

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135320.7 + phase: 0
(190 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
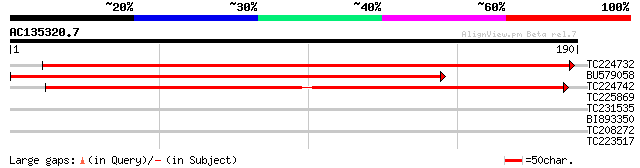
Score E
Sequences producing significant alignments: (bits) Value
TC224732 272 6e-74
BU579058 191 1e-49
TC224742 weakly similar to UP|Q9LJ90 (Q9LJ90) Alanine acetyl tra... 156 6e-39
TC225869 similar to UP|Q84P56 (Q84P56) TGB12K interacting protei... 28 3.3
TC231535 homologue to UP|Q91H08 (Q91H08) Minor structural protei... 27 4.4
BI893350 27 4.4
TC208272 homologue to UP|Q84TL5 (Q84TL5) At1g75760, partial (21%) 27 7.4
TC223517 weakly similar to UP|Q9LUM1 (Q9LUM1) Gb|AAF02862.1, par... 26 9.7
>TC224732
Length = 672
Score = 272 bits (696), Expect = 6e-74
Identities = 129/178 (72%), Positives = 155/178 (86%)
Frame = +3
Query: 12 AKEERIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFL 71
A+EE +DL +I LRPL +SDLDDL++WT+DEKVA FCSW+ Y+SKD+GINFI+NIA+KFL
Sbjct: 90 AEEESVDLGEICLRPLQVSDLDDLLVWTSDEKVAAFCSWDPYSSKDEGINFIQNIASKFL 269
Query: 72 WCKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFC 131
WC+AIC+ DRAIGC+SLSS+S DKSR++ AELGYVLGSKYWGKGVAT VK+VVK A
Sbjct: 270 WCRAICLKDRAIGCISLSSNSEHDKSRSRSAELGYVLGSKYWGKGVATVAVKKVVKAALS 449
Query: 132 ELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTDPQ 189
EL +LER+EALVDV N GSQRVLEKAGFQKEG+LRKY+ KGK RDM+I S+L TDP+
Sbjct: 450 ELPHLERIEALVDVFNVGSQRVLEKAGFQKEGILRKYIFQKGKPRDMVIFSLLSTDPK 623
>BU579058
Length = 444
Score = 191 bits (486), Expect = 1e-49
Identities = 92/146 (63%), Positives = 114/146 (78%)
Frame = +2
Query: 1 MEITSISSKPGAKEERIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGI 60
ME +I S + EE DLTQI+LRP++L DLDDLM+WTTDEKVA++C+WE YTSK+DGI
Sbjct: 5 MEGATICSTTDSIEEGFDLTQISLRPISLDDLDDLMLWTTDEKVARYCTWEPYTSKEDGI 184
Query: 61 NFIENIATKFLWCKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATC 120
NFI+NIA K LW +AIC+N+RAIGC+ S ++R+K ELGY L S YWGKG+AT
Sbjct: 185 NFIQNIAWKSLWFRAICLNNRAIGCIDFFSCEGQRRNRHKSVELGYALASIYWGKGIATH 364
Query: 121 VVKQVVKVAFCELSYLERLEALVDVE 146
VKQV+KVAF E +LERL+ALVDVE
Sbjct: 365 AVKQVIKVAFSEFPHLERLQALVDVE 442
>TC224742 weakly similar to UP|Q9LJ90 (Q9LJ90) Alanine acetyl
transferase-like protein, partial (89%)
Length = 704
Score = 156 bits (394), Expect = 6e-39
Identities = 75/175 (42%), Positives = 117/175 (66%)
Frame = +1
Query: 13 KEERIDLTQITLRPLNLSDLDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFLW 72
K R+DL++I+LRP +SD+DD +IW D++V + W+ S+++ + FI ++ W
Sbjct: 43 KMARVDLSRISLRPFKMSDVDDFLIWAGDDQVTRNLRWKTCGSREEALAFIRDVCIPHPW 222
Query: 73 CKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFCE 132
++IC++DR+IG VS+ S ++ + A++GY +G+ YWG+G+AT + V F +
Sbjct: 223 RRSICLDDRSIGFVSVYPWSGDERCK---ADIGYAIGTNYWGQGIATKALMTAVPQVFKD 393
