

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135416.15 - phase: 0 /pseudo
(662 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
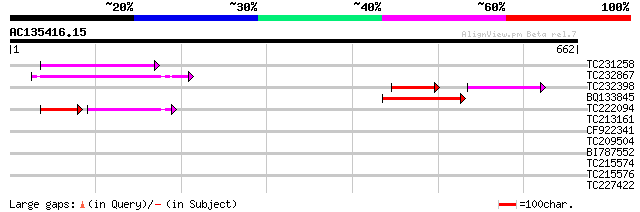
TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kin... 113 2e-25
TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial ... 105 8e-23
TC232398 70 1e-20
BQ133845 82 6e-16
TC222094 54 3e-15
TC213161 33 0.31
CF922341 32 0.91
TC209504 similar to UP|MLO8_ARATH (O22757) MLO-like protein 8 (A... 30 3.4
BI787552 30 4.5
TC215574 similar to UP|O04076 (O04076) ACC-oxidase, partial (98%) 30 4.5
TC215576 similar to UP|O04076 (O04076) ACC-oxidase, partial (97%) 29 5.9
TC227422 similar to UP|P92973 (P92973) CCA1 (MYB-related transcr... 29 5.9
>TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kinase/response
regulator, partial (3%)
Length = 1374
Score = 113 bits (283), Expect = 2e-25
Identities = 58/138 (42%), Positives = 82/138 (59%)
Frame = +2
Query: 37 QEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKTAKLRDQITRFN 96
++ + L FP SL A+ W + L SI +WDD++R FL +FFP S+T +R I+
Sbjct: 641 EDHIFLKAFPHSLEGVAKDWLYYLAPRSIFNWDDLKRVFLEKFFPASRTTTIRKDISGIR 820
Query: 97 QKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKNH 156
Q E LYE WERFK+ CPHH + K L++ FY LS + +DAA+ GAL +
Sbjct: 821 QLSRESLYEYWERFKKSCASCPHHQISKQLLL*YFYEELSNMKRSMIDAASGGALGDMTP 1000
Query: 157 TEAYSLIEDMAQNHYQWT 174
TEA +LI+ MA N Q++
Sbjct: 1001TEARNLIKKMASNSQQFS 1054
>TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial (3%)
Length = 1144
Score = 105 bits (261), Expect = 8e-23
Identities = 67/189 (35%), Positives = 105/189 (55%)
Frame = -3
Query: 26 SLRLSGAIKENQEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKT 85
++RL+G +A+RL L FSL A+ W HS + S+ SWD++ FL ++FP SKT
Sbjct: 719 TIRLAGV---PADAIRLSLLSFSLSGEAKRWLHSFKGNSLKSWDEVVEKFLKKYFPESKT 549
Query: 86 AKLRDQITRFNQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDA 145
A+ + I+ F+Q E L EA ERF+ +LR P HG + + ++ F + L +K VDA
Sbjct: 548 AEGKAAISSFHQFPDESLSEALERFRGLLRKTPTHGFSEPIQLNIFIDELRPESKQLVDA 369
Query: 146 AACGALMNKNHTEAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYEVSEYNHLAAKVE 205
+ G + K EA LIE MA + +R A P+KK + E++ + L A+ +
Sbjct: 368 SVGGKIKMKTPDEAMDLIESMAASDIAILRDR---AHIPTKK--SLLELTSQDTLLAQNK 204
Query: 206 ALTQKFEKL 214
L+++ E L
Sbjct: 203 LLSKQLETL 177
>TC232398
Length = 1054
Score = 70.5 bits (171), Expect(2) = 1e-20
Identities = 37/91 (40%), Positives = 55/91 (59%)
Frame = +1
Query: 535 RLGLGELKLTETTLKLADRSDIQPVGNVEDIPVKIEGIDIPTDFMVLDIDEDNECPIILG 594
R+G +++ T TL+LAD S + G VEDI VK+ + DF+++DI+ED E +ILG
Sbjct: 175 RIGNQKIEPTRMTLQLADHSITRSFGVVEDILVKVHQLIFLVDFVIMDIEEDAEIRLILG 354
