

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135317.6 + phase: 0
(432 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
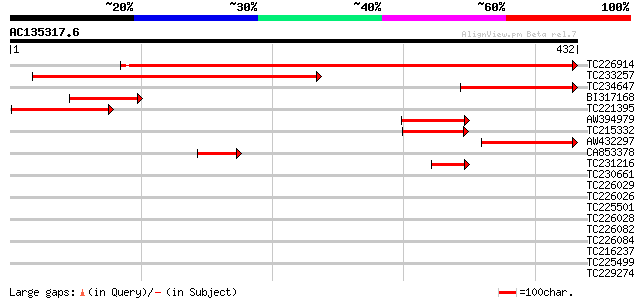
Score E
Sequences producing significant alignments: (bits) Value
TC226914 weakly similar to UP|Q93Z33 (Q93Z33) AT3g63150/T20O10_2... 364 e-101
TC233257 weakly similar to UP|Q93Z33 (Q93Z33) AT3g63150/T20O10_2... 345 3e-95
TC234647 115 4e-26
BI317168 similar to GP|19571035|dbj rac-GTP binding protein -lik... 95 7e-20
TC221395 similar to UP|Q8S298 (Q8S298) Rac-GTP binding protein-l... 87 2e-17
AW394979 73 2e-13
TC215332 similar to UP|P93344 (P93344) Aldehyde dehydrogenase (N... 70 2e-12
AW432297 63 3e-10
CA853378 weakly similar to GP|16648803|gb| AT3g63150/T20O10_250 ... 56 3e-08
TC231216 43 3e-04
TC230661 homologue to UP|RAC2_ARATH (Q38903) RAC-like GTP bindin... 39 0.006
TC226029 homologue to UP|RAC1_BETVU (Q39435) RAC-like GTP bindin... 37 0.023
TC226026 homologue to UP|Q9SWE8 (Q9SWE8) RAC-like G-protein Rac1... 37 0.023
TC225501 homologue to UP|Q9SXT7 (Q9SXT7) Rac-type small GTP-bind... 37 0.023
TC226028 homologue to GB|AAD00118.1|4097583|NTU64924 NTGP3 {Nico... 35 0.066
TC226082 homologue to UP|Q8S2V3 (Q8S2V3) Small G-protein ROP9, c... 35 0.066
TC226084 UP|Q8S2V3 (Q8S2V3) Small G-protein ROP9, partial (44%) 35 0.086
TC216237 similar to UP|Q70WD8 (Q70WD8) RAC-ROP-like G-protein, p... 34 0.11
TC225499 homologue to UP|Q9M628 (Q9M628) Small GTP binding prote... 34 0.15
TC229274 homologue to UP|RAC2_LOTJA (Q40220) RAC-like GTP bindin... 34 0.15
>TC226914 weakly similar to UP|Q93Z33 (Q93Z33) AT3g63150/T20O10_250, partial
(48%)
Length = 1426
Score = 364 bits (934), Expect = e-101
Identities = 180/349 (51%), Positives = 247/349 (70%), Gaps = 1/349 (0%)
Frame = +3
Query: 85 DDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDKDQLLRPAEVDKLFDAAPESPWKD 144
DD +P P K A DQSVEL+ A+EFL+ +F D D D +LRP E+++LF APESPW
Sbjct: 3 DDLIP-PIKCAPDQSVELTNEAVEFLRAIFDAFDGDGDGMLRPRELEELFSTAPESPWTG 179
Query: 145 APYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSLANLICIGYRDNPSAALRLTSRRSE 204
PY DAAE G +SL+ FLS WALMTLL+P +S+ NLI IGY + S+A+R+T RR
Sbjct: 180 IPYEDAAEKNAFGGLSLEAFLSEWALMTLLNPTFSVENLIYIGYPGDSSSAIRVTRRRRL 359
Query: 205 DRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTK 264
DRK Q ++RNV QC+VFG + AGKSALL + +GRP+S Y PTT +RYA N++++ +
Sbjct: 360 DRKKQHSDRNVLQCFVFGPRKAGKSALLNSFIGRPYSESYNPTTEDRYAVNVVDISMENR 539
Query: 265 KTLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGH 324
K L+LREIPED V LSNK+ LA+CD+A FVHD SD SW+ S +LL ++ + GE TG
Sbjct: 540 KYLVLREIPEDGVRKLLSNKESLASCDIAVFVHDRSDESSWRTSSELLVEIASHGEDTGF 719
Query: 325 RFPSLLIAAKDDLTPFPRAVLDSVKVAQELKIDAPIRLSIKSDDSNNVYSKIINAAEHPH 384
P L++AAKDDL FP A+ +S +V+Q++ ++API +S+K D N+++ KI+ AAEHPH
