

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135318.3 + phase: 1 /pseudo
(726 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
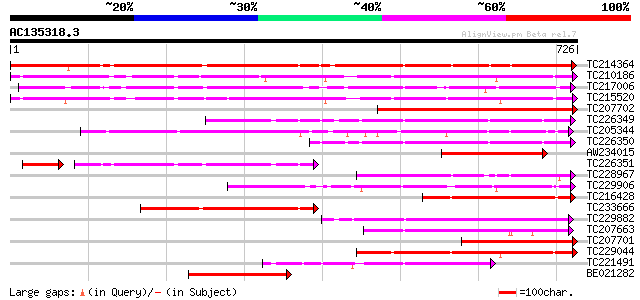
Sequences producing significant alignments: (bits) Value
TC214364 UP|Q6WNU4 (Q6WNU4) Subtilisin-like protease, complete 692 0.0
TC210186 UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, complete 457 e-129
TC217006 UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1, complete 456 e-128
TC215520 GB|AAG38994.1|11611651|AF160513 subtilisin-type proteas... 449 e-126
TC207702 weakly similar to UP|Q8S896 (Q8S896) Subtilisin-like se... 410 e-114
TC226349 similar to PIR|JC7519|JC7519 subtilisin-like serine pro... 340 1e-93
TC205344 similar to UP|Q9SZV5 (Q9SZV5) Proteinase-like protein (... 294 1e-79
TC226350 similar to PIR|JC7519|JC7519 subtilisin-like serine pro... 254 9e-68
AW234015 242 5e-64
TC226351 similar to PIR|JC7519|JC7519 subtilisin-like serine pro... 219 2e-63
TC228967 similar to GB|BAB09208.1|9758669|AB012245 subtilisin-li... 234 1e-61
TC229906 weakly similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like pr... 233 3e-61
TC216428 similar to UP|Q9AX30 (Q9AX30) Subtilisin-like protease,... 228 9e-60
TC233666 weakly similar to UP|Q9FJF3 (Q9FJF3) Serine protease-li... 228 9e-60
TC229882 weakly similar to UP|Q75I27 (Q75I27) Putaive subtilisin... 227 1e-59
TC207663 similar to UP|P93204 (P93204) SBT1 protein (Subtilisin-... 227 1e-59
TC207701 similar to UP|Q84TR6 (Q84TR6) Subtilase, partial (6%) 221 7e-58
TC229044 similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease,... 209 3e-54
TC221491 similar to UP|P93204 (P93204) SBT1 protein (Subtilisin-... 205 7e-53
BE021282 similar to GP|21593457|gb| subtilisin-like serine prote... 201 7e-52
>TC214364 UP|Q6WNU4 (Q6WNU4) Subtilisin-like protease, complete
Length = 2667
Score = 692 bits (1787), Expect = 0.0
Identities = 376/730 (51%), Positives = 481/730 (65%), Gaps = 5/730 (0%)
Frame = +1
Query: 1 DLLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTT 60
D LGS LGS AK++I YSY + INGFAA L+EE A +IAK+PKV+SVF ++ KLHTT
Sbjct: 208 DFLGSFLGSSNTAKDSIFYSYTRHINGFAATLDEEVAVEIAKHPKVLSVFENRGRKLHTT 387
Query: 61 RSWEFLGLRGNDI---NSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGG 117
RSW+F+ L N + +S W+K RFGE IIGN+DTGVWPESKSFS++G+GPIP+KWRG
Sbjct: 388 RSWDFMELEHNGVIQSSSIWKKARFGEGVIIGNLDTGVWPESKSFSEQGLGPIPSKWRG- 564
Query: 118 NICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGG 177
IC +T CNRKLIGAR+FNK Y G L S + RD GHGTHTLSTAGG
Sbjct: 565 -ICHNGIDHTFH---CNRKLIGARYFNKGYASVAGPLNSSFDSPRDNEGHGTHTLSTAGG 732
Query: 178 NFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDII 237
N V S+F G+GT KGGSP ARVA YKVCW CF AD+L+A D AI DGVD++
Sbjct: 733 NMVARVSVFGQGHGTAKGGSPMARVAAYKVCWPPVAGDECFDADILAAFDLAIHDGVDVL 912
Query: 238 SVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAA 297
SVS GG SST F D ++IG+FHA R +++V SAGN GP + N+APW TVAA
Sbjct: 913 SVSLGGSSST----FFKDSVAIGSFHAAKRGVVVVCSAGNSGPAEATAENLAPWHVTVAA 1080
Query: 298 STLDRDFSSVMTIGNK-TLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDARFCRPRTL 356
ST+DR F + + +GN T G SL ++ + I+ +TDAKLA+A DA C+ TL
Sbjct: 1081STMDRQFPTYVVLGNDITFKGESLSATKLAHKFYPIIKATDAKLASARAEDAVLCQNGTL 1260
Query: 357 DPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISY 416
DP+K GKIV C R G V +G++A AGA G++L N + G ++++PHVL
Sbjct: 1261DPNKAKGKIVVCLR-GINARVDKGEQAFLAGAVGMVLAND-KTTGNEIIADPHVL----- 1419
Query: 417 PGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKP 476
P +H T S + ++ KT KPAP MA++SS+GPN + P ILKP
Sbjct: 1420PASHINFTDGSAVFKYINSTKFPVAYITHPKTQLDTKPAPFMAAFSSKGPNTIVPEILKP 1599
Query: 477 DVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWS 536
D+TAPGV+++AAY+ +N + D RR PFN + GTSMSCPHV+G GL++ L+P WS
Sbjct: 1600DITAPGVSVIAAYTEAQGPTNQVFDKRR-IPFNSVSGTSMSCPHVSGIVGLLRALYPTWS 1776
Query: 537 PAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDY 596
AAIKSAIMTTATT DN +P+ +A D A PF+YG+GH++PN AMDPGLVYD+ I DY
Sbjct: 1777TAAIKSAIMTTATTLDNEVEPLLNATDGK-ATPFSYGAGHVQPNRAMDPGLVYDITIDDY 1953
Query: 597 LNFLCASGYNQQLISALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTNVGP 656
LNFLCA GYN+ IS + C S+ +LNYPSIT+P L SVTVTRT+ NVG
Sbjct: 1954LNFLCALGYNETQISVFT-EGPYKCRKKFSLLNLNYPSITVPKLS-GSVTVTRTLKNVGS 2127
