

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
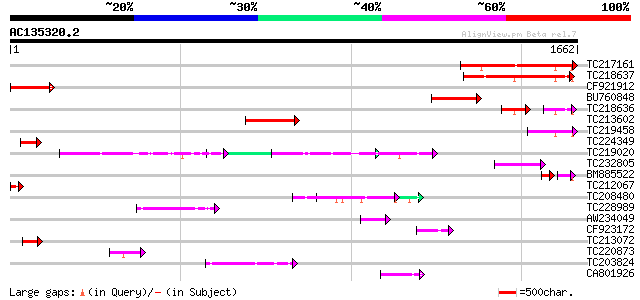
Query= AC135320.2 + phase: 0
(1662 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC217161 351 2e-96
TC218637 similar to UP|Q9XEY9 (Q9XEY9) NT3, partial (6%) 306 6e-83
CF921912 196 5e-50
BU760848 167 5e-41
TC218636 similar to UP|Q9XEY9 (Q9XEY9) NT3, partial (6%) 91 6e-36
TC213602 weakly similar to UP|Q9LUI2 (Q9LUI2) Centromere protein... 121 2e-27
TC219458 similar to UP|Q7X9F2 (Q7X9F2) Centromere protein (Fragm... 88 3e-17
TC224349 similar to UP|Q9LR76 (Q9LR76) F21B7.9, partial (20%) 86 1e-16
TC219020 similar to UP|SRB7_SCHPO (O94376) RNA polymerase II med... 76 1e-13
TC232805 similar to UP|Q7X9F2 (Q7X9F2) Centromere protein (Fragm... 70 8e-12
BM885522 similar to GP|4894628|gb|A NT3 {Nicotiana tabacum}, par... 42 2e-09
TC212067 62 2e-09
TC208480 59 2e-08
TC228989 similar to UP|Q9LSS5 (Q9LSS5) Myosin heavy chain-like, ... 57 5e-08
AW234049 55 3e-07
CF923172 54 7e-07
TC213072 53 1e-06
TC220873 homologue to UP|Q9SFF4 (Q9SFF4) T26F17.2, partial (5%) 45 2e-04
TC203824 similar to UP|Q9V3T8 (Q9V3T8) SR family splicing factor... 45 3e-04
CA801926 similar to GP|9828634|gb F1N21.5 {Arabidopsis thaliana}... 44 8e-04
>TC217161
Length = 1526
Score = 351 bits (900), Expect = 2e-96
Identities = 208/391 (53%), Positives = 268/391 (68%), Gaps = 49/391 (12%)
Frame = +1
Query: 1321 KDVVIETCLRKH-SHQSARENEIVLIPDGVSDLLTLQTRIRAVEKIMMEELKRRVKQKSL 1379
+DV IETCL++ S +S + +E L PDGV+DLL++QTRIR VEK MMEEL+RRVK++SL
Sbjct: 49 QDVAIETCLQEELSQKSLKGSESSLTPDGVADLLSMQTRIRVVEKFMMEELERRVKKESL 228
Query: 1380 T--------TESTPYSSLEIATYPKVENRKKEIELVEENVF-DRNSWRKKPKIRLLMKDI 1430
T TE +S+LE+ TYP++++RK +++ ++N N+WR K + RL+M DI
Sbjct: 229 TANVKAEAVTEMNEHSNLEVGTYPEIDDRKVVMKIKKDNSKRGHNAWRTKSQKRLIMIDI 408
Query: 1431 PLDRNVDDQN-SKYLKREHRRTNDHVLELCENNEHEPL-----SAPTVDHAMICHRSDDS 1484
PLD DD + +K KR+H R ++H+LELCE ++H+ ++ +++ + CH S+
Sbjct: 409 PLDDYKDDPDFNKDGKRDHTRIDNHMLELCETDQHDVTEENKQNSVSLEDVITCHESE-- 582
Query: 1485 GRYLNYSSELDIEKELGVDKLELSKSVKEKT-EDDKRRILERLSSDGQKLAILKMALQDL 1543
R NYSSEL+ EKELGVDKLEL K+ KE T ED KR+ILERL+SD Q+LAILKM LQDL
Sbjct: 583 -RCQNYSSELETEKELGVDKLELWKTRKETTSEDSKRKILERLASDSQRLAILKMTLQDL 759
Query: 1544 KKKTETKKKSKQGNDIEYETVKRHIEEVEEAVMQQVSINDQMAKNVEEGASSLD------ 1597
KKK ET+KKS + N+IEYETVKRHIE+VEEAVM+Q+ I DQ+AK+ EE SS
Sbjct: 760 KKKPETQKKSSKVNEIEYETVKRHIEDVEEAVMKQIGIYDQLAKDTEECTSSSSDTSTMQ 939
Query: 1598 -------------REIPRRGSEQIGKLQFEVQNIQYILLKLAEENNNKVKNRISRKTGIL 1644
E RRGSEQIG+LQFEVQNIQYILLKLA+ NNK KN+ SR TG+L
Sbjct: 940 LEKQGGQTQRKKLTEQARRGSEQIGRLQFEVQNIQYILLKLADVKNNKCKNKNSRPTGVL 1119
Query: 1645 L-------------RRKLRVCGCSGPSTNED 1662
L RRK VCGCS PSTNED
Sbjct: 1120 LKDFIRIGRKNSRRRRKGCVCGCSRPSTNED 1212
>TC218637 similar to UP|Q9XEY9 (Q9XEY9) NT3, partial (6%)
Length = 1103
Score = 306 bits (783), Expect = 6e-83
Identities = 184/370 (49%), Positives = 245/370 (65%), Gaps = 43/370 (11%)
Frame = +2
Query: 1329 LRKHSHQSARENEIVLIPDGVSDLLTLQTRIRAVEKIMMEELKRRVKQKSLTTESTPYSS 1388
L+ + +QS+ +N+ LIPDGVSDLL+++ RIRAVEK M+EE++R VK+++LTT + +
Sbjct: 8 LQGNGYQSSTDNKSALIPDGVSDLLSVKARIRAVEKSMVEEIERHVKEQNLTTTANLGAL 187
Query: 1389 LEIATYPKVENRKKEIELVEENVFDRNSWRKKPKIRLLMKDIPLDRNVDDQNSKYLKREH 1448
++ P VENR ++ EL +E+ D NSWR + + LMKDIPLD D+ SK +RE+
Sbjct: 188 TKV---PNVENRNRK-ELKDESTHDVNSWRTRTENGSLMKDIPLDHISDNSASKSGRREN 355
Query: 1449 RRTNDHVLELCENNEHEPLSAPTVDHAM-----------ICHRSDDSGRYLNYSSELDIE 1497