Sbjct: 720 EVPCLIVAAKDDLDSFPMAIQESTRVSQDMGVEAPIPISVKLGDFNSLFRKIVTAAEHPH 899
Query: 385 LSIPETEFVRKRKQHQQLLHTFIFALA-GAAMALAGFTARRARANRNSS 432
LSIPETE R RKQ+ +L++ + A++ GAA+ + G A R A R ++
Sbjct: 900 LSIPETEAGRSRKQYHRLINRSLMAVSVGAAVVVVGLAAYRVYAARKNA 1046
>TC233257 weakly similar to UP|Q93Z33 (Q93Z33) AT3g63150/T20O10_250, partial
(23%)
Length = 661
Score = 345 bits (884), Expect = 3e-95
Identities = 169/220 (76%), Positives = 189/220 (85%)
Frame = +2
Query: 18 FGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLKKGRTETFWAVLRKFGY 77
F A LQQ ++ +KT+V + VPEGVNS+GLTFPGFI +HN+FLKKGRTET WAVLRKF Y
Sbjct: 2 FNAPLQQFEVANIKTIVEQKVPEGVNSIGLTFPGFIYVHNLFLKKGRTETLWAVLRKFEY 181
Query: 78 GNDLKLRDDFLPVPSKKASDQSVELSGAAIEFLKGVFRLVDTDKDQLLRPAEVDKLFDAA 137
NDLKLRDDFLPVPSK+ASDQSVEL+ A+EFL G+FRL+DTDKD+ LRPAEVDKLFD A
Sbjct: 182 DNDLKLRDDFLPVPSKQASDQSVELTSEAVEFLNGIFRLLDTDKDRYLRPAEVDKLFDIA 361
Query: 138 PESPWKDAPYMDAAETTDMGYISLKGFLSRWALMTLLDPRYSLANLICIGYRDNPSAALR 197
PESPW DAPY DAAE TDMGYISL GFLS+WALMTLLDP+ SLANLI IGY NP+ ALR
Sbjct: 362 PESPWNDAPYKDAAEKTDMGYISLNGFLSQWALMTLLDPKRSLANLIYIGYNGNPAEALR 541
Query: 198 LTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLG 237
+T RRS DRK Q TERNVFQCYVFGSK AGKSAL+ +LLG
Sbjct: 542 VTRRRSVDRKKQTTERNVFQCYVFGSKHAGKSALMYSLLG 661
>TC234647
Length = 482
Score = 115 bits (288), Expect = 4e-26
Identities = 60/90 (66%), Positives = 72/90 (79%), Gaps = 1/90 (1%)
Frame = +2
Query: 344 VLDSVKVAQELKIDAPIRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLL 403
VLDSVKV Q L I API +S+K DS+NVY+KI+NAAEHPHLSIPETE R++KQH QLL
Sbjct: 2 VLDSVKVTQALGIKAPIHVSMKLGDSSNVYNKIVNAAEHPHLSIPETETARRKKQHNQLL 181
Query: 404 -HTFIFALAGAAMALAGFTARRARANRNSS 432
H+ IFAL GAAMA+AG TA R RA + ++
Sbjct: 182 HHSLIFALVGAAMAVAGLTACRVRAVKKNT 271
>BI317168 similar to GP|19571035|dbj rac-GTP binding protein -like {Oryza
sativa (japonica cultivar-group)}, partial (3%)
Length = 279
Score = 94.7 bits (234), Expect = 7e-20
Identities = 42/56 (75%), Positives = 48/56 (85%)
Frame = +2
Query: 46 GLTFPGFIEIHNMFLKKGRTETFWAVLRKFGYGNDLKLRDDFLPVPSKKASDQSVE 101
GLTFPGFI +HNMFLKKGR ET WAVLR FGY N+LKL DDFLP+PSK+A DQ ++
Sbjct: 2 GLTFPGFIYVHNMFLKKGRPETLWAVLRDFGYDNNLKLMDDFLPIPSKRALDQVID 169
>TC221395 similar to UP|Q8S298 (Q8S298) Rac-GTP binding protein-like, partial
(30%)
Length = 608
Score = 86.7 bits (213), Expect = 2e-17
Identities = 39/78 (50%), Positives = 54/78 (69%)
Frame = +3
Query: 2 DGTLTDHEVNEFQIRSFGASLQQPDITQLKTMVLRNVPEGVNSLGLTFPGFIEIHNMFLK 61
DG L+D E+N+FQ++ F A LQ +I +K +V + EGVN GLT GF+ +H +F++
Sbjct: 375 DGALSDAELNDFQVKCFNAPLQPSEIVGVKKVVQEKLSEGVNERGLTLTGFLFLHALFIE 554
Query: 62 KGRTETFWAVLRKFGYGN 79
KGR ET W VLRKFGY +
Sbjct: 555 KGRLETTWTVLRKFGYND 608
>AW394979
Length = 407
Score = 73.2 bits (178), Expect = 2e-13
Identities = 37/52 (71%), Positives = 42/52 (80%)
