

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135311.8 + phase: 0
(189 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
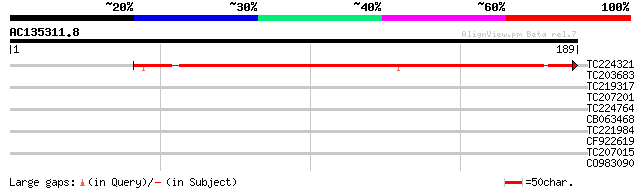
Score E
Sequences producing significant alignments: (bits) Value
TC224321 similar to UP|Q6H5X0 (Q6H5X0) Ripening-related protein-... 160 4e-40
TC203683 homologue to UP|R37A_GOSHI (Q9XHE4) 60S ribosomal prote... 35 0.016
TC219317 similar to GB|AAP13385.1|30023704|BT006277 At3g56360 {A... 30 0.67
TC207201 30 0.87
TC224764 weakly similar to PIR|A96697|A96697 protein F1N21.18 [i... 28 3.3
CB063468 28 3.3
TC221984 similar to GB|AAP13431.1|30023796|BT006323 At5g02530 {A... 27 5.6
CF922619 27 5.6
TC207015 weakly similar to UP|Q8LAH2 (Q8LAH2) Polygalacturonase-... 26 9.6
CO983090 26 9.6
>TC224321 similar to UP|Q6H5X0 (Q6H5X0) Ripening-related protein-like,
partial (52%)
Length = 743
Score = 160 bits (404), Expect = 4e-40
Identities = 83/151 (54%), Positives = 99/151 (64%), Gaps = 3/151 (1%)
Frame = +3
Query: 42 PP--GKCNQENDSDCCVQGKMYTTYVCSPSVSTHTKAYLTLNSFQKGGDGGGPSECDKQY 99
PP G C +C GK Y Y CSP V++ T A LTLN F +GGDGG PS+CD+++
Sbjct: 222 PPSGGSCQSSGTLNC--GGKSYPQYRCSPPVTSSTPAILTLNDFSEGGDGGDPSQCDEKF 395
Query: 100 HSDDTPVVALSTGWFNHKSRCLNNITISA-NGRNVVAMVVDECDSRKGCDEQHDYQPPCP 158
H++ VVALSTGWFN S CL I I+A NGRNV A VVD+CDS GCD H QPPC
Sbjct: 396 HNNTERVVALSTGWFNGSSLCLKMIRITAGNGRNVTAKVVDQCDSVNGCDRAHAGQPPCR 575
Query: 159 NNIVDASKAVWKALGVPKEQWGGLDITWSDA 189
NNIVD S+AVW ALG+ + G +TWS A
Sbjct: 576 NNIVDGSQAVWDALGLDSDV-GEARVTWSMA 665
>TC203683 homologue to UP|R37A_GOSHI (Q9XHE4) 60S ribosomal protein L37a,
complete
Length = 1919
Score = 35.4 bits (80), Expect = 0.016
Identities = 14/54 (25%), Positives = 24/54 (43%)
Frame = +3
Query: 126 ISANGRNVVAMVVDECDSRKGCDEQHDYQPPCPNNIVDASKAVWKALGVPKEQW 179
I +N +V ++ C + C H Y PCP ++ ++ + WK E W
Sbjct: 87 IVSNNNKLVDGFLNSCQGQSNCSCHHHY*DPCPKLLLKSAGSKWKNKNTEPETW 248
>TC219317 similar to GB|AAP13385.1|30023704|BT006277 At3g56360 {Arabidopsis
thaliana;} , partial (33%)
Length = 829
Score = 30.0 bits (66), Expect = 0.67
Identities = 16/36 (44%), Positives = 19/36 (52%), Gaps = 1/36 (2%)
Frame = +2
Query: 86 GGDGGGPSECDKQYHSDDTPVVAL-STGWFNHKSRC 120
GGD G E + YHSDD A+ T W + K RC
Sbjct: 245 GGDDHGHGEGARDYHSDDDGNGAVRETRWRSQKRRC 352
>TC207201
Length = 1431
Score = 29.6 bits (65), Expect = 0.87
Identities = 20/71 (28%), Positives = 31/71 (43%), Gaps = 2/71 (2%)
Frame = -3
Query: 65 VCSPSVSTHTKAYLTLNSFQKGGDGGGPSECDKQYHSDD--TPVVALSTGWFNHKSRCLN 122
+C+PS TH+ YLT N+ + G+ PS H P++ + T K+
Sbjct: 952 LCNPSTPTHSPHYLT-NTQKIAGESIRPSNAVTSTHCKHFWVPIIIVITMIICFKATSFE 776