+D +LEL E E + +P V AM H+SD SG++ N SSELD+E
Sbjct: 356 SGADDQMLELWETAEQDCFDSPMVSEAMKQSSVPTEDVITYHQSDHSGKFQNTSSELDVE 535
Query: 1498 KELGVDKLELSKSVKEKTEDDKRR-ILERLSSDGQKLAILKMALQDLKKKTETKKKSKQG 1556
KELGVD+L+LS+S+KE+T+D KRR ILERLSSD QKL ILK A+QDLK+KTETKK+SK+G
Sbjct: 536 KELGVDRLQLSRSIKERTQDGKRRKILERLSSDAQKLTILKTAVQDLKQKTETKKRSKKG 715
Query: 1557 NDIEYETVKRHIEEVEEAVMQQVSINDQMAKNVEEGASSLDR------------------ 1598
+ EYETVKR I+EVE AV++ V NDQ+ K++EE A SL+R
Sbjct: 716 VETEYETVKRQIDEVEGAVVKLVDTNDQLTKDLEESAPSLNRQTSAELEKSRHIQRKRIT 895
Query: 1599 EIPRRGSEQIGKLQFEVQNIQYILLKLAEENNNKVKNRISRKTGILLR------------ 1646
E R+GSEQIG+LQFEVQNIQY LLKLA+E +K K+R + KT +LLR
Sbjct: 896 EQARKGSEQIGRLQFEVQNIQYTLLKLADE--SKGKSRFTGKTVVLLRDFIHSGSKRTSK 1069
Query: 1647 -RKLRVCGCS 1655
R CGCS
Sbjct: 1070 KRNKGFCGCS 1099
Score = 37.0 bits (84), Expect = 0.072
Identities = 67/320 (20%), Positives = 138/320 (42%), Gaps = 30/320 (9%)
Frame = +2
Query: 238 LESMLSQKDNIEA--KNLKQELTRVVVQKDTVLLQYKQCLEKIPMLE--NKIALAEENSR 293
+ +LS K I A K++ +E+ R V +++ L K+P +E N+ L +E++
Sbjct: 68 VSDLLSVKARIRAVEKSMVEEIERHVKEQNLTTTANLGALTKVPNVENRNRKELKDESTH 247
Query: 294 MLNDQIERTE-------LEVETLRKNLAEMNEERDSLSVLYHHCLEKISKMENEILHVQE 346
+N RTE + ++ + N A + R++ S ++++L + E
Sbjct: 248 DVNSWRTRTENGSLMKDIPLDHISDNSASKSGRREN------------SGADDQMLELWE 391
Query: 347 NAEQ--LKNKIEKEAEKLEI----------SEKHRGMLEKSNQNLQLEAENLVQRIASKD 394
AEQ + + EA K H G + ++ L +E E V R+
Sbjct: 392 TAEQDCFDSPMVSEAMKQSSVPTEDVITYHQSDHSGKFQNTSSELDVEKELGVDRLQLS- 568
Query: 395 HELLEKHTEIERLQTLMHGEHSNFIQIESALQALQKLYSQSQKEQRN-LALELKYGLLLL 453
+++ T+ + + ++ S+ ++ A+Q L +++ ++R+ +E +Y
Sbjct: 569 -RSIKERTQDGKRRKILERLSSDAQKLTILKTAVQDLKQKTETKKRSKKGVETEY----- 730
Query: 454 KDLELSKQDFKEEMQG----IVEENKTLHELNFSSTRSLKKQ-QMEISKLKEIKEK-LER 507
E K+ +E++G +V+ N L + S SL +Q E+ K + I+ K +
Sbjct: 731 ---ETVKRQI-DEVEGAVVKLVDTNDQLTKDLEESAPSLNRQTSAELEKSRHIQRKRITE 898
Query: 508 EFHTSTEESNVLQRETHQIK 527
+ +E+ LQ E I+
Sbjct: 899 QARKGSEQIGRLQFEVQNIQ 958
>CF921912
Length = 715
Score = 196 bits (499), Expect = 5e-50
Identities = 97/129 (75%), Positives = 106/129 (81%)
Frame = +3
Query: 1 MATLSESESRRLYSWWWDSHNSPKNSKWLLENLTDIDTKVKSMIKLIEEEADSFARRAEM 60
MATLS ++SRR+YSWWWDSH SPKNSKWL ENLTD+D+KV+ MIKLIEE+ADSFARRAEM
Sbjct: 216 MATLSHADSRRMYSWWWDSHISPKNSKWLQENLTDMDSKVRQMIKLIEEDADSFARRAEM 395
Query: 61 YYKKRPELMKLVEEFYRAYRALAERYDHAMGELRHAHKTMPEAFPNSAYYILNDDSPCGS 120
YYKKRPELMKLVEEFYRAYRALAERYDHA G +R AH TM EAFPN DDSP S
Sbjct: 396 YYKKRPELMKLVEEFYRAYRALAERYDHATGVIRQAHHTMAEAFPNQGPPAPADDSPVVS 575
Query: 121 LGPDAESHT 129
+ E HT
Sbjct: 576 -SMETEPHT 599
>BU760848
Length = 451
Score = 167 bits (422), Expect = 5e-41
Identities = 91/146 (62%), Positives = 111/146 (75%), Gaps = 1/146 (0%)
Frame = +3
Query: 1237 LLESKVSELTGVCKILDDESSAKSLENEHMREIISLLESEIGGLKEQLSAYVPLVSSLKE 1296
LL+ KV+ELTGVCK LDDES+ K L E M E I LLE EI GLK QLSAY P ++SLKE
Sbjct: 3 LLKGKVTELTGVCKRLDDESAGKGLVIEQMIERIGLLEKEIRGLKGQLSAYTPTITSLKE 182
Query: 1297 DFNSLEHISLLWTKRNSVVGNGAQKDVVIETCLRK-HSHQSARENEIVLIPDGVSDLLTL 1355
DF SLEH L T + VGN QKDVV E CL++ +S++S + NE L PDGV+DLL++
Sbjct: 183 DFASLEHTYFLLTNKTFAVGNIEQKDVVTEICLQEANSYRSLKGNESTLTPDGVTDLLSM 362
Query: 1356 QTRIRAVEKIMMEELKRRVKQKSLTT 1381
QTRIRAVEK+MMEEL+R V+++SLTT
Sbjct: 363 QTRIRAVEKLMMEELERHVEEESLTT 440
>TC218636 similar to UP|Q9XEY9 (Q9XEY9) NT3, partial (6%)
Length = 919
Score = 90.9 bits (224), Expect(2) = 6e-36