Query: 657 PSTYFAKVQLA-GYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGK 715
P TY A VQ G ++V PS L FK +GE+K+F++ +A Y FG+L W++GK
Sbjct: 2128PGTYIAHVQNPYGITVSVKPSILKFKNVGEEKSFKLTFKAMQGKATNNYAFGKLIWSDGK 2307
Query: 716 HIVRSPVTVR 725
H V SP+ V+
Sbjct: 2308HYVTSPIVVK 2337
>TC210186 UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, complete
Length = 2401
Score = 457 bits (1177), Expect = e-129
Identities = 274/746 (36%), Positives = 425/746 (56%), Gaps = 21/746 (2%)
Frame = +1
Query: 2 LLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTR 61
+L S+L ENA ++ +Y +GFAA L ++EA IA+ P VVSVF KLHTTR
Sbjct: 244 VLNSVLRRNENA---LVRNYKHGFSGFAARLSKKEATSIAQKPGVVSVFPGPVLKLHTTR 414
Query: 62 SWEFLGLRGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQ 121
SW+FL + +++IG +DTG+WPE+ SFSD+G+GP+P++W+G +
Sbjct: 415 SWDFLKYQTQVKIDTKPNAVSKSSSVIGILDTGIWPEAASFSDKGMGPVPSRWKGTCMKS 594
Query: 122 LDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAGGNFVP 181
D +++ CNRKLIGAR++ TARD GHGTH TA G V
Sbjct: 595 QDFYSSN----CNRKLIGARYYADPNDS-------GDNTARDSNGHGTHVAGTAAGVMVT 741
Query: 182 GASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSA 241
AS + + G KGGSP +R+A Y+VC + C G+ +L+A D AI DGVD++SVS
Sbjct: 742 NASYYGVATGCAKGGSPESRLAVYRVCSNF----GCRGSSILAAFDDAIADGVDLLSVSL 909
Query: 242 GGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLD 301
G S+ ++ +D IS+GAFHA+ IL+V SAGN+GP+ ++VN APW+ TVAAST+D
Sbjct: 910 GA-STGFRPDLTSDPISLGAFHAMEHGILVVCSAGNDGPSSYTLVNDAPWILTVAASTID 1086
Query: 302 RDFSSVMTIG-NKTLTGASLFVNLPP---NQDFTIVTSTDAKLANATNRDARFCRPRTLD 357
R+F S + +G NK + G + +NL P + + ++ AK + + +AR C P +LD
Sbjct: 1087RNFLSNIVLGDNKIIKGKA--INLSPLSNSPKYPLIYGESAKANSTSLVEARQCHPNSLD 1260
Query: 358 PSKVNGKIVAC-DREGKIKSVAEGQEALSAGAKGVI-LRNQPEINGK-------TLLSEP 408
+KV GKIV C D+ K + + + G G++ + +Q E T++S
Sbjct: 1261GNKVKGKIVVCDDKNDKYSTRKKVATVKAVGGIGLVHITDQNEAIASNYGDFPATVISSK 1440
Query: 409 HVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNK 468
++ + Y + S L ++ KPAP++ ++SSRGP+
Sbjct: 1441DGVTILQYINSTSNPVATIL----------------ATTSVLDYKPAPLVPNFSSRGPSS 1572
Query: 469 VQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLI 528
+ +ILKPD+ APGVNILAA+ + + ++ ++ + ++ GTSM+CPHV+G A +
Sbjct: 1573LSSNILKPDIAAPGVNILAAW--IGNGTEVVPKGKKPSLYKIISGTSMACPHVSGLASSV 1746
Query: 529 KTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLV 588
KT +P WS ++IKSAIMT+A +N PI+ ++A P+ YG+G + + + PGLV
Sbjct: 1747KTRNPTWSASSIKSAIMTSAIQSNNLKAPITTE-SGSVATPYDYGAGEMTTSEPLQPGLV 1923
Query: 589 YDLGIKDYLNFLCASGYNQQLISALNFNM--TFTCS---GTSSIDDLNYPSITLPNLGLN 643
Y+ DYLNFLC G+N + ++ + F C + I ++NYPSI + G
Sbjct: 1924YETSSVDYLNFLCYIGFNVTTVKVISKTVPRNFNCPKDLSSDHISNINYPSIAINFSGKR 2103
Query: 644 SVTVTRTVTNVG-PPSTYFAKVQLA--GYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTP 700
+V ++RTVTNVG T ++ + A G + + P+ L F K +K +++VI +T +T
Sbjct: 2104AVNLSRTVTNVGEDDETVYSPIVDAPSGVHVTLTPNKLRFTKSSKKLSYRVIFSST-LTS 2280
Query: 701 RRKYQFGELRWTNGKHIVRSPVTVRR 726
++ FG + W+NGK++VRSP + +
Sbjct: 2281LKEDLFGSITWSNGKYMVRSPFVLTK 2358
>TC217006 UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1, complete
Length = 2422
Score = 456 bits (1172), Expect = e-128
Identities = 305/733 (41%), Positives = 416/733 (56%), Gaps = 20/733 (2%)
Frame = +3
Query: 12 NAKEAII-YSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLG--L 68
NA+ ++ + + + +GF AML EEEA ++A++ +VV+VF +K+ +LHTTRSW+F+G L
Sbjct: 207 NAEPKLVQHHFKRSFSGFVAMLTEEEADRMARHDRVVAVFPNKKKQLHTTRSWDFIGFPL 386
Query: 69 RGNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTS 128
+ N + + II D+G+WPES+SF+D+G GP P+KW+G CQ TS
Sbjct: 387 QAN-------RAPAESDVIIAVFDSGIWPESESFNDKGFGPPPSKWKG--TCQ-----TS 524
Query: 129 KKVPCNRKLIGARFFNKAYQKRNGKLPRSQ-QTARDFVGHGTHTLSTAGGNFVPGASIFN 187
K CN K+IGA+ + K +G + ++ RD GHGTH STA GN V AS+
Sbjct: 525 KNFTCNNKIIGAKIY-----KVDGFFSKDDPKSVRDIDGHGTHVASTAAGNPVSTASMLG 689
Query: 188 IGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSST 247
+G GT +GG +AR+A YKVCW C AD+L+A D AI DGVDII+VS GG S
Sbjct: 690 LGQGTSRGGVTKARIAVYKVCWF----DGCTDADILAAFDDAIADGVDIITVSLGGFSDE 857
Query: 248 NSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSV 307
N F D I+IGAFHA+ +L V SAGN GP P S+ N +PW +VAAST+DR F +
Sbjct: 858 N---YFRDGIAIGAFHAVRNGVLTVTSAGNSGPRPSSLSNFSPWSISVAASTIDRKFVTK 1028
Query: 308 MTIGNK-TLTGASLFVNLPPNQDFTIVTSTDA--KLANATNRDARFCRPRTLDPSKVNGK 364
