+ AL + +K +P ++ Y
Sbjct: 540 -SRRSKGPFFRVREEFMKFFLKKHELDAAVRHLEAALSEVKGDKWRPSPQVVGAFLKYYE 716
Query: 435 KRGDVHNAEKIFYRLRQANY 454
+ DV +++ L+ N+
Sbjct: 717 EETDVDGVDELSKILKANNF 776
>TC219850 weakly similar to UP|Q9FKI5 (Q9FKI5) Emb|CAB69839.1, partial (53%)
Length = 1153
Score = 58.5 bits (140), Expect = 6e-09
Identities = 47/224 (20%), Positives = 102/224 (44%), Gaps = 4/224 (1%)
Frame = +2
Query: 231 DIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCP 290
++D ++V K + D L Y GL EKT ++KE+E ++K + +
Sbjct: 68 EVDVAEELVREAKGKTTIKDPEVYLKLVHMYIEEGLLEKTLEVVKEMEDADVKVSDCILC 247
Query: 291 TLLRLYAILGRADEVERIWKVCESK---PRVEDCLAAIEAWGRLKKIEEAEAVF-EMMSN 346
T++ ++ ++++ SK P + I A+ RL + +AE VF EM
Sbjct: 248 TVVNGFSKKRGFSAAVKVFEELISKGNEPGQVTYASVINAYWRLGQYSKAEEVFLEMEQK 427
Query: 347 KWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKA 406
+ Y +++ +Y R + L+ M + GC +++L+ ++ + +++
Sbjct: 428 GFDKCVYAYSTMIVMYGRTGRVRSAMKLVAKMKERGCKPNVWIYNSLIDMHGRDKNLKQL 607
Query: 407 DTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLR 450
+ L K +++ ++ P ++ +I+ Y+K G+ K+F R
Sbjct: 608 EK-LWKEMKRRRVAPDKVSYTSIIGAYSKAGEFETCVKLFNEYR 736
Score = 56.2 bits (134), Expect = 3e-08
Identities = 45/211 (21%), Positives = 95/211 (44%), Gaps = 3/211 (1%)
Frame = +2
Query: 125 AKLRGLPKAEKYLEHVPNSFR--GELLYRTLLANCASLENLRKTEETFNKMRELGFPVTA 182
+K RG A K E + + G++ Y +++ L K EE F +M + GF
Sbjct: 266 SKKRGFSAAVKVFEELISKGNEPGQVTYASVINAYWRLGQYSKAEEVFLEMEQKGFDKCV 445
Query: 183 FACNQLLLIYKKIDKKKIADVLLMMEKEN-VKPSSYTYKILIDVKGLSNDIDGMSQIVET 241
+A + ++++Y + + + A L+ KE KP+ + Y LID+ G ++ + ++ +
Sbjct: 446 YAYSTMIVMYGRTGRVRSAMKLVAKMKERGCKPNVWIYNSLIDMHGRDKNLKQLEKLWKE 625
Query: 242 MKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKENMWVCPTLLRLYAILGR 301
MK D ++ S+ Y+ AG E + E + + ++ +++ +G+
Sbjct: 626 MKRRRVAPDKVSYTSIIGAYSKAGEFETCVKLFNEYRMNGGLIDRAMAGIMVGVFSKVGQ 805
Query: 302 ADEVERIWKVCESKPRVEDCLAAIEAWGRLK 332
DE+ ++ + +++ D AW K
Sbjct: 806 VDELVKLLQDMKAEGTRLDQRLYQSAWNAFK 898
Score = 35.0 bits (79), Expect = 0.076
Identities = 51/250 (20%), Positives = 99/250 (39%), Gaps = 46/250 (18%)
Frame = +2
Query: 63 DSALDKWVEKGKELSRQEIGLA-------LNSLRRRKMYGRALQALDWLESNKKLEFTEK 115
+ L+K +E KE+ ++ ++ +N +++ + A++ + L S K E +
Sbjct: 167 EGLLEKTLEVVKEMEDADVKVSDCILCTVVNGFSKKRGFSAAVKVFEELIS-KGNEPGQV 343
Query: 116 EYASKLDLIAKLRGLPKAEK-YLEHVPNSF-RGELLYRTLLANCASLENLRKTEETFNKM 173
YAS ++ +L KAE+ +LE F + Y T++ +R + KM
Sbjct: 344 TYASVINAYWRLGQYSKAEEVFLEMEQKGFDKCVYAYSTMIVMYGRTGRVRSAMKLVAKM 523