Identities = 59/128 (46%), Positives = 75/128 (58%), Gaps = 31/128 (24%)
Frame = +2
Query: 1565 KRHIEEVEEAVMQQVSINDQMAKNVEEGASSLDR------------------EIPRRGSE 1606
+R I+EVE AV++ V NDQ+ K++EE A SL+R E R+GSE
Sbjct: 275 ERQIDEVEGAVVKLVDTNDQLTKDLEESAPSLNRQTSVELEKSRHIQRKRITEQARKGSE 454
Query: 1607 QIGKLQFEVQNIQYILLKLAEENNNKVKNRISRKTGILLR-------------RKLRVCG 1653
QIG+LQFEVQNIQY LLKLA+E +K K+R + KT +LLR R CG
Sbjct: 455 QIGRLQFEVQNIQYTLLKLADE--SKGKSRFTGKTVVLLRDFIHSGSKRTSKKRNKGFCG 628
Query: 1654 CSGPSTNE 1661
CS PST+E
Sbjct: 629 CSRPSTDE 652
Score = 80.5 bits (197), Expect(2) = 6e-36
Identities = 45/98 (45%), Positives = 61/98 (61%), Gaps = 11/98 (11%)
Frame = +1
Query: 1441 SKYLKREHRRTNDHVLELCENNEHEPLSAPTVDHAM-----------ICHRSDDSGRYLN 1489
SK +RE+ +D +LEL E E + +P V AM H+SD SG++ N
Sbjct: 1 SKSGRRENSGADDQMLELWETAEQDCFDSPMVSEAMKQSSVPTEDVITYHQSDHSGKFQN 180
Query: 1490 YSSELDIEKELGVDKLELSKSVKEKTEDDKRRILERLS 1527
SSELD+EKELGVD+L+LS+S+KE+T+D KRR R S
Sbjct: 181 TSSELDVEKELGVDRLQLSRSIKERTQDGKRRKTNRRS 294
>TC213602 weakly similar to UP|Q9LUI2 (Q9LUI2) Centromere protein, partial
(3%)
Length = 597
Score = 121 bits (304), Expect = 2e-27
Identities = 64/161 (39%), Positives = 107/161 (65%)
Frame = +3
Query: 690 LNDEKNNLQNERSMLISQLEIVEEKLSNLEKKVTNLEEKYADVEKDKESAVNQVEELFAS 749
L+ EK+++ E+ L+SQL I + L +LE+ + LE K+ + + ++ESA+ +VEEL S
Sbjct: 18 LDHEKSSIFQEKETLVSQLNITHQTLKDLEELHSLLELKHLEQKGERESALQKVEELLVS 197
Query: 750 ILVQKENHSNHKHSSEARLANLENIVRVLQEEQRLGKVEFEQELDRVVNAQIEMFILQNC 809
+ ++E +S +E LA E + +LQE+ K E+E+ELDR ++A +++FILQ C
Sbjct: 198 LYSEREENSRVLKLNEDELAEKELQIHILQEDANCKKKEYEEELDRAIHAHVQIFILQKC 377
Query: 810 IEELELKNFVLLTECEKLVEASKFSDKVISELESENLMQLI 850
+++LE KNF LL E ++L+EASK S K+I +L ++QL+
Sbjct: 378 VDDLEKKNFSLLVEWQRLLEASKMSYKMICKL**RLVLQLV 500
Score = 32.7 bits (73), Expect = 1.4
Identities = 36/150 (24%), Positives = 65/150 (43%), Gaps = 5/150 (3%)
Frame = +3
Query: 849 LIEEEFLLHRIRKFKMDIHKVCGVLQIDSDGGGDNEIKKEEIPISRILDKIESLESSLVK 908
L+ + + H+ K ++H + + ++ G ++ L K+E L SL
Sbjct: 60 LVSQLNITHQTLKDLEELHSLLELKHLEQKGERESA-----------LQKVEELLVSLYS 206
Query: 909 SQEENQQLLV--ENSVLLGSLQQHQSEGEKLKLEKKTVEQEFENMREQNV---ILQKDKV 963
+EEN ++L E+ + LQ H + E +KK E+E + +V ILQK
Sbjct: 207 EREENSRVLKLNEDELAEKELQIHILQ-EDANCKKKEYEEELDRAIHAHVQIFILQKCVD 383
Query: 964 ELLEENRQLRIEVVNGVEKENRSKSTLAAL 993
+L ++N L +E +E S + L
Sbjct: 384 DLEKKNFSLLVEWQRLLEASKMSYKMICKL 473
>TC219458 similar to UP|Q7X9F2 (Q7X9F2) Centromere protein (Fragment), partial
(91%)
Length = 816
Score = 88.2 bits (217), Expect = 3e-17
Identities = 59/179 (32%), Positives = 86/179 (47%), Gaps = 34/179 (18%)
Frame = +2
Query: 1518 DKRRILERLSSDGQKLAILKMALQDLKKKTETKKKSKQGNDIEYETVKRHIEEVEEAVMQ 1577
+ ++ ERL D QKL L++ +Q KK E +K +G +E+ VK +E +E + +
Sbjct: 62 EPKQXXERLDXDAQKLTXLQITIQXXMKKVEINEKXSKGKSVEFGEVKGQLEAAQENITK 241
Query: 1578 QVSINDQMAKNVEEG-ASSLDR-----------------EIPRRGSEQIGKLQFEVQNIQ 1619
N ++ KNVEEG SS+ + E RR SE+IG+L EVQ +Q
Sbjct: 242 LFDANRKLMKNVEEGTVSSVGKDAAELGEIGSVSRRRVSEQARRESEKIGQLHLEVQRLQ 421
Query: 1620 YILLKLAEENNNKVKNRIS-RKTGILLR---------------RKLRVCGCSGPSTNED 1662
++LLKL E NK K + + R +LLR +KL C C P T D
Sbjct: 422 FLLLKLGEGKENKEKTKTADRSPRVLLRDYLYGGTRTNNQKKKKKLPFCSCVRPPTKGD 598
>TC224349 similar to UP|Q9LR76 (Q9LR76) F21B7.9, partial (20%)
Length = 652
Score = 85.9 bits (211), Expect = 1e-16
Identities = 37/59 (62%), Positives = 51/59 (85%)
Frame = +1