+ +GNK T G S+ + + I+ DA K +R+C +LD V GK
Sbjct: 1029VELGNKITYEGTSINTFDLKGELYPIIYGGDAPNKGEGIDGSSSRYCSSGSLDKKLVKGK 1208
Query: 365 IVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNH-SRT 423
IV C+ K AGA G +++ Q + ++ PG++ +
Sbjct: 1209IVLCESRSKALG------PFDAGAVGALIQGQ---------GFRDLPPSLPLPGSYLALQ 1343
Query: 424 TGRSLDIIPSDIKSGTKLRMSPAKTLNRRKP--APVMASYSSRGPNKVQPSILKPDVTAP 481
G S+ D + T+ ++ + K APV+AS+SSRGPN V P ILKPD+ AP
Sbjct: 1344DGASV----YDYINSTRTPIATIFKTDETKDTIAPVVASFSSRGPNIVTPEILKPDLVAP 1511
Query: 482 GVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIK 541
GV+ILA++S + S++ D R FN++ GTSM+CPHV+G A +K+ HP WSPAAI+
Sbjct: 1512GVSILASWSPASPPSDVEGDNRT-LNFNIISGTSMACPHVSGAAAYVKSFHPTWSPAAIR 1688
Query: 542 SAIMTTATTRDNTNKPISDAFDKT-LANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFL 600
SA+MTTA K +S KT L FAYG+G I P+ A+ PGLVYD G DY+ FL
Sbjct: 1689SALMTTA-------KQLS---PKTHLLAEFAYGAGQIDPSKAVYPGLVYDAGEIDYVRFL 1838
Query: 601 CASGYNQ---QLISALNFNMTFTCSGTSSIDDLNYPSITL--PNLGLNSV--TVTRTVTN 653
C GY+ QLI+ N + T +G S DLNY S L P NSV + RTVTN
Sbjct: 1839CGQGYSTRTLQLITGDNSSCPETKNG--SARDLNYASFALFVPPYNSNSVSGSFNRTVTN 2012
Query: 654 VG-PPSTYFAKV-QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRW 711
VG P STY A V G KI V PS L F + +K+TF + + P G L W
Sbjct: 2013VGSPKSTYKATVTSPKGLKIEVNPSVLPFTSLNQKQTFVLTITGKLEGP---IVSGSLVW 2183
Query: 712 TNGKHIVRSPVTV 724
+GK+ VRSP+ V
Sbjct: 2184DDGKYQVRSPIVV 2222
>TC215520 GB|AAG38994.1|11611651|AF160513 subtilisin-type protease precursor
{Glycine max;} , complete
Length = 2392
Score = 449 bits (1154), Expect = e-126
Identities = 287/751 (38%), Positives = 419/751 (55%), Gaps = 26/751 (3%)
Frame = +2
Query: 2 LLGSILGSKENAKEAIIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTR 61
+L S+L ENA ++ +Y +GFAA L +EEA IA+ P VVSVF KLHTTR
Sbjct: 212 ILNSVLRRNENA---LVRNYKHGFSGFAARLSKEEANSIAQKPGVVSVFPDPILKLHTTR 382
Query: 62 SWEFLGLR-----GNDINSAWQKGRFGENTIIGNIDTGVWPESKSFSDRGIGPIPAKWRG 116
SW+FL + N+ + I+G +DTG+WPE+ SFSD+G GP+P++W+G
Sbjct: 383 SWDFLKSQTRVNIDTKPNTLSGSSFSSSDVILGVLDTGIWPEAASFSDKGFGPVPSRWKG 562
Query: 117 GNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKRNGKLPRSQQTARDFVGHGTHTLSTAG 176
+ D N+S CNRK+IGARF+ +K TARDF GHGTH STA
Sbjct: 563 TCMTSKD-FNSSC---CNRKIIGARFYPNPEEK----------TARDFNGHGTHVSSTAV 700
Query: 177 GNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDI 236
G V GAS + + GT +GGSP +R+A YKVC + SC G+ +L+ D AI DGVDI
Sbjct: 701 GVPVSGASFYGLAAGTARGGSPESRLAVYKVCGAF---GSCPGSAILAGFDDAIHDGVDI 871
Query: 237 ISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVA 296
+S+S GG T + ++ TD I+IGAFH++ R IL+V +AGN+G P +V+N APW+ TVA
Sbjct: 872 LSLSLGGFGGTKT-DLTTDPIAIGAFHSVQRGILVVCAAGNDG-EPFTVLNDAPWILTVA 1045
Query: 297 ASTLDRDFSSVMTIG-NKTLTGASL-FVNLPPNQDFTIVTSTDAKLANATN-RDARFCRP 353
AST+DRD S + +G N+ + G ++ F L + D+ ++ + A AN +N DAR C P
Sbjct: 1046ASTIDRDLQSDVVLGNNQVVKGRAINFSPLLNSPDYPMIYAESAARANISNITDARQCHP 1225
Query: 354 RTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGK----------T 403
+LDP KV GKIV CD + I + + + G+ L + + +G T
Sbjct: 1226DSLDPKKVIGKIVVCDGKNDIYYSTDEKIVIVKALGGIGLVHITDQSGSVAFYYVDFPVT 1405
Query: 404 LLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSS 463
+ H + + Y + S G L + T+ KPAP + +SS
Sbjct: 1406EVKSKHGDAILQYINSTSHPVGTILATV----------------TIPDYKPAPRVGYFSS 1537
Query: 464 RGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAG 523
RGP+ + ++LKPD+ APGVNILAA+ F + ++ + R+ + ++ GTSM+ PHV+G
Sbjct: 1538RGPSLITSNVLKPDIAAPGVNILAAW--FGNDTSEVPKGRKPSLYRILSGTSMATPHVSG 1711
Query: 524 TAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAM 583
A +K +P WS +AIKSAIMT+A DN PI+ +A P+ YG+G I + +
Sbjct: 1712LACSVKRKNPTWSASAIKSAIMTSAIQNDNLKGPIT-TDSGLIATPYDYGAGAITTSEPL 1888
Query: 584 DPGLVYDLGIKDYLNFLCASGYNQQLISALNFNM--TFTCSGTSS---IDDLNYPSITLP 638
PGLVY+ DYLN+LC +G N +I ++ + F C SS I +NYPSI +
Sbjct: 1889QPGLVYETNNVDYLNYLCYNGLNITMIKVISGTVPENFNCPKDSSSDLISSINYPSIAVN 2068
Query: 639 NLGLNSVTVTRTVTNVG--PPSTYFAKVQLAGYKIAVV-PSSLNFKKIGEKKTFQVIVQA 695
G V+RTVTNV + YF V+ I + P +L F +K+++ + +
Sbjct: 2069FTGKADAVVSRTVTNVDEEDETVYFPVVEAPSEVIVTLFPYNLEFTTSIKKQSYNITFRP 2248
Query: 696 TSVTPRRKYQFGELRWTNGKHIVRSPVTVRR 726
T +K FG + W+N K++VR P + +
Sbjct: 2249K--TSLKKDLFGSITWSNDKYMVRIPFVLTK 2335
>TC207702 weakly similar to UP|Q8S896 (Q8S896) Subtilisin-like serine