Frame = +1
Query: 299 SSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVKV 350
SSD +SWK+S DLLEKV QG+LTG+R LLI AKD LTP+PRAV DSVKV
Sbjct: 250 SSDEHSWKKSRDLLEKVGRQGDLTGYRVSCLLIGAKDYLTPYPRAVQDSVKV 405
>TC215332 similar to UP|P93344 (P93344) Aldehyde dehydrogenase (NAD+) ,
partial (6%)
Length = 751
Score = 70.1 bits (170), Expect = 2e-12
Identities = 35/50 (70%), Positives = 40/50 (80%)
Frame = +3
Query: 300 SDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKDDLTPFPRAVLDSVK 349
SD +SWK+S DLLEKV QG+LTG+R LLI AKD LTP+PRAV DSVK
Sbjct: 54 SDEHSWKKSRDLLEKVGRQGDLTGYRVSCLLIGAKDYLTPYPRAVQDSVK 203
>AW432297
Length = 390
Score = 62.8 bits (151), Expect = 3e-10
Identities = 31/74 (41%), Positives = 49/74 (65%), Gaps = 1/74 (1%)
Frame = +3
Query: 360 IRLSIKSDDSNNVYSKIINAAEHPHLSIPETEFVRKRKQHQQLLHTFIFALA-GAAMALA 418
I +S+K D N+++ KI+ AAEHPHLSIPETE R RKQ+ +L++ + A++ GAA+ +
Sbjct: 3 IPISVKLGDFNSLFRKIVTAAEHPHLSIPETEAGRSRKQYHRLINRSLMAVSVGAAVVIX 182
Query: 419 GFTARRARANRNSS 432
G A + ++
Sbjct: 183 GLATYXVYAAKKNA 224
>CA853378 weakly similar to GP|16648803|gb| AT3g63150/T20O10_250 {Arabidopsis
thaliana}, partial (4%)
Length = 138
Score = 56.2 bits (134), Expect = 3e-08
Identities = 25/33 (75%), Positives = 28/33 (84%)
Frame = +1
Query: 144 DAPYMDAAETTDMGYISLKGFLSRWALMTLLDP 176
DA Y DAAE TDMGYI + GFL++WALMTLLDP
Sbjct: 1 DALYKDAAERTDMGYIXINGFLAQWALMTLLDP 99
>TC231216
Length = 726
Score = 42.7 bits (99), Expect = 3e-04
Identities = 21/29 (72%), Positives = 23/29 (78%)
Frame = +3
Query: 322 TGHRFPSLLIAAKDDLTPFPRAVLDSVKV 350
TG+R LLI AKD LTP+PRAV DSVKV
Sbjct: 3 TGYRVSCLLIGAKDYLTPYPRAVQDSVKV 89
>TC230661 homologue to UP|RAC2_ARATH (Q38903) RAC-like GTP binding protein
ARAC2, partial (74%)
Length = 632
Score = 38.5 bits (88), Expect = 0.006
Identities = 18/62 (29%), Positives = 30/62 (48%)
Frame = +1
Query: 195 ALRLTSRRSEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAA 254
AL ++ E+ K T R + +C G GK+ +L + F DY PT + ++A
Sbjct: 85 ALGYFAKEQEENKTMSTARFI-KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSA 261
Query: 255 NI 256
N+
Sbjct: 262 NV 267
>TC226029 homologue to UP|RAC1_BETVU (Q39435) RAC-like GTP binding protein
RHO1, complete
Length = 995
Score = 36.6 bits (83), Expect = 0.023
Identities = 47/197 (23%), Positives = 78/197 (38%), Gaps = 10/197 (5%)
Frame = +2
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 218 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VDGSTVNLGLWDTAGQE 391
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + P +L+ K
Sbjct: 392 DYNRLRPLSYRGADVFILAFSLISKASYE-----NIAKKWIPELRHYAPGVPIILVGTKL 556
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +EL+ I AP + S NV + +AA +
Sbjct: 557 DLRDDKQFFMDHPGAVPITTAQGEELRKLIGAPAYIECSSKTQQNV-KAVFDAAIKVVIQ 733
Query: 387 IPETEFVRKRKQHQQLL 403