Query: 33 LTDIDTKVKSMIKLIEEEADSFARRAEMYYKKRPELMKLVEEFYRAYRALAERYDHAMG 91
+ +++ K K+M+KLIEE+ADSFA+RAEMYYKKRP+L+ +VE+FYR +R+LAERYD G
Sbjct: 133 MAELNEKTKAMLKLIEEDADSFAQRAEMYYKKRPQLVSMVEDFYRTHRSLAERYDQVTG 309
>TC219020 similar to UP|SRB7_SCHPO (O94376) RNA polymerase II mediator
complex protein srb7, partial (14%)
Length = 1446
Score = 76.3 bits (186), Expect = 1e-13
Identities = 113/515 (21%), Positives = 219/515 (41%), Gaps = 18/515 (3%)
Frame = +2
Query: 145 EESNGEVQTLREALAKMQSDKDALFLQYQESLENLSKMETDLNKAQNNARGLDDRASEAE 204
E +Q L + ++ ++ + A Q + L+++S + ++L Q A L+ A
Sbjct: 5 EAEKYRIQELEQQISTLEEKRGASEGQANKYLDDVSNLTSELEAIQARASTLETTLQAAN 184
Query: 205 IQVEILKESLMQLKADKDAGEVLYNQCLETIARLESMLS-QKDNIEAKNLKQELTRVVVQ 263
+ + L++SL + +K E E +A E++L +D++ K + T ++
Sbjct: 185 ERGKELEDSLNAVTEEKKNLEDASISLNEKLAEKENLLEILRDDLNLTQDKLQSTESDLR 364
Query: 264 KDTVLLQYKQCLEKIPMLENKIALAEENSRMLNDQIERTELEVETLRKN----LAEMNEE 319
+ L+ + +EK+ E + + + + +L E+L ++ L E E+
Sbjct: 365 EAE--LRESEIIEKLKASEENLVVRGRDIEETATRHSELQLLHESLTRDSEQKLQEAIEK 538
Query: 320 RDSLSVLYHHCLEKISKMENEILHVQENAEQLKNKIEKEAEKLEISEKHRGMLEKSNQNL 379
++ LEKI +E +I E + LKN+ E+ KL +L
Sbjct: 539 FNNKDSEVQSLLEKIKILEEQIAKAGEQSTSLKNEFEESLSKL--------------TSL 676
Query: 380 QLEAENLVQRIASKDHELLEKHTEIERLQTLMHGEHSNFIQIESALQALQKLYSQSQKEQ 439
+ E E+L ++I + + + +E E L+ G + IQ+++ + L++ + + E+
Sbjct: 677 ESENEDLKRQILDAESKSSQSFSENE----LLVGTN---IQLKTKIDELEESLNHALSEK 835
Query: 440 RNLALEL---KYGLLLLKDLELSKQDFKEEMQGIVEENKTLHELNFSSTRSLKKQQMEIS 496
A EL K + L DL+ + + E +TL ++ +L++ + S
Sbjct: 836 EAAAQELVSHKNSITELNDLQSKSSEIQR-----ANEARTL-KVESQLQEALQRHTEKES 997
Query: 497 KLKEIKEKL----------EREFHTSTEESNVLQRETHQIKDDIQHLNERYQAMLEQLQS 546
+ KE+ EKL E + S + E Q ++HL + ++E+LQ+
Sbjct: 998 ETKELNEKLNTLEGQIKLFEEHAREAVATSGTHKAELEQSLIKLKHL----EIVIEELQN 1165
Query: 547 LGLNPNSFAASVRDLQNENFMLKETCKKEHSEKEALREKSKDMNEVLMENACMEFSLLGL 606
L+ A L EN L + S+ L+EK ++ L+E + LL L
Sbjct: 1166KSLHHEKETAG---LNEENSKLNQEIASYESKLSDLQEK---LSAALVEKEETDKELLTL 1327
Query: 607 NDELDGLRGTVKEIQQFCQVLQEEKSILADEKSTL 641
D ++ L GT + Q L + S L DEK+ L
Sbjct: 1328KDAMEKL-GTKHSAE--VQTLNSQISSLVDEKNLL 1423
Score = 71.2 bits (173), Expect = 3e-12
Identities = 100/514 (19%), Positives = 203/514 (39%), Gaps = 3/514 (0%)
Frame = +2
Query: 577 SEKEALREKSKDMNEVLMENACMEFSLLGLNDELDGLRGTVKEIQQFCQVLQEEKSILAD 636
+EK ++E + ++ + + E D++ L ++ IQ L+ +
Sbjct: 8 AEKYRIQELEQQISTLEEKRGASEGQANKYLDDVSNLTSELEAIQARASTLETTLQAANE 187
Query: 637 EKSTLLSQLQIITESMQKILENNTVLEKSLSDAKIEFEGLRIKSGDLEDCCKLLNDEKNN 696
L L +TE + + + + L + L++ + E LR +D + +
Sbjct: 188 RGKELEDSLNAVTEEKKNLEDASISLNEKLAEKENLLEILRDDLNLTQDKLQSTESDLRE 367
Query: 697 LQNERSMLISQLEIVEEKLSNLEKKVTNLEEKYADVEKDKESAVNQVEELFASILVQKEN 756
+ S +I +L+ EE L + + ++++++ ES E+ + + N
Sbjct: 368 AELRESEIIEKLKASEENLVVRGRDIEETATRHSELQLLHESLTRDSEQKLQEAIEKFNN 547
Query: 757 HSNHKHSSEARLANLENIVRVLQEEQRLGKVEFEQELDRVVNAQIEMFILQNCIEELELK 816
+ S ++ LE + E+ K EFE+ L ++ + + E L+ I + E K
Sbjct: 548 KDSEVQSLLEKIKILEEQIAKAGEQSTSLKNEFEESLSKLTSLESENEDLKRQILDAESK 727
Query: 817 NFVLLTECEKLVEASKFSDKVISELESENLMQLIEEEFLLHRIRKFKMDIHKVCGVLQID 876
+ +E E LV + I ELE L E+E + K I ++
Sbjct: 728 SSQSFSENELLVGTNIQLKTKIDELEESLNHALSEKEAAAQELVSHKNSITEL------- 886
Query: 877 SDGGGDNEIKKEEIPISRILDKIESLESSLVKSQEENQQLLVENSVLLGSLQQHQSEGEK 936
N+++ + SS ++ E + L VE S L +LQ+H +
Sbjct: 887 ------NDLQSK---------------SSEIQRANEARTLKVE-SQLQEALQRHTEK--- 991
Query: 937 LKLEKKTVEQEFENMREQNVILQKDKVELLEENRQLRIEVVNGVEKENRSKSTLAALQAE 996