protease AIR3 (Fragment), partial (20%)
Length = 1025
Score = 410 bits (1053), Expect = e-114
Identities = 202/256 (78%), Positives = 223/256 (86%), Gaps = 1/256 (0%)
Frame = +2
Query: 472 SILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTL 531
SILKPDVTAPGVNILAAYS ASASNL+ D RRGF FNV+QGTS+SCPHVAG AGLIKTL
Sbjct: 2 SILKPDVTAPGVNILAAYSELASASNLLVDNRRGFKFNVLQGTSVSCPHVAGIAGLIKTL 181
Query: 532 HPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDL 591
HPNWSPAAIKSAIMTTATT DNTN+PI DAFD +A+ FAYGSGH++P+ A+DPGLVYDL
Sbjct: 182 HPNWSPAAIKSAIMTTATTLDNTNRPIQDAFDNKVADAFAYGSGHVQPDLAIDPGLVYDL 361
Query: 592 GIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTV 651
+ DYLNFLCASGY+QQLISALNFN TF C G+ S+ DLNYPSITLPNLGL VT+TRTV
Sbjct: 362 SLADYLNFLCASGYDQQLISALNFNGTFICKGSHSVTDLNYPSITLPNLGLKPVTITRTV 541
Query: 652 TNVGPPSTYFAKVQL-AGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELR 710
TNVGPP+TY A V AGY I VVP SL F KIGEKK FQVIVQA+SVT RRKYQFG+LR
Sbjct: 542 TNVGPPATYTANVHSPAGYTIVVVPRSLTFTKIGEKKKFQVIVQASSVTTRRKYQFGDLR 721
Query: 711 WTNGKHIVRSPVTVRR 726
WT+GKHIVRSP+TV+R
Sbjct: 722 WTDGKHIVRSPITVKR 769
>TC226349 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(56%)
Length = 1773
Score = 340 bits (873), Expect = 1e-93
Identities = 208/484 (42%), Positives = 284/484 (57%), Gaps = 10/484 (2%)
Frame = +1
Query: 251 EIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTI 310
+ + D ++IGAF A+ + IL+ SAGN GP P S+ NVAPW+ TV A TLDRDF + + +
Sbjct: 1 DYYRDSVAIGAFSAMEKGILVSCSAGNSGPGPYSLSNVAPWITTVGAGTLDRDFPAYVAL 180
Query: 311 GNK-TLTGASLFV-NLPPNQDFTIVTSTDAKLANATN--RDARFCRPRTLDPSKVNGKIV 366
GN +G SL+ N P+ +V + N +N + C TL P KV GKIV
Sbjct: 181 GNGLNFSGVSLYRGNALPDSSLPLVYA-----GNVSNGAMNGNLCITGTLSPEKVAGKIV 345
Query: 367 ACDREGKIKSVAEGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGR 426
CDR G V +G SAGA G++L N NG+ L+++ H+L + + G
Sbjct: 346 LCDR-GLTARVQKGSVVKSAGALGMVLSNTAA-NGEELVADAHLLPATAV----GQKAGD 507
Query: 427 SLD-IIPSDIKSGTKLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNI 485
++ + SD K K+ K +P+PV+A++SSRGPN + P ILKPD+ APGVNI
Sbjct: 508 AIKKYLVSDAKPTVKIFFEGTKV--GIQPSPVVAAFSSRGPNSITPQILKPDLIAPGVNI 681
Query: 486 LAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIM 545
LA +S + L D RR FN++ GTSMSCPHV+G A LIK+ HP+WSPAA++SA+M
Sbjct: 682 LAGWSKAVGPTGLPVDNRR-VDFNIISGTSMSCPHVSGLAALIKSAHPDWSPAAVRSALM 858
Query: 546 TTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGY 605
TTA T T + + D+ + PF +GSGH+ P +A++PGLVYDL + DYL FLCA Y
Sbjct: 859 TTAYTVYKTGEKLQDSATGKPSTPFDHGSGHVDPVAALNPGLVYDLTVDDYLGFLCALNY 1038
Query: 606 NQQLISALNFNMTFTCSGTS--SIDDLNYPSITLPNLGLNSVTV-TRTVTNVGPPSTYFA 662
+ IS L F C S+ DLNYPS + SV TRT+TNVGP TY A
Sbjct: 1039SAAEISTL-AKRKFQCDAGKQYSVTDLNYPSFAVLFESSGSVVKHTRTLTNVGPAGTYKA 1215
Query: 663 KV--QLAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRS 720
V A KI+V P L+FK+ EKKTF V ++ + FG + W++GKH+V S
Sbjct: 1216SVTSDTASVKISVEPQVLSFKE-NEKKTFTVTFSSSGSPQHTENAFGRVEWSDGKHLVGS 1392
Query: 721 PVTV 724
P++V
Sbjct: 1393PISV 1404
>TC205344 similar to UP|Q9SZV5 (Q9SZV5) Proteinase-like protein (AT4g30020
protein) (AT4g30020/F6G3_50), partial (78%)
Length = 2192
Score = 294 bits (752), Expect = 1e-79
Identities = 232/672 (34%), Positives = 332/672 (48%), Gaps = 40/672 (5%)
Frame = +2
Query: 91 IDTGVWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGARFFNKAYQKR 150
+D+G++P SF+ P R C++D +KK CN K++GA+ F +A
Sbjct: 8 VDSGIYPHHPSFTTHNTEPYGPVSRYRGKCEVDP--DTKKSFCNGKIVGAQHFAQAAIAA 181
Query: 151 NGKLPRSQ-QTARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCW 209
P + D GHG+HT S A G + G G +PRAR+A YK +
Sbjct: 182 GAFNPFIVFDSPLDGDGHGSHTASIAAGRNGIPVRMHGHEFGKASGMAPRARIAVYKALY 361
Query: 210 SLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSS-TNSEEIFTDEISIGAFHALARN 268
L F ADV++AIDQA+ DGVDI+S+S G S +N++ F + A+
Sbjct: 362 RLFGG---FIADVVAAIDQAVHDGVDILSLSVGPNSPPSNTKTTFLNPFDATLLGAVKAG 532
Query: 269 ILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-KTLTGASLFVNLPPN 327
+ + +AGN GP P S+V+ +PW+ TVAA+ DR + + + +GN K L G L + N
Sbjct: 533 VFVAQAAGNGGPFPKSLVSYSPWIATVAAAIDDRRYKNHLILGNGKILAGLGLSPSTRLN 712
Query: 328 QDFTIVTSTDAKL-ANATNRDARFC-RPRTLDPSKVNGKIVACDRE-------GKIKSVA 378
Q +T+V +TD L ++ T C RP L+ + + G I+ C IK V+
Sbjct: 713 QTYTLVAATDVLLDSSVTKYSPTDCQRPELLNKNLIKGNILLCGYSYNFVIGSASIKQVS 892