P+ + RK ++ +L
Sbjct: 734 PPKLKKKRKTQKACSIL 784
>TC226026 homologue to UP|Q9SWE8 (Q9SWE8) RAC-like G-protein Rac1, complete
Length = 1339
Score = 36.6 bits (83), Expect = 0.023
Identities = 46/192 (23%), Positives = 76/192 (38%), Gaps = 10/192 (5%)
Frame = +3
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 93 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VDGSTVNLGLWDTAGQE 266
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + P +L+ K
Sbjct: 267 DYNRLRPLSYRGADVFILAFSLISKASYE-----NIAKKWIPELRHYAPGVPIILVGTKL 431
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +EL+ I AP + S NV + +AA L
Sbjct: 432 DLREDKQFFIDHPGAVPITTTQGEELRKLIGAPAYIECSSKTQQNV-KAVFDAAIKVVLQ 608
Query: 387 IPETEFVRKRKQ 398
P+ + +++ Q
Sbjct: 609 PPKQKKKKRKAQ 644
>TC225501 homologue to UP|Q9SXT7 (Q9SXT7) Rac-type small GTP-binding protein,
complete
Length = 1075
Score = 36.6 bits (83), Expect = 0.023
Identities = 48/202 (23%), Positives = 81/202 (39%), Gaps = 9/202 (4%)
Frame = +2
Query: 206 RKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKK 265
RK R + +C G GK+ +L + F DY PT + ++AN++ + G+
Sbjct: 83 RKAMSASRFI-KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV--VDGSTV 253
Query: 266 TLILREIPEDEVSNFLSNKDCLAACDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHR 325
L L + E N L A DV S++ ++ +K + +
Sbjct: 254 NLGLWDTAGQEDYNRLRPLSYRGA-DVFLLAFSLISRASYE---NVAKKWIPELRHYAPG 421
Query: 326 FPSLLIAAKDDL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKI 376
P +L+ K DL P AV + +EL+ I API + S NV +
Sbjct: 422 VPIILVGTKLDLRDDKQFFQDHPGAVPITTAQGEELRKLIGAPIYIECSSKTQQNV-KAV 598
Query: 377 INAAEHPHLSIPETEFVRKRKQ 398
+AA L P+ + +++ Q
Sbjct: 599 FDAAIKVVLQPPKQKKKKRKGQ 664
>TC226028 homologue to GB|AAD00118.1|4097583|NTU64924 NTGP3 {Nicotiana
tabacum;} , partial (63%)
Length = 494
Score = 35.0 bits (79), Expect = 0.066
Identities = 19/65 (29%), Positives = 30/65 (45%)
Frame = +3
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 144 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VDGSTVNLALWDTAGQE 317
Query: 277 VSNFL 281
N L
Sbjct: 318 DYNRL 332
>TC226082 homologue to UP|Q8S2V3 (Q8S2V3) Small G-protein ROP9, complete
Length = 1127
Score = 35.0 bits (79), Expect = 0.066
Identities = 46/192 (23%), Positives = 77/192 (39%), Gaps = 10/192 (5%)
Frame = +3
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILREIPEDE 276
+C G GK+ LL + F DY PT + ++AN++ + G+ L L + E
Sbjct: 228 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV--VNGSIVNLGLWDTAGQE 401
Query: 277 VSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSLLIAAKD 335
N L A + AF S Y ++ +K + + + P +L+ K
Sbjct: 402 DYNRLRPLSYRGADVFILAFSLISKASYE-----NVSKKWIPELKHYAPGVPIILVGTKL 566
Query: 336 DL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAAEHPHLS 386
DL P AV + +EL+ I+AP + S NV + +AA L