+ E K + ++ + Q + ++ E + + + E+ + K + + LQ +
Sbjct: 992 -ESETKELNEKLNTLEGQIKLFEEHAREAVATSGTHKAELEQSLIKLKHLEIVIEELQNK 1168
Query: 997 MIELRQTNQVFQEENGKMLDEKNSLCRNVSDLKDAKSSAEDENSVMFHDVLALSNLNLVY 1056
+ + EEN K+ E S +SDL++ S+A E ++L L +
Sbjct: 1169 SLHHEKETAGLNEENSKLNQEIASYESKLSDLQEKLSAALVEKEETDKELLTLKD---AM 1339
Query: 1057 EIFFTENMVEKRALCEHLGNL---SHLNNDLNQE 1087
E T++ E + L + +L +L ND NQ+
Sbjct: 1340 EKLGTKHSAEVQTLNSQISSLVDEKNLLNDTNQD 1441
Score = 67.8 bits (164), Expect = 4e-11
Identities = 110/508 (21%), Positives = 211/508 (40%), Gaps = 20/508 (3%)
Frame = +2
Query: 767 RLANLENIVRVLQEEQRLGKVEFEQELDRVVNAQIEMFILQNCIEELELKNFVLLTECEK 826
R+ LE + L+E++ + + + LD V N E+ +Q LE ++
Sbjct: 20 RIQELEQQISTLEEKRGASEGQANKYLDDVSNLTSELEAIQARASTLETTLQAANERGKE 199
Query: 827 LVEASKFSDKVISELESENLM---QLIEEEFLLHRIRKFKMDIHKVCGVLQIDSDGGGDN 883
L ++ + LE ++ +L E+E LL +R D++ LQ ++
Sbjct: 200 LEDSLNAVTEEKKNLEDASISLNEKLAEKENLLEILRD---DLNLTQDKLQ-----STES 355
Query: 884 EIKKEEIPISRILDKIESLESSLVKSQEENQQLLVENSVLLGSLQQHQSEGEKLKLEKKT 943
++++ E+ S I++K+++ E +LV + ++ +S L L + + + KL++
Sbjct: 356 DLREAELRESEIIEKLKASEENLVVRGRDIEETATRHSELQ-LLHESLTRDSEQKLQEAI 532
Query: 944 VEQEFENMREQNVILQKDKVELLEENRQLRIEVVNGVEKE-NRSKSTLAALQAEMIELRQ 1002
++F N ++ V +K+++LEE E ++ E S S L +L++E +L+
Sbjct: 533 --EKFNN-KDSEVQSLLEKIKILEEQIAKAGEQSTSLKNEFEESLSKLTSLESENEDLK- 700
Query: 1003 TNQVFQEENGKMLDEKNSLCRNVSDLKDAKSSAEDENSVMFHDVLALSNLNLVYEIFFTE 1062
R + D + S + EN ++ + L E
Sbjct: 701 --------------------RQILDAESKSSQSFSENELLVGTNIQLKTKIDELEESLNH 820
Query: 1063 NMVEKRALCEHLGNLSHLN-----NDLNQEFGVLRKNFEVKEAE-NVYLNESIER---MD 1113
+ EK A + L +SH N NDL + +++ E + + L E+++R +
Sbjct: 821 ALSEKEAAAQEL--VSHKNSITELNDLQSKSSEIQRANEARTLKVESQLQEALQRHTEKE 994
Query: 1114 KELLEMDKRLKAAETSNAEFSRHIEEL-------KMEQEESTKIKENLDRQILEQSENCM 1166
E E++++L E F H E K E E+S ++L+ I E +
Sbjct: 995 SETKELNEKLNTLEGQIKLFEEHAREAVATSGTHKAELEQSLIKLKHLEIVIEELQNKSL 1174
Query: 1167 NHKKEIEHLNEANETLQFEMKTLLHEVEQHRVREEALNLELLNKENEFKLWENEAAAFYH 1226
+H+KE LNE N L E+ + ++ +E L+ L+ KE K A
Sbjct: 1175 HHEKETAGLNEENSKLNQEIASYESKLSD---LQEKLSAALVEKEETDKELLTLKDAMEK 1345
Query: 1227 DLQMSSICLALLESKVSELTGVCKILDD 1254
S + L S++S L +L+D
Sbjct: 1346 LGTKHSAEVQTLNSQISSLVDEKNLLND 1429
>TC232805 similar to UP|Q7X9F2 (Q7X9F2) Centromere protein (Fragment), partial
(66%)
Length = 502
Score = 70.1 bits (170), Expect = 8e-12
Identities = 53/157 (33%), Positives = 85/157 (53%), Gaps = 9/157 (5%)
Frame = +3
Query: 1422 KIRLLMKDIPLDRNVDDQNSKYL--KREHRRTNDHVLELCENNEHEPL----SAPTVDHA 1475
+I +L KDI LD+ + + +S + +RE +D +LEL E + + + T A
Sbjct: 30 EIEVLPKDIMLDQ-ISECSSYGISRRREILEADDQMLELWETADKDATIGKQAEKTQKMA 206
Query: 1476 MICHRSDDSGRYLNYSSELD--IEKELGVDKLELSKSVK-EKTEDDKRRILERLSSDGQK 1532
H+ + N D +EKEL VDKLE+S+ + + E ++ +ILERL SD QK
Sbjct: 207 AGNHQRGTTKEPKNRYPSTDSLVEKELSVDKLEVSRRLTLPREEGNQSKILERLDSDAQK 386
Query: 1533 LAILKMALQDLKKKTETKKKSKQGNDIEYETVKRHIE 1569
L L++ +QDL K + +KS +G +E+ VK +E
Sbjct: 387 LTNLQITIQDLMNKVQINEKSTKGKSVEFGDVKGQLE 497
>BM885522 similar to GP|4894628|gb|A NT3 {Nicotiana tabacum}, partial (4%)
Length = 421
Score = 41.6 bits (96), Expect(2) = 2e-09
Identities = 26/72 (36%), Positives = 36/72 (49%), Gaps = 17/72 (23%)
Frame = +3
Query: 1605 SEQIGKLQFEVQNIQYILLKLAEENNNKVKNRI-SRKTGILLR----------------R 1647
SE+IG+LQ EVQ +Q++LLKL +E K K + R + +LLR +
Sbjct: 201 SEKIGRLQLEVQRLQFLLLKLNDEKEGKGKAMMDERNSKVLLRDYLYAGGTRRNYQKRKK 380