Query: 379 EGQEALSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDII------- 431
E +AL GA G +L + G P + PG +S ++I
Sbjct: 893 ETAKAL--GAVGFVLYVENVSPGTKFDPVP-----VGIPGILITDASKSKELIDYYNIST 1051
Query: 432 PSDIKSGTKLRMSPAKTLNRRKP-----APVMASYSSRGPNKV-----QPSILKPDVTAP 481
P D K K + P AP +A +S+RGPN + +LKPD+ AP
Sbjct: 1052PRDWTGRVKTFEGTGKIEDGLMPILHKSAPQVAMFSARGPNIKDFSFQEADLLKPDILAP 1231
Query: 482 GVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIK 541
G I AA+SL + G F ++ GTSM+ PH+AG A LIK HP+WSPAAIK
Sbjct: 1232GSLIWAAWSLNGTDE----PNYAGEGFAMISGTSMAAPHIAGIAALIKQKHPHWSPAAIK 1399
Query: 542 SAIMTTATTRDNTNKPI-------SDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIK 594
SA+MTT+TT D PI ++A A PF YGSGH+ P +A+DPGL++D G +
Sbjct: 1400SALMTTSTTLDRAGNPILAQLYSETEAMKLVKATPFDYGSGHVNPRAALDPGLIFDAGYE 1579
Query: 595 DYLNFLCAS-GYNQQLISALNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTN 653
DYL FLC + G + I N+ + + +LN PSIT+ +L + S VTRTVTN
Sbjct: 1580DYLGFLCTTPGIDVHEIK--NYTNSPCNNTMGHPSNLNTPSITISHL-VRSQIVTRTVTN 1750
Query: 654 VG-PPSTYFAKVQL-AGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRW 711
V TY ++ I V P ++ K ++ F V + SVT Y FGE+
Sbjct: 1751VADEEETYVITARMQPAVAIDVNPPAMTIKASASRR-FTVTLTVRSVT--GTYSFGEVLM 1921
Query: 712 TNGK-HIVRSPV 722
+ H VR PV
Sbjct: 1922KGSRGHKVRIPV 1957
>TC226350 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(34%)
Length = 1312
Score = 254 bits (649), Expect = 9e-68
Identities = 148/344 (43%), Positives = 206/344 (59%), Gaps = 4/344 (1%)
Frame = +1
Query: 385 SAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMS 444
SAGA G++L N NG+ L+++ H+L + + L SD K K+
Sbjct: 7 SAGALGMVLSNTAA-NGEELVADAHLLPATAVGQKAGDAIKKYLF---SDAKPTVKILFE 174
Query: 445 PAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRR 504
K +P+PV+A++SSRGPN + P ILKPD+ APGVNILA +S + L D RR
Sbjct: 175 GTKL--GIQPSPVVAAFSSRGPNSITPQILKPDLIAPGVNILAGWSKAVGPTGLPVDNRR 348
Query: 505 GFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDK 564
FN++ GTSMSCPHV+G A LIK+ HP+WSPAA++SA+MTTA T T + + D+
Sbjct: 349 -VDFNIISGTSMSCPHVSGLAALIKSAHPDWSPAAVRSALMTTAYTVYKTGEKLQDSATG 525
Query: 565 TLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGT 624
+ PF +GSGH+ P +A++PGLVYDL + DYL FLCA Y+ I+ L F C
Sbjct: 526 KPSTPFDHGSGHVDPVAALNPGLVYDLTVDDYLGFLCALNYSASEINTL-AKRKFQCDAG 702
Query: 625 S--SIDDLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYFAKV--QLAGYKIAVVPSSLNF 680
S+ DLNYPS + V TRT+TNVGP TY A V +A KI+V P L+F
Sbjct: 703 KQYSVTDLNYPSFAVLFESGGVVKHTRTLTNVGPAGTYKASVTSDMASVKISVEPQVLSF 882
Query: 681 KKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTV 724
K+ EKK+F V ++ +R FG + W++GKH+V +P+++
Sbjct: 883 KE-NEKKSFTVTFSSSGSPQQRVNAFGRVEWSDGKHVVGTPISI 1011
>AW234015
Length = 411
Score = 242 bits (617), Expect = 5e-64
Identities = 115/136 (84%), Positives = 125/136 (91%)
Frame = +3
Query: 553 NTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISA 612
NTNKPI DAFDKTLANPFAYGSGH++PNSA+DPGL+YDL I DYLNFLCASGY+QQLISA
Sbjct: 3 NTNKPIGDAFDKTLANPFAYGSGHVQPNSAIDPGLIYDLSIVDYLNFLCASGYDQQLISA 182
Query: 613 LNFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTVTRTVTNVGPPSTYFAKVQLAGYKIA 672
LNFN TFTCSG+ SI DLNYPSITLPNLGLN++TVTRTVTNVGP STYFAK QL GY I
Sbjct: 183 LNFNSTFTCSGSHSITDLNYPSITLPNLGLNAITVTRTVTNVGPASTYFAKAQLRGYNIV 362
Query: 673 VVPSSLNFKKIGEKKT 688
VVPSSL+FKKIGEK+T
Sbjct: 363 VVPSSLSFKKIGEKRT 410
>TC226351 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(48%)
Length = 1490
Score = 219 bits (559), Expect(2) = 2e-63
Identities = 134/320 (41%), Positives = 191/320 (58%), Gaps = 7/320 (2%)
Frame = +3
Query: 83 GENTIIGNIDTG-VWPESKSFSDRGIGPIPAKWRGGNICQLDKLNTSKKVPCNRKLIGAR 141
G + IIG ++T + ++ + ++ G+GP+P+ W+G C+ T+ CNRKLIGAR
Sbjct: 585 GSDVIIGVLNTWRLAGKAGASTNTGLGPVPSTWKGA--CETGTNLTASN--CNRKLIGAR 752
Query: 142 FFNKAYQKRNGKLPRSQQT--ARDFVGHGTHTLSTAGGNFVPGASIFNIGNGTIKGGSPR 199
FF+K + G + ++++ ARD GHGTHT STA G+ V AS+F +GT +G + R
Sbjct: 753 FFSKGVEAILGPINETEESRSARDDDGHGTHTASTAAGSVVSDASLFGYASGTARGMATR 932
Query: 200 ARVATYKVCWSLTDATSCFGADVLSAIDQAIDDGVDIISVSAGGPSSTNSEEIFTDEISI 259
ARVA YKVCW CF +D+L+AI++AI D V+++S+S GG S + + D ++I
Sbjct: 933 ARVAAYKVCWK----GGCFSSDILAAIERAILDNVNVLSLSLGGGMS----DYYRDSVAI 1088
Query: 260 GAFHALARNILLVASAGNEGPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-KTLTGA 318
GAF A+ IL+ SAGN GP+P S+ NVAPW+ TV A TLDRDF + + +GN +G