Query: 174 RELGFPVTAFACNQLLLIYKKIDK--KKIADVLLMMEKENVKPSSYTYK----------- 220
+E G + N L+ ++ + DK K++ + M++ V P +Y
Sbjct: 524 KERGCKPNVWIYNSLIDMHGR-DKNLKQLEKLWKEMKRRRVAPDKVSYTSIIGAYSKAGE 700
Query: 221 ------------------------ILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRAS 256
I++ V +D + ++++ MKAEG LD S
Sbjct: 701 FETCVKLFNEYRMNGGLIDRAMAGIMVGVFSKVGQVDELVKLLQDMKAEGTRLDQRLYQS 880
Query: 257 LARHYAAAGL 266
+ AGL
Sbjct: 881 AWNAFKDAGL 910
Score = 34.7 bits (78), Expect = 0.099
Identities = 30/184 (16%), Positives = 69/184 (37%), Gaps = 1/184 (0%)
Frame = +2
Query: 163 LRKTEETFNKMRELGFPVT-AFACNQLLLIYKKIDKKKIADVLLMMEKENVKPSSYTYKI 221
L KT E +M + V+ C + KK V + + +P TY
Sbjct: 176 LEKTLEVVKEMEDADVKVSDCILCTVVNGFSKKRGFSAAVKVFEELISKGNEPGQVTYAS 355
Query: 222 LIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGEN 281
+I+ ++ M+ +G + +++ Y G ++ +++
Sbjct: 356 VINAYWRLGQYSKAEEVFLEMEQKGFDKCVYAYSTMIVMYGRTGRVRSAMKLVAKMKERG 535
Query: 282 LKENMWVCPTLLRLYAILGRADEVERIWKVCESKPRVEDCLAAIEAWGRLKKIEEAEAVF 341
K N+W+ +L+ ++ ++E++WK + + D ++ G K E E
Sbjct: 536 CKPNVWIYNSLIDMHGRDKNLKQLEKLWKEMKRRRVAPDKVSYTSIIGAYSKAGEFETCV 715
Query: 342 EMMS 345
++ +
Sbjct: 716 KLFN 727
>BM178048
Length = 422
Score = 57.0 bits (136), Expect = 2e-08
Identities = 40/119 (33%), Positives = 60/119 (49%)
Frame = +2
Query: 39 KKSFSSRRRSELFKAIVSVSGLSVDSALDKWVEKGKELSRQEIGLALNSLRRRKMYGRAL 98
KK +LFK S L V +L+ +V+ K + + E+G L LR RK+Y AL
Sbjct: 50 KKYLEEALYMKLFKD--GSSQLIVRQSLNNFVKSRKRVYKWEVGDTLKKLRDRKLYQPAL 223
Query: 99 QALDWLESNKKLEFTEKEYASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANC 157
+ + + ++ T ++A LDL+AK RG+ AE Y +P + L Y LL NC
Sbjct: 224 KLSETMAKRNMIK-TVSDHAIHLDLLAKARGITAAENYFVSLPEPSKNHLCYGALL-NC 394
>TC210395 similar to UP|Q9FGR7 (Q9FGR7) Similarity to salt-inducible protein,
partial (33%)
Length = 954
Score = 56.6 bits (135), Expect = 2e-08
Identities = 49/233 (21%), Positives = 95/233 (40%), Gaps = 2/233 (0%)
Frame = +1
Query: 212 VKPSSYTYKILIDVKGLSNDIDGMS--QIVETMKAEGCELDHLTRASLARHYAAAGLTEK 269
+KP++ +Y LI G ++ M+ MK G + + +L Y+ +GL EK
Sbjct: 13 LKPNATSYTCLIIAYGKQKNMSDMAAADAFLKMKKVGVKPTSQSYTALIHAYSVSGLHEK 192
Query: 270 TEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKVCESKPRVEDCLAAIEAWG 329
A + ++ E +K ++ TLL + G A + IWK+
Sbjct: 193 AYAAFENMQNEGIKPSIETYTTLLNAFRHAGDAQTLMEIWKL------------------ 318
Query: 330 RLKKIEEAEAVFEMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTT 389
M+S K + T + L+ + + + + +++I G G T
Sbjct: 319 -------------MISEKVEGTGATFNILVDGFAKQGLFMEAREVISEFGKVGLKPTVVT 459