Query: 1648 KLRVCGCSGPST 1659
K C C P T
Sbjct: 381 KTHFCACMQPPT 416
Score = 40.4 bits (93), Expect(2) = 2e-09
Identities = 18/38 (47%), Positives = 25/38 (65%)
Frame = +2
Query: 1558 DIEYETVKRHIEEVEEAVMQQVSINDQMAKNVEEGASS 1595
D EY+TVK +E +EA+ + N ++ KNVEEG SS
Sbjct: 5 DSEYDTVKGQLEATQEAITKLFDANQKLKKNVEEGTSS 118
>TC212067
Length = 623
Score = 62.0 bits (149), Expect = 2e-09
Identities = 29/40 (72%), Positives = 30/40 (74%)
Frame = +3
Query: 1 MATLSESESRRLYSWWWDSHNSPKNSKWLLENLTDIDTKV 40
MATLS SESRR YSWWWDSH PKNSKWL ENL + V
Sbjct: 462 MATLSHSESRRSYSWWWDSH-LPKNSKWLQENLAGLSLPV 578
>TC208480
Length = 1576
Score = 58.9 bits (141), Expect = 2e-08
Identities = 78/334 (23%), Positives = 137/334 (40%), Gaps = 18/334 (5%)
Frame = +3
Query: 828 VEASKFSDKVISELESENLMQLIEEEFLLHRIRKFKMDIHKVCGVLQIDSDGGGDNEIKK 887
V+ S+ I L+ +N + + +E I+K K + + I D IK
Sbjct: 72 VQKEAISEDTIRNLKEKNDVHVQKETLSQETIKKLKEE-----NEVHIQEDFLSKETIKN 236
Query: 888 EEIPISRILDKIESLESSLVKSQEENQQLLVENSVLLGSLQQHQSEGEKLKLEKKTVEQE 947
+ ++L K+ SLE + Q +N+ +++ L + Q QSE L ++ + ++
Sbjct: 237 LKEENDKLLQKVVSLEEVINNLQTDNELQTQKHTSLEMRIAQLQSENNSLLQKEAGLVEK 416
Query: 948 FENMREQNVILQKDKVELLEENRQLRIEVVNGVEKENRSKSTLAALQAEMIELRQTNQVF 1007
+ + V+L L + L E+ + EKE ++ +A LQ+E L Q
Sbjct: 417 TNQLLNEKVVLSLKAESLERKINLLENELSSFSEKEVGLETRIAQLQSENNSLLQKEATL 596
Query: 1008 QEENGKMLDEKNSLCRNVSDLK--------DAKSSAEDENSVMFHDVLALSNLNLVYEIF 1059
E ++L+EK L L+ D S + ENS +SNLN I
Sbjct: 597 VERTNQLLNEKAVLSLKGESLEQKIYLLESDLNSLVKKENSTK----ETISNLN--GNIA 758
Query: 1060 FTENMVEKRALCEHLGNLSHLNNDLNQEFGVLR---KNFEVKEAENVYLNESIERMDKEL 1116
+ VE+ L E NL N L ++ L+ +N E + + S++ + E
Sbjct: 759 VLQAQVEE--LEESRNNLFLENQQLREKVSSLQSTVQNHENSNTSSCSWDASVKDLASEN 932
Query: 1117 LEMDKRLKAAET-------SNAEFSRHIEELKME 1143
++ ++AA T NAE + EL +E
Sbjct: 933 EDLKSEIEAAFTLVEKLMAENAELVEKVTELCVE 1034
Score = 50.4 bits (119), Expect = 6e-06
Identities = 60/339 (17%), Positives = 133/339 (38%), Gaps = 25/339 (7%)
Frame = +3
Query: 900 ESLESSLVKSQEENQQLLVENSVLLGSLQQHQSEGEKLKLEKKTVEQE-FENMREQNVI- 957
E ++ +E + V+ + ++ E + ++K+T+ QE + ++E+N +
Sbjct: 18 EIASEETIRKLKEKHDMDVQKEAISEDTIRNLKEKNDVHVQKETLSQETIKKLKEENEVH 197
Query: 958 -----LQKDKVELLEENRQLRI-------EVVNGVEKENRSKSTL-AALQAEMIELRQTN 1004
L K+ ++ L+E + EV+N ++ +N ++ +L+ + +L+ N
Sbjct: 198 IQEDFLSKETIKNLKEENDKLLQKVVSLEEVINNLQTDNELQTQKHTSLEMRIAQLQSEN 377
Query: 1005 QVFQEENGKMLDEKNSLCRNVSDLKDAKSSAEDENSVMFHDVLALSNLNLVYEIFFTENM 1064
++ ++++ N L L S E + +++ +++ + S + E +
Sbjct: 378 NSLLQKEAGLVEKTNQLLNEKVVLSLKAESLERKINLLENELSSFSEKEVGLETRIAQLQ 557
Query: 1065 VEKRALCEHLGNLSHLNNDLNQEFGVLRKNFEVKEAENVYLNESIERMDKELLEMDKRLK 1124
E +L + L N L E VL E E + L + + K+ + +
Sbjct: 558 SENNSLLQKEATLVERTNQLLNEKAVLSLKGESLEQKIYLLESDLNSLVKKENSTKETIS 737
Query: 1125 AAETSNAEFSRHIEELKMEQEESTKIKENLDRQILEQSENCMNHKK----------EIEH 1174
+ A +EEL+ + + L ++ NH+ ++
Sbjct: 738 NLNGNIAVLQAQVEELEESRNNLFLENQQLREKVSSLQSTVQNHENSNTSSCSWDASVKD 917
Query: 1175 LNEANETLQFEMKTLLHEVEQHRVREEALNLELLNKENE 1213
L NE L+ E++ VE + A N EL+ K E
Sbjct: 918 LASENEDLKSEIEAAFTLVE----KLMAENAELVEKVTE 1022
Score = 38.1 bits (87), Expect = 0.033
Identities = 61/276 (22%), Positives = 116/276 (41%), Gaps = 40/276 (14%)
Frame = +3
Query: 1144 QEESTKIKENLDRQILEQ--SENCMNHKKEIEHLNEANETLQFEMKTLLHEVEQHRVREE 1201
+E K+KE D + ++ SE+ + + KE ++ ETL E L E + ++E+
Sbjct: 30 EETIRKLKEKHDMDVQKEAISEDTIRNLKEKNDVHVQKETLSQETIKKLKEENEVHIQED 209