Query: 133 LSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMIISSVLFTD 187
+ L RL+A VDVEN SQRVLEKAGF +EGVLRKY +KG +D+++ S L TD
Sbjct: 394 FNELLRLQAFVDVENKASQRVLEKAGFLREGVLRKYTYLKGVVKDLVLYSFLSTD 558
>TC225869 similar to UP|Q84P56 (Q84P56) TGB12K interacting protein 2, partial
(78%)
Length = 1445
Score = 27.7 bits (60), Expect = 3.3
Identities = 32/114 (28%), Positives = 48/114 (42%), Gaps = 7/114 (6%)
Frame = -1
Query: 73 CKAICINDRAIGCVSLSSSSPGDKSRNKCAELGYVLGSKYWGKGVATCVVKQVVKVAFC- 131
C AI +ND + S +S G SR AE L GK VA +V VA C
Sbjct: 383 CSAIWLNDGSFAICSANSLILGSLSRPDIAEKSKGLP----GKPVAG---PEVAVVALCA 225
Query: 132 ------ELSYLERLEALVDVENAGSQRVLEKAGFQKEGVLRKYLVMKGKSRDMI 179
++S L+ L + V ++ + LE ++E RK L + + R +
Sbjct: 224 AGESLDDVSSLDALFSAVPALSSAGKSFLESEAMREEN--RKLLNLSERGRGSV 69
>TC231535 homologue to UP|Q91H08 (Q91H08) Minor structural protein, partial
(6%)
Length = 1441
Score = 27.3 bits (59), Expect = 4.4
Identities = 15/52 (28%), Positives = 26/52 (49%), Gaps = 8/52 (15%)
Frame = +1
Query: 17 IDLTQITLRPLNLSDLDDLM--------IWTTDEKVAKFCSWELYTSKDDGI 60
IDL Q R +N +++D + IW + ++ K C W+ T K++ I
Sbjct: 1159 IDLKQ*NNRLINTTEVDGTIEKH*AMEPIWCSGDEATKTCVWDTNT*KNNSI 1314
>BI893350
Length = 423
Score = 27.3 bits (59), Expect = 4.4
Identities = 12/31 (38%), Positives = 16/31 (50%)
Frame = +1
Query: 47 FCSWELYTSKDDGINFIENIATKFLWCKAIC 77
FC L T + DGI+ EN A + W +C
Sbjct: 1 FCK*RLSTLQADGISMFENSAMRSHWSLELC 93
>TC208272 homologue to UP|Q84TL5 (Q84TL5) At1g75760, partial (21%)
Length = 463
Score = 26.6 bits (57), Expect = 7.4
Identities = 14/63 (22%), Positives = 28/63 (44%), Gaps = 2/63 (3%)
Frame = +1
Query: 32 LDDLMIWTTDEKVAKFCSWELYTSKDDGINFIENIATKFL--WCKAICINDRAIGCVSLS 89
+D ++W +K+ K W+++ + + +N+ FL WC D + C +
Sbjct: 67 MDCGLLWFFYQKLCKLSFWQIFATTTSRVLLGDNLFFDFLQEWCNLTFK*DVPLTCSTQG 246
Query: 90 SSS 92
S S
Sbjct: 247 SQS 255
>TC223517 weakly similar to UP|Q9LUM1 (Q9LUM1) Gb|AAF02862.1, partial (25%)
Length = 445
Score = 26.2 bits (56), Expect = 9.7
Identities = 8/22 (36%), Positives = 14/22 (63%)
Frame = -2
Query: 59 GINFIENIATKFLWCKAICIND 80
G+ I+ + T LWC+ +C+ D
Sbjct: 297 GLPSIQELVTSSLWCRELCLVD 232
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,317,147
Number of Sequences: 63676
Number of extensions: 114093
Number of successful extensions: 499
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 497
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 498
length of query: 190
length of database: 12,639,632
effective HSP length: 92
effective length of query: 98
effective length of database: 6,781,440
effective search space: 664581120
effective search space used: 664581120
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC135320.7