Query: 390 WDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNA 442
++ L++ Y + G+ K +L K + K+KP T+ T++ + + D A
Sbjct: 460 YNMLINAYARGGQHSKLPQLL-KEMAVLKLKPDSVTYSTMIFAFVRVRDFRRA 615
>TC232781 similar to UP|Q9ZU27 (Q9ZU27) F5F19.2 protein, partial (3%)
Length = 1118
Score = 51.2 bits (121), Expect = 1e-06
Identities = 40/163 (24%), Positives = 77/163 (46%), Gaps = 1/163 (0%)
Frame = +2
Query: 150 YRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLLLIYKKIDK-KKIADVLLMME 208
+ LL+ + + ++ K EE N+M + G + + N +L +Y ++ + K+ +VL +ME
Sbjct: 347 HMVLLSAYSKMGSVNKCEEILNQMCKSGLKLDTYVLNSMLNLYGRLGQFGKMEEVLRVME 526
Query: 209 KENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTE 268
K + TY ILI+ G + I+ M + + + ++G + D +T S Y+ L
Sbjct: 527 KGSYVADISTYNILINRYGQAGFIERMEDLFQLLPSKGLKPDVVTWTSRIGAYSKKKLYL 706
Query: 269 KTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERIWKV 311
K I +E+ + + LL A D+ E++ V
Sbjct: 707 KCLEIFEEMIDDGCYPDGGTAKVLL---AACSNEDQTEQVTTV 826
>TC231677 similar to UP|O23677 (O23677) T7I23.8 protein, partial (14%)
Length = 833
Score = 48.1 bits (113), Expect = 9e-06
Identities = 41/168 (24%), Positives = 76/168 (44%), Gaps = 8/168 (4%)
Frame = +2
Query: 295 LYAILGRADEVERIWKVCESK-PRVEDC--LAAIEAWGRLKKIEEAEAVFE-MMSNKWKL 350
LY +G+ DEV R+W +S P + + A I + +L IE AE ++E +S K
Sbjct: 5 LYGSVGKKDEVCRVWNTYKSIFPSIPNLGYHAIISSLVKLDDIEVAEKLYEEWISVKSSY 184
Query: 351 TARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTIGPTTWDALVSLYVQAGEVEKADTVL 410
R L+ Y++ +K + M + GC TW+ L ++ + +A + L
Sbjct: 185 DPRIGNLLMGWYVKKGDTDKALSFFEQMLNDGCIPNSNTWEILSEGHIADKRISEALSCL 364
Query: 411 QKALQ----QNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYRLRQANY 454
++A +P + +E ++ D+ +AE + LRQ+ +
Sbjct: 365 KEAFMVAGGSKSWRPKPSYLSAFLELCQEQDDMESAEVLIGLLRQSKF 508
>BG882595 weakly similar to GP|3107905|dbj| leaf protein {Ipomoea nil},
partial (14%)
Length = 400
Score = 47.8 bits (112), Expect = 1e-05
Identities = 26/83 (31%), Positives = 45/83 (53%)
Frame = +2
Query: 389 TWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYR 448
T++ + Y Q GE EKA+ ++ K +Q NK+KP T I+ Y K G++ A + YR
Sbjct: 20 TYNTMARAYAQNGETEKAERLILK-MQYNKVKPNERTCGIIISGYCKEGNMTEALRFLYR 196
Query: 449 LRQANYISRISPFHALAQAYKNA 471
+++ F++L + Y +A
Sbjct: 197 MKELGVHPNPVVFNSLIKGYLDA 265
>TC217956 similar to UP|O65793 (O65793) Leaf protein, partial (30%)
Length = 1224
Score = 43.9 bits (102), Expect = 2e-04
Identities = 35/149 (23%), Positives = 65/149 (43%), Gaps = 1/149 (0%)
Frame = +1