Query: 1202 ALNLELLN--KENEFKLWE---------------NEAAAFYHD------LQMSSICLALL 1238
L+ E + KE KL + NE H Q+ S +LL
Sbjct: 210 FLSKETIKNLKEENDKLLQKVVSLEEVINNLQTDNELQTQKHTSLEMRIAQLQSENNSLL 389
Query: 1239 ESKVSELTGVCKILDDESSAKSLENEHMREIISLLESEIGGLKEQLSAYVPLVSSLKEDF 1298
+ + + ++L+ E SL+ E + I+LLE+E+ E+ ++ L+ +
Sbjct: 390 QKEAGLVEKTNQLLN-EKVVLSLKAESLERKINLLENELSSFSEKEVGLETRIAQLQSEN 566
Query: 1299 NS-LEHISLLWTKRNSVVGNGA----------QKDVVIETCLRKHSHQSARENEIVLIPD 1347
NS L+ + L + N ++ A QK ++E+ L + E + +
Sbjct: 567 NSLLQKEATLVERTNQLLNEKAVLSLKGESLEQKIYLLESDLNSLVKKENSTKETISNLN 746
Query: 1348 GVSDLLTLQTRIRAVEK----IMMEELKRRVKQKSL 1379
G ++ LQ ++ +E+ + +E + R K SL
Sbjct: 747 G--NIAVLQAQVEELEESRNNLFLENQQLREKVSSL 848
>TC228989 similar to UP|Q9LSS5 (Q9LSS5) Myosin heavy chain-like, partial
(23%)
Length = 884
Score = 57.4 bits (137), Expect = 5e-08
Identities = 61/247 (24%), Positives = 115/247 (45%), Gaps = 3/247 (1%)
Frame = +1
Query: 371 MLEKSNQNLQLEAENLVQRIASKDHELLEKHTEIERLQTLMHGEH-SNFIQIESALQALQ 429
+L++ QLEAE + S E+ + + IE +T E + +QI +A + ++
Sbjct: 7 ILKEGEDPNQLEAE-----VVSLKSEVEQLRSAIETAETKYQEEQIRSMVQIRNAYELVE 171
Query: 430 KLYSQSQKEQRNLALELKYGLLLLKDLELSKQDFKEEMQGIVEENKTLH-ELNFS-STRS 487
++ S+S K + L ELK +++L+ + D + E+QGIVEEN L+ +L S S+++
Sbjct: 172 QIKSESGKRECELEAELKKKKADIEELKANLMDKETELQGIVEENDNLNLKLEESMSSKN 351
Query: 488 LKKQQMEISKLKEIKEKLEREFHTSTEESNVLQRETHQIKDDIQHLNERYQAMLEQLQSL 547
K + EI +L E +L+ + + E +K +I + + + + +++
Sbjct: 352 EHKLKREIKRLAECVAELKADMMDKETTLQSISEENEMLKSEINNSGKAREEVAAEVE-- 525
Query: 548 GLNPNSFAASVRDLQNENFMLKETCKKEHSEKEALREKSKDMNEVLMENACMEFSLLGLN 607
+ AA L +++E K S K+A R ++ + N ME L L
Sbjct: 526 ----GAKAAEHEALMKLGIVMEEADK---SNKKAAR-VAEQLEAAQAANLEMEAELRRLK 681
Query: 608 DELDGLR 614
+ D R
Sbjct: 682 VQSDQWR 702
>AW234049
Length = 421
Score = 55.1 bits (131), Expect = 3e-07
Identities = 30/90 (33%), Positives = 47/90 (51%)
Frame = +1
Query: 1027 DLKDAKSSAEDENSVMFHDVLALSNLNLVYEIFFTENMVEKRALCEHLGNLSHLNNDLNQ 1086
DL + KS E+E +M HD +A SNL+L+Y+ E + + L + L L +N DL +
Sbjct: 4 DLGEEKSKLEEEICIMIHDTIAQSNLSLLYQNIVLEKLQALKELSKDLDRLCSVNTDLEE 183
Query: 1087 EFGVLRKNFEVKEAENVYLNESIERMDKEL 1116
+ ++ E + EN L ES+ EL
Sbjct: 184 KLKIMMGKLEDVQMENSDLKESLILSSNEL 273
>CF923172
Length = 371
Score = 53.5 bits (127), Expect = 7e-07
Identities = 34/108 (31%), Positives = 58/108 (53%)
Frame = -1
Query: 1192 EVEQHRVREEALNLELLNKENEFKLWENEAAAFYHDLQMSSICLALLESKVSELTGVCKI 1251
E+ + ++REE LN ELL NE + WE +A+ + +LQ S++ L E KV EL + ++
Sbjct: 365 ELXEIKLREEKLNCELLKGTNEIEQWETQASTLFAELQXSAVNETLFEGKVCELNEMFRV 186
Query: 1252 LDDESSAKSLENEHMREIISLLESEIGGLKEQLSAYVPLVSSLKEDFN 1299
L E + E + M E + E + E+ ++ + +SS K+ N
Sbjct: 185 LHTEKT----ELQRMVEDLKTKYDEARVMLEEKASRILKLSSDKDRQN 54
>TC213072
Length = 548
Score = 52.8 bits (125), Expect = 1e-06
Identities = 25/58 (43%), Positives = 38/58 (65%), Gaps = 1/58 (1%)
Frame = +3
Query: 39 KVKSMIKLIEEE-ADSFARRAEMYYKKRPELMKLVEEFYRAYRALAERYDHAMGELRH 95
KV +M EEE D+FA RAE YY+KRP+L+ L+++ Y Y L++RY + + +H
Sbjct: 66 KVLAMSTTEEEEMGDTFAERAETYYQKRPQLLSLLQDLYNGYITLSDRYIQTLAKHKH 239
>TC220873 homologue to UP|Q9SFF4 (Q9SFF4) T26F17.2, partial (5%)
Length = 851
Score = 45.4 bits (106), Expect = 2e-04
Identities = 31/112 (27%), Positives = 52/112 (45%), Gaps = 7/112 (6%)
Frame = +2
Query: 293 RMLNDQIERTELEVETLRKNLAEMNEERDSLSVLYHHC-------LEKISKMENEILHVQ 345
R N ++E + L K L +M E+ L + C L +I ++E E++ +Q
Sbjct: 308 RTKNAEVEAIVQKNAALEKKLEKMEAEKLELEMDLTECQKQLEASLSRIKEVELEVVELQ 487