Sbjct: 567 DLRDDKQFCIDHPGAVPITTAQGEELRKLINAPAYIECSSKTQENV-KAVFDAAIRVVLQ 743
Query: 387 IPETEFVRKRKQ 398
P+ + + + Q
Sbjct: 744 PPKQKKKKGKAQ 779
>TC226084 UP|Q8S2V3 (Q8S2V3) Small G-protein ROP9, partial (44%)
Length = 431
Score = 34.7 bits (78), Expect = 0.086
Identities = 13/41 (31%), Positives = 21/41 (50%)
Frame = +2
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
+C G GK+ LL + F DY PT + ++AN++
Sbjct: 191 KCVTVGDGAVGKTCLLISYTSNTFPTDYVPTVFDNFSANVV 313
>TC216237 similar to UP|Q70WD8 (Q70WD8) RAC-ROP-like G-protein, partial (90%)
Length = 1651
Score = 34.3 bits (77), Expect = 0.11
Identities = 45/198 (22%), Positives = 76/198 (37%), Gaps = 10/198 (5%)
Frame = +2
Query: 211 TERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANIIELIAGTKKTLILR 270
T +C G GK+ +L F DY PT + ++AN++ + G L L
Sbjct: 218 TAPRFIKCVTVGDGAVGKTCMLICYTSNKFPTDYIPTVFDNFSANVV--VEGITVNLGLW 391
Query: 271 EIPEDEVSNFLSNKDCLAA-CDVAAFVHDSSDGYSWKRSIDLLEKVVNQGELTGHRFPSL 329
+ E N L A V AF S Y ++L+K + + + P +
Sbjct: 392 DTAGQEDYNRLRPLSYRGADVFVLAFSLVSRASYE-----NVLKKWIPELQHFAPGIPLV 556
Query: 330 LIAAKDDL-------TPFPRAVLDSVKVAQELK--IDAPIRLSIKSDDSNNVYSKIINAA 380
L+ K DL P V + +EL+ I A + S NV + +AA
Sbjct: 557 LVGTKLDLREDKHYMADHPSLVPVTTDQGEELRKHIGATYYIECSSKTQQNV-KAVFDAA 733
Query: 381 EHPHLSIPETEFVRKRKQ 398
+ P+ + +++K+
Sbjct: 734 IRMVIKPPQKQNEKRKKK 787
>TC225499 homologue to UP|Q9M628 (Q9M628) Small GTP binding protein RACDP,
partial (65%)
Length = 554
Score = 33.9 bits (76), Expect = 0.15
Identities = 12/41 (29%), Positives = 21/41 (50%)
Frame = +3
Query: 217 QCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
+C G GK+ +L + F DY PT + ++AN++
Sbjct: 192 KCVTVGDGAVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV 314
>TC229274 homologue to UP|RAC2_LOTJA (Q40220) RAC-like GTP binding protein
RAC2, complete
Length = 973
Score = 33.9 bits (76), Expect = 0.15
Identities = 13/55 (23%), Positives = 26/55 (46%)
Frame = +2
Query: 203 SEDRKNQKTERNVFQCYVFGSKTAGKSALLRALLGRPFSNDYTPTTVERYAANII 257
++ R + + +C G GK+ +L + F DY PT + ++AN++
Sbjct: 131 NQKRSLKMSTARFIKCVTVGYVVVGKTCMLISYTSNTFPTDYVPTVFDNFSANVV 295
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.135 0.390
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,072,301
Number of Sequences: 63676
Number of extensions: 194630
Number of successful extensions: 868
Number of sequences better than 10.0: 44
Number of HSP's better than 10.0 without gapping: 866
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 867
length of query: 432
length of database: 12,639,632
effective HSP length: 100
effective length of query: 332
effective length of database: 6,272,032
effective search space: 2082314624
effective search space used: 2082314624
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135317.6


