

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135315.2 - phase: 0
(478 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
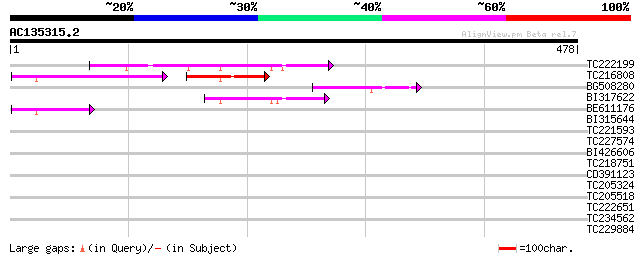
Sequences producing significant alignments: (bits) Value
TC222199 106 2e-23
TC216808 61 3e-21
BG508280 59 6e-09
BI317622 51 1e-06
BE611176 45 5e-05
BI315644 39 0.004
TC221593 similar to UP|Q6NN02 (Q6NN02) At4g14450, partial (30%) 32 0.62
TC227574 similar to UP|RRS1_ARATH (Q9SH88) Ribosome biogenesis r... 32 0.81
BI426606 32 0.81
TC218751 similar to UP|Q6W6U5 (Q6W6U5) Olfactory receptor family... 31 1.4
CD391123 30 3.1
TC205324 similar to UP|Q8L7S4 (Q8L7S4) At1g68060/T23K23_9, parti... 29 5.3
TC205518 similar to UP|SERA_ARATH (O04130) D-3-phosphoglycerate ... 29 5.3
TC222651 homologue to UP|Q9B8W1 (Q9B8W1) NADH dehydrogenase subu... 28 6.9
TC234562 similar to UP|HT22_ARATH (P46604) Homeobox-leucine zipp... 28 6.9
TC229884 weakly similar to UP|Q94AX9 (Q94AX9) At1g72650/F28P22_1... 28 9.0
>TC222199
Length = 691
Score = 106 bits (265), Expect = 2e-23
Identities = 77/231 (33%), Positives = 118/231 (50%), Gaps = 25/231 (10%)
Frame = +2
Query: 68 DFQLVPTLEAYSYLLGSPIAEKTPFTGPGA--SLTPL-VIAKDLYLKTSDVSKHLITKSH 124
DFQLVPT+E + +LG P+ E+ P+ G SL+ + + KD V + T++
Sbjct: 5 DFQLVPTIEEFEEILGCPLGERKPYLSSGCLPSLSRIPTMVKDSARGLDRVKQ---TRNG 175
Query: 125 IRGFTSKYLLVQANLETTCQDALEA--ILALLIYGLILFPNLDNFVDMNAIEIF----HS 178
I G KYL +A D + +LALLI+G++LFPN+D VD+ AI+ F HS
Sbjct: 176 IAGLPRKYLEDKARGIANQGDWVPFMDVLALLIFGVVLFPNVDGLVDLAAIDAFLAYHHS 355
Query: 179 R-NPVPTLLANMYHAIHDRTLKGRGYILCCVPLLYRWFISHL---------PSSFHDNSE 228
+ +PV +LA+++ + R K I+CC+P L W +SHL P H +
Sbjct: 356 KESPVVAVLADLFDTFNRRYEKSSARIICCLPALCVWLVSHLFQQDTRRPCPLLSHRSCT 535
Query: 229 -----DWSYSQRMMALSPNEVVWITPAAQVKE-IITGCGDFLNVPLLGTRG 273
DW Q + + + W + KE ++ CG + N+PL+GTRG
Sbjct: 536 EKRRIDW--DQLLAGIGGRTISWFPRWKEGKEGVLFSCGRYPNIPLVGTRG 682
>TC216808
Length = 1144
Score = 61.2 bits (147), Expect(2) = 3e-21
Identities = 37/134 (27%), Positives = 66/134 (48%), Gaps = 2/134 (1%)
Frame = +2
Query: 2 KKTKSYKFKEVDLVSLRELA--LKVKNQTGFRLRYGGLLTILRTNVEEKLVHTLVQFYDP 59
K+ K K +++ SL EL + + FR YG + + R V + + +L Q+YD
Sbjct: 146 KRFYQVKIKSLEVTSLGELGQLMGQLQR*AFRKAYGKIWDLARVEVSMEPIASLSQYYDQ 325
Query: 60 SFRCFTFPDFQLVPTLEAYSYLLGSPIAEKTPFTGPGASLTPLVIAKDLYLKTSDVSKHL 119
RCFTF DFQLVPT++ + G + + P+ G + +AK + + ++++
Sbjct: 326 PLRCFTFEDFQLVPTVKELEEIRGCQLGGRKPYLFSGFYPSTTRVAKVVKISAQELNRVK 505
Query: 120 ITKSHIRGFTSKYL 133
++ + G K L
Sbjct: 506 QNRNGVVGVPRKCL 547