Query: 123 NITISANGRNV 133
N +S N V
Sbjct: 775 NGRVSTNSERV 743
>TC224764 weakly similar to PIR|A96697|A96697 protein F1N21.18 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(38%)
Length = 1766
Score = 27.7 bits (60), Expect = 3.3
Identities = 10/26 (38%), Positives = 17/26 (64%)
Frame = -2
Query: 5 CPKVSFIFLAITLIFTSFVYSEAQNC 30
CP +F ++ + L+ +SFV S+ NC
Sbjct: 622 CPGCNFRYIIVFLLGSSFVLSQVSNC 545
>CB063468
Length = 410
Score = 27.7 bits (60), Expect = 3.3
Identities = 14/25 (56%), Positives = 17/25 (68%), Gaps = 1/25 (4%)
Frame = -1
Query: 59 KMYTTYVCSPSVSTHTKAY-LTLNS 82
K +Y CS S+S HTK Y L+LNS
Sbjct: 158 KQTNSYTCSLSLSLHTKLY*LSLNS 84
>TC221984 similar to GB|AAP13431.1|30023796|BT006323 At5g02530 {Arabidopsis
thaliana;} , partial (26%)
Length = 554
Score = 26.9 bits (58), Expect = 5.6
Identities = 16/45 (35%), Positives = 20/45 (43%), Gaps = 3/45 (6%)
Frame = +3
Query: 28 QNCRPSGRIRGKKAPPGKCNQENDSDCCVQG---KMYTTYVCSPS 69
Q RPS R + PP C Q + C QG + T YV P+
Sbjct: 159 QGPRPSLGTRTRSPPPQPCRQSRRTVCRCQGAGDSLETRYVREPA 293
>CF922619
Length = 562
Score = 26.9 bits (58), Expect = 5.6
Identities = 12/25 (48%), Positives = 15/25 (60%)
Frame = +2
Query: 11 IFLAITLIFTSFVYSEAQNCRPSGR 35
IFL + FTSF+ S RP+GR
Sbjct: 317 IFLLLYYYFTSFLISNVVTARPNGR 391
>TC207015 weakly similar to UP|Q8LAH2 (Q8LAH2) Polygalacturonase-like
protein, partial (22%)
Length = 733
Score = 26.2 bits (56), Expect = 9.6
Identities = 19/58 (32%), Positives = 28/58 (47%), Gaps = 3/58 (5%)
Frame = +1
Query: 120 CLNNITISANGRNVVAMVVDECDSRKGCDEQHDYQPPCP---NNIVDASKAVWKALGV 174
CL+NIT+S N V+ + EC + G + PCP N D+S + + L V
Sbjct: 181 CLSNITLSTNS---VSPITWECSNVSGFSDS-VLPEPCPELGNPSYDSSSSCFYLLSV 342
>CO983090
Length = 830
Score = 26.2 bits (56), Expect = 9.6
Identities = 9/21 (42%), Positives = 13/21 (61%)
Frame = +2
Query: 46 CNQENDSDCCVQGKMYTTYVC 66
CN++N + C G+M T Y C
Sbjct: 392 CNEQNQTTRCSLGRMTTIYQC 454
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.134 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,376,901
Number of Sequences: 63676
Number of extensions: 174333
Number of successful extensions: 1047
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1044
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1046
length of query: 189
length of database: 12,639,632
effective HSP length: 92
effective length of query: 97
effective length of database: 6,781,440
effective search space: 657799680
effective search space used: 657799680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 56 (26.2 bits)
Medicago: description of AC135311.8