Sbjct: 1089GAFSAMENGILVSCSAGNAGPSPYSLSNVAPWITTVGAGTLDRDFPAYVALGNGLNFSGV 1268
Query: 319 SLF-VNLPPNQDFTIVTSTDAKLANATN--RDARFCRPRTLDPSKVNGKIVACDREGKIK 375
SL+ N P+ V + N +N + C TL P KV GKIV CDR G
Sbjct: 1269SLYRGNAVPDSPLPFVYA-----GNVSNGAMNGNLCITGTLSPEKVAGKIVLCDR-GLTA 1430
Query: 376 SVAEGQEALSAGAKGVILRN 395
V +G SAGA G++L N
Sbjct: 1431RVQKGSVVKSAGALGMVLSN 1490
Score = 42.0 bits (97), Expect(2) = 2e-63
Identities = 20/52 (38%), Positives = 32/52 (61%)
Frame = +2
Query: 17 IIYSYNKQINGFAAMLEEEEAAQIAKNPKVVSVFLSKEHKLHTTRSWEFLGL 68
I+Y+Y+ I+G+A L EEA + +++V ++LHTTR+ FLGL
Sbjct: 392 IMYTYDNAIHGYATRLTAEEARVLETQAGILAVLPETRYELHTTRTPMFLGL 547
>TC228967 similar to GB|BAB09208.1|9758669|AB012245 subtilisin-like protease
{Arabidopsis thaliana;} , partial (21%)
Length = 1106
Score = 234 bits (597), Expect = 1e-61
Identities = 138/289 (47%), Positives = 173/289 (59%), Gaps = 9/289 (3%)
Frame = +2
Query: 445 PAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRR 504
P T+ KPAP MAS+SSRGPN V P+ILKPD+TAPGV+ILAA++ + + + +R
Sbjct: 41 PGTTVLETKPAPSMASFSSRGPNIVDPNILKPDITAPGVDILAAWTAEDGPTRMTFNDKR 220
Query: 505 GFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDK 564
+N+ GTSMSCPHVA A L+K +HP WS AAI+SA+MTTA T DNT P++D
Sbjct: 221 VVKYNIFSGTSMSCPHVAAAAVLLKAIHPTWSTAAIRSALMTTAMTTDNTGHPLTDETGN 400
Query: 565 TLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGT 624
A PFA GSGH P A DPGLVYD YL + C G Q NFN+T+ C
Sbjct: 401 P-ATPFAMGSGHFNPKRAADPGLVYDASYMGYLLYTCNLGVTQ------NFNITYNCP-K 556
Query: 625 SSID--DLNYPSITLPNLGLNSVTVTRTVTNVG-PPSTY-FAKVQLAGYKIAVVPSSLNF 680
S ++ +LNYPSI + L + T+ RTVTNVG S Y F+ V Y I P+ L F
Sbjct: 557 SFLEPFELNYPSIQIHRL-YYTKTIKRTVTNVGRGRSVYKFSAVSPKEYSITATPNILKF 733
Query: 681 KKIGEKKTFQVIVQAT-SVTPRR----KYQFGELRWTNGKHIVRSPVTV 724
+G+K F + V A S P + KY FG WT+ HIVRSPV V
Sbjct: 734 NHVGQKINFTITVTANWSQIPTKHGPDKYYFGWYAWTHQHHIVRSPVAV 880
>TC229906 weakly similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1,
partial (40%)
Length = 1392
Score = 233 bits (593), Expect = 3e-61
Identities = 172/460 (37%), Positives = 241/460 (52%), Gaps = 14/460 (3%)
Frame = +2
Query: 279 GPTPGSVVNVAPWVFTVAASTLDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTD 337
GP S+ +PW+ +VAAST+ R F + + +GN G S+ N+ F +V + D
Sbjct: 5 GPGLSSITTYSPWILSVAASTIGRKFLTKVQLGNGMVFEGVSINTFDLKNKMFPLVYAGD 184
Query: 338 A-KLANATNRD-ARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRN 395
A+ N +RFC ++D V GKIV CD K V + +GA G++L
Sbjct: 185 VPNTADGYNSSTSRFCYVNSVDKHLVKGKIVLCDGNASPKKVGD-----LSGAAGMLL-- 343
Query: 396 QPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTL-----N 450
G T + + + +Y + + R+ +I S + S LR S A N
Sbjct: 344 -----GATDVKD----APFTYALPTAFISLRNFKLIHSYMVS---LRNSTATIFRSDEDN 487
Query: 451 RRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNV 510
P + S+SSRGPN + P+ LKPD+ APGVNILAA+S + S D +R +N+
Sbjct: 488 DDSQTPFIVSFSSRGPNPLTPNTLKPDLAAPGVNILAAWSPVYTISEFKGD-KRAVQYNI 664
Query: 511 MQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPF 570
GTSM+CPHV+ A +K+ HPNWSPA IKSA+MTTAT T P ++ F
Sbjct: 665 ESGTSMACPHVSAAAAYVKSFHPNWSPAMIKSALMTTATPMSPTLNPDAE---------F 817
Query: 571 AYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCS---GTSSI 627
AYG+G I P A +PGLVYD+ DY+ FLC GY +++ L + + CS ++
Sbjct: 818 AYGAGLINPLKAANPGLVYDISEADYVKFLCGEGYTDEMLRVLTKDHS-RCSKHAKKEAV 994
Query: 628 DDLNYPSITL-PNLGLNSVTVTRTVTNVG-PPSTYFAKVQLAG-YKIAVVPSSLNFKKIG 684
DLN PS+ L N+ S RTVTNVG S+Y AKV I V P+ L+F IG
Sbjct: 995 YDLNLPSLALYVNVSSFSRIFHRTVTNVGLATSSYKAKVVSPSLIDIQVKPNVLSFTSIG 1174
Query: 685 EKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTV 724
+KK+F VI++ +V P L W +G VRSP+ V
Sbjct: 1175QKKSFSVIIEG-NVNP--DILSASLVWDDGTFQVRSPIVV 1285
>TC216428 similar to UP|Q9AX30 (Q9AX30) Subtilisin-like protease, partial
(12%)
Length = 810
Score = 228 bits (580), Expect = 9e-60
Identities = 118/198 (59%), Positives = 141/198 (70%), Gaps = 2/198 (1%)
Frame = +1
Query: 529 KTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRPNSAMDPGLV 588
KT HP WSPAAIKSAIMTTATT DNTN+PI +AF K +A PF YG+GHI+PN A+DPGLV
Sbjct: 1 KTYHPTWSPAAIKSAIMTTATTLDNTNQPIRNAFHK-VATPFEYGAGHIQPNLAIDPGLV 177
Query: 589 YDLGIKDYLNFLCASGYNQQLISAL-NFNMTFTCSGTSSIDDLNYPSITLPNLGLNSVTV 647
YDL DYLNFLCASGYNQ L++ +TC + I+D NYPSIT+ + G +++V
Sbjct: 178 YDLRTTDYLNFLCASGYNQALLNLFAKLKFPYTCPKSYRIEDFNYPSITVRHPGSKTISV 357