Query: 346 ENAEQLKNKIEKEAEKLEISEKHRGMLEKSNQNLQLEAENLVQRIASKDHEL 397
N E+ EKLE +E+ + + E + EAE LV +I S + E+
Sbjct: 488 TKLALANNSNEEAYEKLEATEEKKEIAESKLRVAHTEAEELVSKICSLEEEI 643
>TC203824 similar to UP|Q9V3T8 (Q9V3T8) SR family splicing factor SC35
(CG5442-PB) (LD32469p), partial (6%)
Length = 1532
Score = 45.1 bits (105), Expect = 3e-04
Identities = 63/274 (22%), Positives = 130/274 (46%), Gaps = 3/274 (1%)
Frame = +2
Query: 574 KEHSEKEALREKSKDMNEVLMENACMEFSLLGLNDELDGLRGTVKEIQQFCQVLQEE--K 631
+E + K A ++ +D E++ ENA + + L ELDGLR + + + LQ E +
Sbjct: 275 EESATKIAALQRERD--ELVSENAARKEEIKKLTAELDGLRSDGADRSEKIEELQREVAQ 448
Query: 632 SILADEKSTLLSQLQIITESMQKILENNTVLEKSLS-DAKIEFEGLRIKSGDLEDCCKLL 690
S A + + +++ E+ L ++ V E + + +A+ + LR G+ E + L
Sbjct: 449 SRDAAKATEVIAARAADLETEVARLHHDMVAEMTAAEEARADAAELRKVLGETESRVESL 628
Query: 691 NDEKNNLQNERSMLISQLEIVEEKLSNLEKKVTNLEEKYADVEKDKESAVNQVEELFASI 750
+E L+ ++ ++ +E K+ LE K +EE+ + ++E+ ++++E I
Sbjct: 629 ENELAGLKKVKAESDVRVRDLERKIGVLETK--EIEERNKRIRFEEETR-DKIDEKEREI 799
Query: 751 LVQKENHSNHKHSSEARLANLENIVRVLQEEQRLGKVEFEQELDRVVNAQIEMFILQNCI 810
++ + + + L+ +V+ Q+L E +E + V +E ILQ
Sbjct: 800 KGYRQKVEELEKVAAEKKTELDELVK-----QKLRLEEALRESEEKVTG-LESSILQLRK 961
Query: 811 EELELKNFVLLTECEKLVEASKFSDKVISELESE 844
E E + V+++ EK VE + D+ ++ + SE
Sbjct: 962 EAKEAER-VIISLNEKAVETVESIDRGLNGVHSE 1060
>CA801926 similar to GP|9828634|gb F1N21.5 {Arabidopsis thaliana}, partial (8%)
Length = 478
Score = 43.5 bits (101), Expect = 8e-04
Identities = 34/131 (25%), Positives = 64/131 (47%), Gaps = 2/131 (1%)
Frame = +3
Query: 1087 EFGVLRKNFEV--KEAENVYLNESIERMDKELLEMDKRLKAAETSNAEFSRHIEELKMEQ 1144
E G+ K EV KEAE + E + + ++ L + ++LK E + + ++E +
Sbjct: 15 EHGLKNKLVEVEKKEAEITHAEEKVVKREQALGKKAEKLKEKEIEYEQKVKALKEKEKLI 194
Query: 1145 EESTKIKENLDRQILEQSENCMNHKKEIEHLNEANETLQFEMKTLLHEVEQHRVREEALN 1204
+ K E R+I + E + HK E+E + NE E+ + E+++ +V EE +
Sbjct: 195 KSEEKSLETEKRKIESEREELLTHKAEVEKIRANNEE---ELLRINEEIDRLKVTEEERS 365
Query: 1205 LELLNKENEFK 1215
E L +++ K
Sbjct: 366 -EYLRLQSQLK 395
Score = 42.0 bits (97), Expect = 0.002
Identities = 39/148 (26%), Positives = 69/148 (46%), Gaps = 17/148 (11%)
Frame = +3
Query: 331 LEKISKMENEILHVQENAEQLKNKIEKEAEKL-----EISEKHRGMLEKS----NQNLQL 381
L ++ K E EI H +E + + + K+AEKL E +K + + EK ++ L
Sbjct: 36 LVEVEKKEAEITHAEEKVVKREQALGKKAEKLKEKEIEYEQKVKALKEKEKLIKSEEKSL 215
Query: 382 EAENLVQRIASKDHELLEKHTEIERLQTLMHGEHSNFIQIESALQALQKLYSQSQKEQRN 441
E E ++I S+ ELL E+E+++ ++I + L K+ + + E
Sbjct: 216 ETEK--RKIESEREELLTHKAEVEKIRA---NNEEELLRINEEIDRL-KVTEEERSEYLR 377
Query: 442 LALELKYGL--------LLLKDLELSKQ 461
L +LK+ + LLLK+ E +Q
Sbjct: 378 LQSQLKHEVDQYRHQKELLLKEAEDLRQ 461
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.310 0.128 0.336
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 54,836,054
Number of Sequences: 63676
Number of extensions: 619748
Number of successful extensions: 3877
Number of sequences better than 10.0: 121
Number of HSP's better than 10.0 without gapping: 3474
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3738
length of query: 1662
length of database: 12,639,632
effective HSP length: 110
effective length of query: 1552
effective length of database: 5,635,272
effective search space: 8745942144
effective search space used: 8745942144
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 66 (30.0 bits)
Medicago: description of AC135320.2