Score = 58.9 bits (141), Expect(2) = 3e-21
Identities = 33/75 (44%), Positives = 45/75 (60%), Gaps = 5/75 (6%)
Frame = +3
Query: 150 ILALLIYGLILFPNLDNFVDMNAIEIF----HSR-NPVPTLLANMYHAIHDRTLKGRGYI 204
ILALLI+G++LFPN++ VD+ AI+ F H + +PV +LA MY K I
Sbjct: 603 ILALLIFGVVLFPNMEGLVDLAAIDAFLAFHHGKESPVVAILA-MYDTFDRGCEKSSARI 779
Query: 205 LCCVPLLYRWFISHL 219
+CC LY W +SHL
Sbjct: 780 VCCALALYVWLVSHL 824
>BG508280
Length = 418
Score = 58.5 bits (140), Expect = 6e-09
Identities = 34/95 (35%), Positives = 52/95 (53%), Gaps = 3/95 (3%)
Frame = +1
Query: 256 IITGCGDFLNVPLLGTRGGINYNPELAMRQFGFPMKTKPINLATSPEF---FYYSNAPTG 312
++ CG F NVPL+GTRG I+YNP LA++Q G+PM P+ +P F +N T
Sbjct: 70 VLISCGGFPNVPLMGTRGCISYNPVLAIKQLGYPM*GVPLEDELAPVISRGFNKTNVETL 249
Query: 313 QREAFTRAWSKVRRKSVRHLGVRSGIAHEAYTQWV 347
Q+ +AW +++K G +G Y +W+
Sbjct: 250 QK--VRKAWEVLQKKDKELRGSNNG-PIGGYRKWL 345
>BI317622
Length = 370
Score = 51.2 bits (121), Expect = 1e-06
Identities = 37/125 (29%), Positives = 53/125 (41%), Gaps = 20/125 (16%)
Frame = -2
Query: 165 DNFVDMNAIEIF-----HSRNPVPTLLANMYHAIHDRTLKGRGYILCCVPLLYRWFISHL 219
D VD+ AI F + P+ +LA++Y + R K I+CC P LY W +SHL
Sbjct: 369 DGLVDLAAIGAFLAYHDYKEIPIVAMLADLYDTFNRRCEKSSTRIVCCTPALYVWLVSHL 190
Query: 220 ---------PSSFH-----DNSEDWSYSQRMMALSPNEVVWITPAAQVK-EIITGCGDFL 264
P H E+W Q + V W + + ++ CG FL
Sbjct: 189 FRQEVRHACPLESHCSCTEKGGENW--GQLLANKEGASVNWFPRWKEGRTRVLISCGVFL 16
Query: 265 NVPLL 269
NVPL+
Sbjct: 15 NVPLM 1
>BE611176
Length = 424
Score = 45.4 bits (106), Expect = 5e-05
Identities = 24/72 (33%), Positives = 37/72 (51%), Gaps = 2/72 (2%)
Frame = -2
Query: 2 KKTKSYKFKEVDLVSLRELA--LKVKNQTGFRLRYGGLLTILRTNVEEKLVHTLVQFYDP 59
K+ K K +D+ S++EL + FR Y +L + + + + +L Q+YD
Sbjct: 360 KRFYQVKIKGLDITSIKELGRLMGPLQMQAFRKAYEKILDLTIAEISTEAIASLTQYYD* 181
Query: 60 SFRCFTFPDFQL 71
RCFTF DFQL
Sbjct: 180 PLRCFTFGDFQL 145
>BI315644
Length = 421
Score = 39.3 bits (90), Expect = 0.004
Identities = 17/31 (54%), Positives = 23/31 (73%)
Frame = +3
Query: 271 TRGGINYNPELAMRQFGFPMKTKPINLATSP 301
TRG INYNP LA+RQ G+PM+ P + + +P
Sbjct: 276 TRGCINYNPVLAIRQLGYPMRGVPSDESITP 368
>TC221593 similar to UP|Q6NN02 (Q6NN02) At4g14450, partial (30%)
Length = 514
Score = 32.0 bits (71), Expect = 0.62
Identities = 33/119 (27%), Positives = 51/119 (42%), Gaps = 7/119 (5%)
Frame = +1
Query: 287 GFPMKTKPINLATSPEFFYYS-------NAPTGQREAFTRAWSKVRRKSVRHLGVRSGIA 339
G P T +L+ + + F Y +AP E+ +R RRK R +
Sbjct: 13 GAPYFTFQFSLSHTLKHFSYFLSLRRRYHAPPHMAESHSRTRPS-RRKPSR-------LQ 168