Query: 595 RPFLATAGAIVDVQNGRIIFQVSDELIGFEL 625
PF+ TA +VD+ G + V D+ F L
Sbjct: 355 WPFMVTAKCVVDMGKGNLEMSVEDQKATFNL 447
Score = 48.1 bits (113), Expect(2) = 1e-20
Identities = 25/56 (44%), Positives = 34/56 (60%)
Frame = +2
Query: 446 IKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENSAIIKKPPTKLRDPG 501
I IPF E + +MPLY KFLK+I KK ++ET + A+I+K P K +D G
Sbjct: 8 ITIPFGERIQQMPLYKKFLKDILIKKGKYINSETIVVGEYCRALIQKLPPKFKDLG 175
>BQ133845
Length = 389
Score = 82.4 bits (202), Expect = 6e-16
Identities = 43/98 (43%), Positives = 64/98 (64%), Gaps = 1/98 (1%)
Frame = +2
Query: 436 KFLNILQKICIKIPFAEALSRMPLYAKFLKEIFSKKKAIDHNETKALTRENSAIIKK-PP 494
KFL+I +K+ I +PF EAL +MPLYA FLK++ +KK H++ + SA+I++ P
Sbjct: 86 KFLDIFKKLEITLPFEEALQQMPLYANFLKDMLTKKNWYIHSDKIVVEGNCSAVIQRILP 265
Query: 495 TKLRDPGSFAIPCMIGSETLNQALCDLGASVSLLPLPL 532
DPG +PC IG + +AL DLGAS++L+PL +
Sbjct: 266 P*HTDPGFVTMPCSIGEVAVGKALIDLGASINLMPLSM 379
>TC222094
Length = 984
Score = 53.5 bits (127), Expect(2) = 3e-15
Identities = 33/104 (31%), Positives = 56/104 (53%)
Frame = -3
Query: 91 QITRFNQKDGEYLYEAWERFKEMLRLCPHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGA 150
+I+ F+Q E L EA +RF +L P HG + + ++ F +G+ +K +DA+A G
Sbjct: 472 EISSFHQHPHESLSEALDRFHGLLWKTPTHGFSEPVQLNIFIDGMQPHSKQLLDASAGGK 293
Query: 151 LMNKNHTEAYSLIEDMAQNHYQWTNERAVTAPTPSKKEAGIYEV 194
+ K EA LIE+MA N Y ++ P+P ++ + E+
Sbjct: 292 IKLKTPEEAIELIENMAANDYVILRDQ---EPSPQEESTRLEEL 170
Score = 46.6 bits (109), Expect(2) = 3e-15
Identities = 19/49 (38%), Positives = 30/49 (60%)
Frame = -2
Query: 37 QEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARFFPPSKT 85
++A+ L LF FSL AR W S + ++ +W++ FL ++FP SKT
Sbjct: 635 EDAIHLDLFCFSLAGEARTWLRSFKGNNLRTWNEXXEKFLKKYFPESKT 489
>TC213161
Length = 502
Score = 33.5 bits (75), Expect = 0.31
Identities = 18/63 (28%), Positives = 29/63 (45%), Gaps = 4/63 (6%)
Frame = +1
Query: 118 PHHGLEKWLIVHTFYNGLSYTTKMSVDAAACGALMNKNH----TEAYSLIEDMAQNHYQW 173
PH+ L WL V +N ++T +S ACG+ N + E +++D+ W
Sbjct: 109 PHNNLSSWLDVSEDHNQKQFST-LSPTITACGSFSNNGNGRFLREESGVVDDVISPDLAW 285
Query: 174 TNE 176
NE
Sbjct: 286 VNE 294
>CF922341
Length = 675
Score = 32.0 bits (71), Expect = 0.91
Identities = 16/49 (32%), Positives = 28/49 (56%)
Frame = +1
Query: 31 GAIKENQEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDMRRAFLARF 79
GA +++E + +H F SL A W+ +LE + SW D+ AF+ ++
Sbjct: 493 GAYAKDEELL-IHSFQESLTGVAVTWYTNLEPSRVHSWKDLMVAFVRQY 636
>TC209504 similar to UP|MLO8_ARATH (O22757) MLO-like protein 8 (AtMlo8),
partial (24%)
Length = 735
Score = 30.0 bits (66), Expect = 3.4
Identities = 21/58 (36%), Positives = 31/58 (53%), Gaps = 1/58 (1%)