Query: 327 AWGRLKKIEEAEA-VFEMMSNKWKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSGCTI 385
A+ + + E AE + +M N K R ++ Y + + + + M + G
Sbjct: 25 AYDQNGETERAERLILKMPYNIVKPNERTCGIIISGYCKEGNMPEALRFLYRMKELGVDP 204
Query: 386 GPTTWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKI 445
P +++L+ Y+ + D L +++ +KP TF TIM ++ G + N E+I
Sbjct: 205 NPVVFNSLIKGYLDTTDTNGVDEALT-LMEEFGIKPDVVTFSTIMNAWSSAGLMENCEEI 381
Query: 446 FYRLRQANYISRISPFHALAQAYKNAKLP 474
F + +A I + LA+ Y A P
Sbjct: 382 FNDMVKAGIEPDIHAYSILAKGYVRAGQP 468
>TC231779 weakly similar to UP|Q9SGV7 (Q9SGV7) F1N19.15, partial (7%)
Length = 620
Score = 42.4 bits (98), Expect = 5e-04
Identities = 31/160 (19%), Positives = 61/160 (37%), Gaps = 1/160 (0%)
Frame = +1
Query: 150 YRTLLANCASLENLRKTEETFNKMRELGFPVTAFACNQLL-LIYKKIDKKKIADVLLMME 208
Y +L+ + N+M G P LL + K + K + + M+
Sbjct: 43 YNSLIDGLCKSGRITSALNLMNEMHHRGQPADVVTYTSLLDALCKNQNHDKATALFMKMK 222
Query: 209 KENVKPSSYTYKILIDVKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTE 268
K ++P+ YTY LID + ++ + + +GC +D T + G+ +
Sbjct: 223 KRRIQPTMYTYTALIDGLCKGGRLKNAQELFQHLLVKGCCIDVWTYTVMISGLCKEGMLD 402
Query: 269 KTEAILKEIEGENLKENMWVCPTLLRLYAILGRADEVERI 308
+ A+ ++E N ++R D+ E+I
Sbjct: 403 EALAMKSKMEDNGCIPNAVTFEIIIRSLFEKDENDKAEKI 522
>AW597040 weakly similar to GP|1931651|gb| membrane-associated salt-inducible
protein isolog; 88078-84012 {Arabidopsis thaliana},
partial (19%)
Length = 384
Score = 39.3 bits (90), Expect = 0.004
Identities = 33/137 (24%), Positives = 60/137 (43%), Gaps = 1/137 (0%)
Frame = +3
Query: 117 YASKLDLIAKLRGLPKAEKYLEHVPNSFRGELLYRTLLANCASLENLRKTEETFNKMREL 176
YA L+LI +L +H G ++Y T++A CAS + E FN+M++
Sbjct: 3 YAKALELIQEL----------QHNKLQMDG-VIYGTIMAVCASNTKWEEAEYYFNQMKDE 149
Query: 177 GFPVTAFACNQLLLIYKKIDKKKIADVLLM-MEKENVKPSSYTYKILIDVKGLSNDIDGM 235
G + + L+ Y K AD+L+ M+ E + P+ L+ V +
Sbjct: 150 GHTPNVYHYSSLINAYSACGNYKKADMLIQDMKSEGLVPNKVILTTLLKVYVKGGLFEKS 329
Query: 236 SQIVETMKAEGCELDHL 252
+++ +K+ G D +
Sbjct: 330 RELLAELKSLGYAEDEM 380
>AW317633 similar to PIR|T46163|T46 nodulin / glutamate-ammonia ligase-like
protein - Arabidopsis thaliana, partial (26%)
Length = 484
Score = 37.4 bits (85), Expect = 0.015
Identities = 19/83 (22%), Positives = 43/83 (50%)
Frame = +1
Query: 389 TWDALVSLYVQAGEVEKADTVLQKALQQNKMKPMFTTFMTIMEQYAKRGDVHNAEKIFYR 448
T++ ++ ++ +AGE+EK D K ++ +KP T+ +++ Y+K G + + I
Sbjct: 34 TYNTVIEVFGKAGEIEKMDQHFLK-MKHLGVKPNSITYCSLVSAYSKVGCIDKVDSIMRH 210
Query: 449 LRQANYISRISPFHALAQAYKNA 471
+ ++ + F+ + AY A
Sbjct: 211 VENSDVVLDTPFFNCIISAYGQA 279