Query: 340 HEAYTQWVINRAEEIGMPYPAMRHVAASAPSIPLPLPPTTQEMYQEHLAMESRKKQMWK 398
A + INRA E + P + +A+S P PLPLPP ++ Q+ E + W+
Sbjct: 169 SRAPSSLQINRALEWNVAIPLLSPLASSPP--PLPLPPQKEDPPQQQRPTEKVVFKKWQ 339
>TC227574 similar to UP|RRS1_ARATH (Q9SH88) Ribosome biogenesis regulatory
protein homolog, partial (50%)
Length = 803
Score = 31.6 bits (70), Expect = 0.81
Identities = 27/86 (31%), Positives = 39/86 (44%), Gaps = 14/86 (16%)
Frame = +3
Query: 371 IPLPLPPTTQEMYQEHLAMESRKKQ---------MWKAR--YNEA--ENLIMTLDGKDEQ 417
+P P PPT E + + ++ RKK WK R Y+ A E+ I ++ K
Sbjct: 375 LPKPKPPTKWEAFAKKKGIQKRKKDKIVYDEQSGTWKRRHGYDRANDEDAIPIIEAKPTD 554
Query: 418 KTHENLMLK-KELVKVRRELEEKDEL 442
E+ K KE K R E +EK+ L
Sbjct: 555 DPEEDPFAKRKENKKNRIEKQEKNRL 632
>BI426606
Length = 357
Score = 31.6 bits (70), Expect = 0.81
Identities = 28/103 (27%), Positives = 50/103 (48%), Gaps = 11/103 (10%)
Frame = +3
Query: 352 EEIGMPYPAMRHVAASAPSI----PLPLPPTTQEMYQEHLAMESR--KKQMWKAR----- 400
EE+ M RH+ ++ LPLPP + ++ H + R K Q+ + R
Sbjct: 18 EEVTMSKLRHRHLRRHLENLHRHRQLPLPPLLRPLHPHHKLLLPRHPKVQLLRHRLHLRL 197
Query: 401 YNEAENLIMTLDGKDEQKTHENLMLKKELVKVRRELEEKDELL 443
++EAENL M +DG + ++ EL +R + +KD+++
Sbjct: 198 FSEAENLHMNIDG-------DRFLVSNEL--LRSKPCDKDDII 299
>TC218751 similar to UP|Q6W6U5 (Q6W6U5) Olfactory receptor family 8 subfamily
S (Fragment), partial (7%)
Length = 988
Score = 30.8 bits (68), Expect = 1.4
Identities = 20/60 (33%), Positives = 32/60 (53%), Gaps = 4/60 (6%)
Frame = +2
Query: 384 QEHLAMESR----KKQMWKARYNEAENLIMTLDGKDEQKTHENLMLKKELVKVRRELEEK 439
Q +LA ES K + ++ EAE I L+ + + T E +LKKEL ++R++ K
Sbjct: 491 QSNLANESEELKSKLSVMDSKLREAEVTITKLNEERRRNTQEKDLLKKELEVLKRQINTK 670
>CD391123
Length = 613
Score = 29.6 bits (65), Expect = 3.1
Identities = 20/60 (33%), Positives = 31/60 (51%), Gaps = 4/60 (6%)
Frame = -3
Query: 384 QEHLAMESR----KKQMWKARYNEAENLIMTLDGKDEQKTHENLMLKKELVKVRRELEEK 439
Q +LA ES K + ++ EAE I L+ + + T E +LKKEL ++R + K
Sbjct: 470 QSNLAHESEELKSKLSVMDSKLREAEVTITKLNEERRRNTQEKDLLKKELEVLKRPIHTK 291
>TC205324 similar to UP|Q8L7S4 (Q8L7S4) At1g68060/T23K23_9, partial (19%)
Length = 980
Score = 28.9 bits (63), Expect = 5.3
Identities = 19/69 (27%), Positives = 33/69 (47%), Gaps = 4/69 (5%)
Frame = +2
Query: 379 TQEMYQEHLAMESRKKQM----WKARYNEAENLIMTLDGKDEQKTHENLMLKKELVKVRR 434
T Y + +ME K++ WK ++ N T+D D + L+KE++ +R+
Sbjct: 74 TPPSYSFNQSMEETKEREASDNWKGNSDDKPNDFPTVDSVDSVPSVLYDFLQKEVIALRK 253
Query: 435 ELEEKDELL 443
EKD+ L
Sbjct: 254 AGHEKDQSL 280
>TC205518 similar to UP|SERA_ARATH (O04130) D-3-phosphoglycerate