Query: 648 TRTVTNVGPPSTYFAKVQ-LAGYKIAVVPSSLNFKKIGEKKTFQVIVQATSVTPRRKYQF 706
TRTVTNVGPPSTY G K+ V PSSL FK+ GEKK FQVI+Q R+ F
Sbjct: 358 TRTVTNVGPPSTYVVNTHGPKGIKVLVQPSSLTFKRTGEKKKFQVILQPIGA---RRGLF 528
Query: 707 GELRWTNGKHIVRSPVTV 724
G L WT+GKH V SP+T+
Sbjct: 529 GNLSWTDGKHRVTSPITI 582
>TC233666 weakly similar to UP|Q9FJF3 (Q9FJF3) Serine protease-like protein,
partial (30%)
Length = 678
Score = 228 bits (580), Expect = 9e-60
Identities = 118/229 (51%), Positives = 152/229 (65%), Gaps = 1/229 (0%)
Frame = +1
Query: 168 GTHTLSTAGGNFVPGASIFNIGNGTIKGGSPRARVATYKVCWSLTDATSCFGADVLSAID 227
G+HTLST GG FVPGA++F +GNGT +GGSPRARVATYKVCW D CF AD+++A D
Sbjct: 4 GSHTLSTIGGTFVPGANVFGLGNGTAEGGSPRARVATYKVCWPPIDGNECFDADIMAAFD 183
Query: 228 QAIDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVN 287
AI DGVD++S+S GG N+ + F D +SIGAFHA + I ++ SAGN GPTP +V N
Sbjct: 184 MAIHDGVDVLSLSLGG----NATDYFDDGLSIGAFHANMKGIPVICSAGNYGPTPATVFN 351
Query: 288 VAPWVFTVAASTLDRDFSSVMTIGN-KTLTGASLFVNLPPNQDFTIVTSTDAKLANATNR 346
VAPW+ TV ASTLDR F SV+ + N + GASL +P ++ + ++ + DAK AN
Sbjct: 352 VAPWILTVGASTLDRQFDSVVELHNGQRFMGASLSKAMPEDKLYPLINAADAKAANKPVE 531
Query: 347 DARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEALSAGAKGVILRN 395
+A C T+DP K GKI+ C R G V + AL AGA +IL N
Sbjct: 532 NATLCMRGTIDPEKARGKILVCLR-GVTARVEKSLVALEAGAAXMILCN 675
>TC229882 weakly similar to UP|Q75I27 (Q75I27) Putaive subtilisin-like
proteinase, partial (27%)
Length = 1263
Score = 227 bits (579), Expect = 1e-59
Identities = 132/327 (40%), Positives = 185/327 (56%), Gaps = 4/327 (1%)
Frame = +1
Query: 400 NGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKSGTKLRMSPAKTLNRRKPAPVMA 459
+G+ L+++ H+L ++ R G + S + T + + T+ KP+PV+A
Sbjct: 16 SGEELVADSHLLPAVAV----GRIVGDQIRAYASSDPNPT-VHLDFRGTVLNVKPSPVVA 180
Query: 460 SYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQGTSMSCP 519
++SSRGPN V ILKPDV PGVNILA +S S L DTR+ FN+M GTSMSCP
Sbjct: 181 AFSSRGPNMVTRQILKPDVIGPGVNILAGWSEAIGPSGLSDDTRKT-QFNIMSGTSMSCP 357
Query: 520 HVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAYGSGHIRP 579
H++G A L+K HP WS +AIKSA+MTTA DNT + DA +NP+A+G+GH+ P
Sbjct: 358 HISGLAALLKAAHPQWSSSAIKSALMTTADVHDNTKSQLRDAAGGAFSNPWAHGAGHVNP 537
Query: 580 NSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGT-SSIDDLNYPSITLP 638
+ A+ PGLVYD DY+ FLC+ Y + I + C+ S LNYPS ++
Sbjct: 538 HKALSPGLVYDATPSDYIKFLCSLEYTPERIQLITKRSGVNCTKRFSDPGQLNYPSFSVL 717
Query: 639 NLGLNSVTVTRTVTNVGPP-STYFAKVQL-AGYKIAVVPSSLNFKKIGEKKTF-QVIVQA 695
G V TR +TNVG S Y V + + V P++L F K+GE++ + V
Sbjct: 718 FGGKRVVRYTRVLTNVGEAGSVYNVTVDAPSTVTVTVKPAALVFGKVGERQRYTATFVSK 897
Query: 696 TSVTPRRKYQFGELRWTNGKHIVRSPV 722
V +Y FG + W+N +H VRSPV
Sbjct: 898 NGVGDSVRYGFGSIMWSNAQHQVRSPV 978
>TC207663 similar to UP|P93204 (P93204) SBT1 protein (Subtilisin-like
protease) , partial (37%)
Length = 1171
Score = 227 bits (579), Expect = 1e-59
Identities = 127/285 (44%), Positives = 168/285 (58%), Gaps = 15/285 (5%)
Frame = +1
Query: 453 KPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRRGFPFNVMQ 512
+P+PV+A++SSRGPN + P ILKPD+ APGVNILA ++ + L DTR FN++
Sbjct: 73 QPSPVVAAFSSRGPNALTPKILKPDLIAPGVNILAGWTGAVGPTGLTVDTRH-VSFNIIS 249
Query: 513 GTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDKTLANPFAY 572
GTSMSCPHV+G A ++K HP WSPAAI+SA+MTTA T + I D PF Y
Sbjct: 250 GTSMSCPHVSGLAAILKGAHPQWSPAAIRSALMTTAYTSYKNGETIQDISTGQPGTPFDY 429
Query: 573 GSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSS--IDDL 630
G+GH+ P +A+DPGLVYD + DYL F CA Y+ I L +TC ++D
Sbjct: 430 GAGHVDPVAALDPGLVYDANVDDYLGFFCALNYSSFQIK-LAARRDYTCDPKKDYRVEDF 606
Query: 631 NYPSITLP-----NLG-----LNSVTVTRTVTNVGPPSTYFAKVQLAG---YKIAVVPSS 677
NYPS +P +G L +V +R +TNVG P TY A V G K V P++
Sbjct: 607 NYPSFAVPMDTASGIGGGSDTLKTVKYSRVLTNVGAPGTYKASVMSLGDSNVKTVVEPNT 786
Query: 678 LNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPV 722
L+F ++ EKK + V TS+ P F L WT+GKH V SP+
Sbjct: 787 LSFTELYEKKDYTVSFTYTSM-PSGTTSFARLEWTDGKHKVGSPI 918
>TC207701 similar to UP|Q84TR6 (Q84TR6) Subtilase, partial (6%)
Length = 763
Score = 221 bits (564), Expect = 7e-58
Identities = 107/149 (71%), Positives = 125/149 (83%), Gaps = 1/149 (0%)
Frame = +2
Query: 579 PNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTFTCSGTSSIDDLNYPSITLP 638
P+ A++PGLVYDL + DYLNFLCASGY+QQLISALNFN TF CSG+ S++DLNYPSITLP