Frame = +2
Query: 457 MPLYAKFLKEIFSKKKAI-DHNETKALTRENSAIIKKPPTKLRDPGSFAIPCMIGSET 513
+PLYA + + KK+I D +KAL + + A+ KK KL G+ + M GS T
Sbjct: 308 LPLYALVTQMVSRMKKSIFDEQTSKALKKWHMAVKKKQGVKL---GNSKVRAMDGSST 472
>BI787552
Length = 421
Score = 29.6 bits (65), Expect = 4.5
Identities = 17/57 (29%), Positives = 27/57 (46%), Gaps = 1/57 (1%)
Frame = +2
Query: 241 CQLGSAANIEQLNYAQFNQGTRPNQKFYKNPQGSYGQVAPPGY-TNNQRVAKKSSLE 296
C + N++ LNY + Q P K +K + V PP Y T + R+A + + E
Sbjct: 170 CCIPVQENLDDLNYLHWRQHVEPVIKLHKLQRFVVNLVVPPRYFTEDDRIADRVNPE 340
>TC215574 similar to UP|O04076 (O04076) ACC-oxidase, partial (98%)
Length = 1169
Score = 29.6 bits (65), Expect = 4.5
Identities = 20/76 (26%), Positives = 36/76 (47%), Gaps = 3/76 (3%)
Frame = +3
Query: 70 DMRRAFLARFFPPSKTAKLRDQITRFNQKDGEYLYEAWERFKEMLRL-CPHHGLEKWLIV 128
D F R P S +++ D + E+ + + +E+L L C + GLEK +
Sbjct: 282 DWESTFFLRHLPTSNISEIPDLSQEYRDAMKEFAQKLEKLAEELLDLLCENLGLEKGYLK 461
Query: 129 HTFY--NGLSYTTKMS 142
+ FY G ++ TK++
Sbjct: 462 NAFYGSRGPNFGTKVA 509
>TC215576 similar to UP|O04076 (O04076) ACC-oxidase, partial (97%)
Length = 1151
Score = 29.3 bits (64), Expect = 5.9
Identities = 20/76 (26%), Positives = 36/76 (47%), Gaps = 3/76 (3%)
Frame = +2
Query: 70 DMRRAFLARFFPPSKTAKLRDQITRFNQKDGEYLYEAWERFKEMLRL-CPHHGLEKWLIV 128
D F R P S +++ D + E+ + + +E+L L C + GLEK +
Sbjct: 233 DWESTFFLRHLPTSNISEIPDLSQEYRDAMKEFAKKLEKLAEELLDLLCENLGLEKGYLK 412
Query: 129 HTFY--NGLSYTTKMS 142
+ FY G ++ TK++
Sbjct: 413 NAFYGSKGPNFGTKVA 460
>TC227422 similar to UP|P92973 (P92973) CCA1 (MYB-related transcription
factor (CCA1); supported by cDNA: gi:1777442) (MYB
transcription factor), partial (11%)
Length = 901
Score = 29.3 bits (64), Expect = 5.9
Identities = 18/68 (26%), Positives = 31/68 (45%), Gaps = 4/68 (5%)
Frame = -2
Query: 36 NQEAVRLHLFPFSLRDRARAWFHSLEVGSITSWDDM----RRAFLARFFPPSKTAKLRDQ 91
N++ +LH FPF L+ H S+ W + + F++ FF P+ +LR +
Sbjct: 360 NKQITKLHTFPFFLK-------H*NPQNSVQKWTSLATKQKHCFVSLFFYPTLLFQLRIE 202
Query: 92 ITRFNQKD 99
T +D
Sbjct: 201 DTLLRSRD 178
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.394
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,891,834
Number of Sequences: 63676
Number of extensions: 364925
Number of successful extensions: 1781
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1759
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1775
length of query: 662
length of database: 12,639,632
effective HSP length: 103
effective length of query: 559
effective length of database: 6,081,004
effective search space: 3399281236
effective search space used: 3399281236
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135416.15