dehydrogenase, chloroplast precursor (3-PGDH) , partial
(70%)
Length = 1666
Score = 28.9 bits (63), Expect = 5.3
Identities = 22/79 (27%), Positives = 42/79 (52%), Gaps = 2/79 (2%)
Frame = +1
Query: 317 FTRAWSKVRRKSVRHLGVRSGIAHEAYTQWVINRAEEIGMPYPAMRHVAASAP--SIPLP 374
F + S+V R++ + LG+ IAH+ Y +RA IG+ + +A S+ +P
Sbjct: 211 FGKVGSEVARRA-KGLGMHV-IAHDPYAP--ADRARAIGVDLVSFDQAITTADFISLHMP 378
Query: 375 LPPTTQEMYQEHLAMESRK 393
L PTT +++ ++ + +K
Sbjct: 379 LTPTTNKIFNDNTFAKMKK 435
>TC222651 homologue to UP|Q9B8W1 (Q9B8W1) NADH dehydrogenase subunit 6,
partial (8%)
Length = 1006
Score = 28.5 bits (62), Expect = 6.9
Identities = 21/85 (24%), Positives = 42/85 (48%), Gaps = 10/85 (11%)
Frame = -1
Query: 212 YRWFISHLPSSFHDNSEDWSYSQRMMA-LSPNEVVWIT---------PAAQVKEIITGCG 261
Y W I +L FH N+E+ +YS +++ ++++I+ +K I+
Sbjct: 628 YLWLIKYL*KDFHINNENQTYSYKILIYFLSTKLIFISIK*ILKTNLKIIYIK*ILKMNL 449
Query: 262 DFLNVPLLGTRGGINYNPELAMRQF 286
+ L +PL G N+N ++++R F
Sbjct: 448 ETLLLPL-----GSNFNNDISLRDF 389
>TC234562 similar to UP|HT22_ARATH (P46604) Homeobox-leucine zipper protein
HAT22 (HD-ZIP protein 22), partial (33%)
Length = 585
Score = 28.5 bits (62), Expect = 6.9
Identities = 19/67 (28%), Positives = 33/67 (48%), Gaps = 6/67 (8%)
Frame = +3
Query: 375 LPPTTQEMYQEHLAMESRKKQMW------KARYNEAENLIMTLDGKDEQKTHENLMLKKE 428
L P ++ E L ++ R+ ++W + + + E L E+ T ENL LKKE
Sbjct: 366 LNPAQKQALAEQLNLKHRQVEVWFQNRRARTKLKQTEVDCEFLKKCCEKLTDENLRLKKE 545
Query: 429 LVKVRRE 435
L ++R +
Sbjct: 546 LQELRAQ 566
>TC229884 weakly similar to UP|Q94AX9 (Q94AX9) At1g72650/F28P22_16 (MYB
transcription factor), partial (20%)
Length = 1244
Score = 28.1 bits (61), Expect = 9.0
Identities = 15/47 (31%), Positives = 25/47 (52%), Gaps = 1/47 (2%)
Frame = -3
Query: 154 LIYGLILFPNLDNFVDMNAIEIFHSR-NPVPTLLANMYHAIHDRTLK 199
+I+ LI PN + +A FH++ NP P LL ++ A+ L+
Sbjct: 384 MIFQLICIPNSVQCLQQHAFHCFHTQENPPPPLLHHLLQALTSHHLQ 244
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.136 0.405
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,126,690
Number of Sequences: 63676
Number of extensions: 301201
Number of successful extensions: 1903
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1887
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1895
length of query: 478
length of database: 12,639,632
effective HSP length: 101
effective length of query: 377
effective length of database: 6,208,356
effective search space: 2340550212
effective search space used: 2340550212
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC135315.2