Sbjct: 2 PDLAIEPGLVYDLSLTDYLNFLCASGYDQQLISALNFNRTFICSGSHSVNDLNYPSITLP 181
Query: 639 NLGLNSVTVTRTVTNVGPPSTYFAKVQLA-GYKIAVVPSSLNFKKIGEKKTFQVIVQATS 697
NL L VT+ RTVTNVGPPSTY + GY IAVVP SL F KIGE+KTF+VIVQA+S
Sbjct: 182 NLRLKPVTIARTVTNVGPPSTYTVSTRSPNGYSIAVVPPSLTFTKIGERKTFKVIVQASS 361
Query: 698 VTPRRKYQFGELRWTNGKHIVRSPVTVRR 726
RRKY+FG+LRWT+GKHIVRSP+TV+R
Sbjct: 362 AATRRKYEFGDLRWTDGKHIVRSPITVKR 448
>TC229044 similar to UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, partial
(42%)
Length = 984
Score = 209 bits (533), Expect = 3e-54
Identities = 118/290 (40%), Positives = 179/290 (61%), Gaps = 8/290 (2%)
Frame = +1
Query: 445 PAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLITDTRR 504
P T+ KPAPV+ ++SSRGP+ + +ILKPD+ APGVNILAA+ + ++ + R+
Sbjct: 100 PTATVLDYKPAPVVPNFSSRGPSSLSSNILKPDIAAPGVNILAAW--IGNNADDVPKGRK 273
Query: 505 GFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPISDAFDK 564
+N++ GTSM+CPHV+G A +KT +P WS +AIKSAIMT+A +N PI+ +
Sbjct: 274 PSLYNIISGTSMACPHVSGLASSVKTRNPTWSASAIKSAIMTSAIQINNLKAPITTDSGR 453
Query: 565 TLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNM--TFTCS 622
+A P+ YG+G + + ++ PGLVY+ DYLN+LC G N + ++ + F+C
Sbjct: 454 -VATPYDYGAGEMTTSESLQPGLVYETNTIDYLNYLCYIGLNITTVKVISRTVPANFSCP 630
Query: 623 GTSS---IDDLNYPSITLPNLGLNSVTVTRTVTNVG--PPSTYFAKVQL-AGYKIAVVPS 676
SS I ++NYPSI + G +V V+RTVTNVG + Y V+ +G K+ V P
Sbjct: 631 KDSSSDLISNINYPSIAVNFTGKAAVNVSRTVTNVGEEDETAYSPVVEAPSGVKVTVTPD 810
Query: 677 SLNFKKIGEKKTFQVIVQATSVTPRRKYQFGELRWTNGKHIVRSPVTVRR 726
L F K +K +QVI +T +T ++ FG + W+NGK++VRSP + +
Sbjct: 811 KLQFTKSSKKLGYQVIFSST-LTSLKEDLFGSITWSNGKYMVRSPFVLTK 957
>TC221491 similar to UP|P93204 (P93204) SBT1 protein (Subtilisin-like
protease) , partial (34%)
Length = 872
Score = 205 bits (521), Expect = 7e-53
Identities = 126/303 (41%), Positives = 167/303 (54%), Gaps = 4/303 (1%)
Frame = +3
Query: 324 LPPNQDFTIVTSTDAKLANATNRDARFCRPRTLDPSKVNGKIVACDREGKIKSVAEGQEA 383
LPPN IV + AN ++ C TL KV GKIV CDR G + V +G
Sbjct: 15 LPPNSPLPIVYA-----ANVSDESQNLCTRGTLIAEKVAGKIVICDRGGNAR-VEKGLVV 176
Query: 384 LSAGAKGVILRNQPEINGKTLLSEPHVLSTISYPGNHSRTTGRSLDIIPSDIKS----GT 439
SAG G+IL N + G+ L+++ ++L + S + + P+ GT
Sbjct: 177 KSAGGIGMILSNNEDY-GEELVADSYLLPAAALGQKSSNELKKYVFSSPNPTAKLGFGGT 353
Query: 440 KLRMSPAKTLNRRKPAPVMASYSSRGPNKVQPSILKPDVTAPGVNILAAYSLFASASNLI 499
+L + +P+PV+A++SSRGPN + P ILKPD+ APGVNILA ++ + L
Sbjct: 354 QLGV---------QPSPVVAAFSSRGPNVLTPKILKPDLIAPGVNILAGWTGAVGPTGLT 506
Query: 500 TDTRRGFPFNVMQGTSMSCPHVAGTAGLIKTLHPNWSPAAIKSAIMTTATTRDNTNKPIS 559
DTR FN++ GTSMSCPHV G A L+K HP WSPAAI+SA+MTTA + I
Sbjct: 507 EDTRH-VEFNIISGTSMSCPHVTGLAALLKGTHPEWSPAAIRSALMTTAYRTYKNGQTIK 683
Query: 560 DAFDKTLANPFAYGSGHIRPNSAMDPGLVYDLGIKDYLNFLCASGYNQQLISALNFNMTF 619
D A PF YG+GH+ P +A DPGLVYD + DYL+F CA Y+ I L F
Sbjct: 684 DVATGLPATPFDYGAGHVDPVAAFDPGLVYDTTVDDYLSFFCALNYSPYQIK-LVARRDF 860
Query: 620 TCS 622
TCS
Sbjct: 861 TCS 869
>BE021282 similar to GP|21593457|gb| subtilisin-like serine protease
{Arabidopsis thaliana}, partial (5%)
Length = 402
Score = 201 bits (512), Expect = 7e-52
Identities = 98/132 (74%), Positives = 114/132 (86%)
Frame = +1
Query: 230 IDDGVDIISVSAGGPSSTNSEEIFTDEISIGAFHALARNILLVASAGNEGPTPGSVVNVA 289
IDDGVDIIS+SAGG E IFTDE+SIGAFHA+ARN +LVASAGN+GPTPG+V+NVA
Sbjct: 1 IDDGVDIISLSAGGSYVVTPEGIFTDEVSIGAFHAIARNRILVASAGNDGPTPGTVLNVA 180
Query: 290 PWVFTVAASTLDRDFSSVMTIGNKTLTGASLFVNLPPNQDFTIVTSTDAKLANATNRDAR 349
PWVFT+AASTLDRDFSS +TI N+ +TGASLFVNLPPN+ F+++ +TDAKLANAT RDA
Sbjct: 181 PWVFTIAASTLDRDFSSNLTINNRQITGASLFVNLPPNKAFSLILATDAKLANATFRDAE 360
Query: 350 FCRPRTLDPSKV 361
CRP TLDP KV
Sbjct: 361 LCRPGTLDPEKV 396
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.132 0.387
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 29,157,683
Number of Sequences: 63676
Number of extensions: 383985
Number of successful extensions: 2234
Number of sequences better than 10.0: 127
Number of HSP's better than 10.0 without gapping: 2034
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2050
length of query: 726
length of database: 12,639,632
effective HSP length: 104
effective length of query: 622
effective length of database: 6,017,328
effective search space: 3742778016
effective search space used: 3742778016
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135318.3