>TC218630 weakly similar to PIR|D86391|D86391 T1K7.17 protein - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (52%)
Length = 1513
Score = 36.6 bits (83), Expect = 0.026
Identities = 28/129 (21%), Positives = 58/129 (44%), Gaps = 8/129 (6%)
Frame = +1
Query: 128 RGLPKAEKYLEHVPNSFRGELLYRTLLA------NCASLENLRKTEETFNKMRE-LGFPV 180
+G + + N R E Y+++ A CA++ +L + +TF + G
Sbjct: 508 KGFETLDNVYFQLENLNRAEPPYKSVAALNCVILGCANIWDLDRAYQTFESIGSTFGLIP 687
Query: 181 TAFACNQLLLIYKKIDKKKIAD-VLLMMEKENVKPSSYTYKILIDVKGLSNDIDGMSQIV 239
+ N L+ + K+ K A V + +KP++ +Y +L+D ++ D+ ++
Sbjct: 688 DIHSYNGLMYAFGKLKKTHEATRVFEHLVSLGLKPNAKSYSLLVDAHLINRDVKSALAVI 867
Query: 240 ETMKAEGCE 248
+ M+A G E
Sbjct: 868 DDMRAAGYE 894
Score = 29.3 bits (64), Expect = 4.1
Identities = 26/95 (27%), Positives = 48/95 (50%), Gaps = 10/95 (10%)
Frame = +1
Query: 298 ILGRAD--EVERIWKVCESK-------PRVEDCLAAIEAWGRLKKIEEAEAVFE-MMSNK 347
ILG A+ +++R ++ ES P + + A+G+LKK EA VFE ++S
Sbjct: 604 ILGCANIWDLDRAYQTFESIGSTFGLIPDIHSYNGLMYAFGKLKKTHEATRVFEHLVSLG 783
Query: 348 WKLTARNYESLLKIYIRHKMLNKGKDLIKTMGDSG 382
K A++Y L+ ++ ++ + +I M +G
Sbjct: 784 LKPNAKSYSLLVDAHLINRDVKSALAVIDDMRAAG 888
>TC231564 similar to GB|AAP31962.1|30387593|BT006618 At3g06430 {Arabidopsis
thaliana;} , partial (40%)
Length = 797
Score = 36.2 bits (82), Expect = 0.034
Identities = 37/146 (25%), Positives = 57/146 (38%), Gaps = 1/146 (0%)
Frame = +1
Query: 166 TEETFNKMRELGFPVTAFACNQLL-LIYKKIDKKKIADVLLMMEKENVKPSSYTYKILID 224
TE+ K R G N L+ K K++ + M K TY +I+
Sbjct: 28 TEKWXKKFRYFGIXXETRTFNILIGXXXXKRMYDKMSSXMEYMRKLQFPWXXSTYNNVIE 207
Query: 225 VKGLSNDIDGMSQIVETMKAEGCELDHLTRASLARHYAAAGLTEKTEAILKEIEGENLKE 284
+ D M + M+AEG + D T L YA AGL K + ++ + E
Sbjct: 208 AFADAGDAKHMECTFDQMRAEGMKADTKTLCCLINGYANAGLFHKVISSVRLAGKLEIPE 387
Query: 285 NMWVCPTLLRLYAILGRADEVERIWK 310
N+ +L A E+ER++K
Sbjct: 388 NITFYNAVLSACAKAEDLMEMERVFK 465
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.133 0.373
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,514,126
Number of Sequences: 63676
Number of extensions: 206814
Number of successful extensions: 1068
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 1035
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1059
length of query: 488
length of database: 12,639,632
effective HSP length: 101
effective length of query: 387
effective length of database: 6,208,356
effective search space: 2402633772
effective search space used: 2402633772
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC135463.1


