

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135311.2 + phase: 0
(164 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
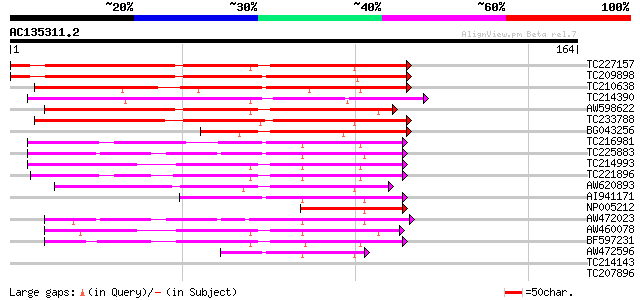
TC227157 similar to UP|Q40329 (Q40329) Serine proteinase inhibit... 134 1e-32
TC209898 weakly similar to UP|Q40329 (Q40329) Serine proteinase ... 122 8e-29
TC210638 weakly similar to PIR|C01300|TIZB1P proteinase inhibito... 109 5e-25
TC214390 homologue to UP|Q9AVS2 (Q9AVS2) Trypsin inhibitor precu... 93 5e-20
AW598622 weakly similar to PIR|S56648|S566 trypsin inhibitor pre... 90 4e-19
TC233788 similar to UP|Q40330 (Q40330) Serine proteinase inhibit... 87 4e-18
BG043256 similar to PIR|S56647|S566 trypsin inhibitor precursor ... 75 2e-14
TC216981 UP|Q8RU22 (Q8RU22) Bowman-Birk type proteinase isoinhib... 74 4e-14
TC225883 UP|Q8RU24 (Q8RU24) Bowman-Birk type proteinase isoinhib... 73 5e-14
TC214993 UP|Q8RU21 (Q8RU21) Bowman-Birk type proteinase isoinhib... 73 5e-14
TC221896 UP|Q8RU21 (Q8RU21) Bowman-Birk type proteinase isoinhib... 72 1e-13
AW620893 weakly similar to PIR|S56648|S566 trypsin inhibitor pre... 69 8e-13
AI941171 homologue to GP|19698251|dbj Bowman-Birk type proteinas... 60 5e-10
NP005212 Bowman-Birk protease inhibitor 53 7e-08
AW472023 similar to GP|19698245|db Bowman-Birk type proteinase i... 45 2e-05
AW460078 similar to GP|19698251|dbj Bowman-Birk type proteinase ... 45 2e-05
BF597231 similar to GP|19698251|db Bowman-Birk type proteinase i... 44 5e-05
AW472596 similar to GP|19698249|dbj Bowman-Birk type proteinase ... 43 6e-05
TC214143 UP|Q39832 (Q39832) Ribulose-1,5-bisphosphate carboxylas... 32 0.10
TC207896 GB|BAA13184.1|1498170|D86929 uricase {Glycine max;} , c... 32 0.14
>TC227157 similar to UP|Q40329 (Q40329) Serine proteinase inhibitor, partial
(89%)
Length = 769
Score = 134 bits (338), Expect = 1e-32
Identities = 69/119 (57%), Positives = 86/119 (71%), Gaps = 3/119 (2%)
Frame = +2
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MEL K +++K+A L+FLLGFT+TVVDARFD +SFITQ + N ++ NNY KST
Sbjct: 212 MELSMK----VLVKVASLLFLLGFTATVVDARFDPSSFITQFLPNAEA--NNYYVKSTTK 373
Query: 61 ACCNTCLC--SVTPELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
ACCN+C C S+ P QCRC+D+G+TCHSACK C C S P C C DIT+FCY+PCN
Sbjct: 374 ACCNSCPCTKSIPP---QCRCSDIGETCHSACKTCICTRSIPPQCHCSDITNFCYEPCN 541
>TC209898 weakly similar to UP|Q40329 (Q40329) Serine proteinase inhibitor,
partial (83%)
Length = 518
Score = 122 bits (306), Expect = 8e-29
Identities = 59/118 (50%), Positives = 82/118 (69%), Gaps = 2/118 (1%)
Frame = +3
Query: 1 MELMNKNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTAT 60
MEL K +++K+A L+FLLGFT+TVVDARFD +SFITQ + N ++ NNY KST
Sbjct: 42 MELSMK----VLVKVASLLFLLGFTATVVDARFDPSSFITQFLPNAEA--NNYYVKSTTN 203
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCECGWSS--PLCTCYDITDFCYKPCN 116
CC+ C C+++ C+CAD+G+TCH +CK+C C + LC C D+T+FCY+ CN
Sbjct: 204 GCCDNCRCTISIS-PMCKCADIGETCHPSCKSCFCDIPTFPGLCQCIDVTNFCYELCN 374
>TC210638 weakly similar to PIR|C01300|TIZB1P proteinase inhibitor
(Bowman-Birk) I-A'' - adzuki bean {Vigna angularis;} ,
partial (47%)
Length = 513
Score = 109 bits (273), Expect = 5e-25
Identities = 61/114 (53%), Positives = 73/114 (63%), Gaps = 5/114 (4%)
Frame = +1
Query: 8 KKAMVMKLALLVFLLGFTSTVVDA-RFDSTSFITQVISNGDSTTNNY--DAKSTATACCN 64
KK ++K+ALL+FLLGF++T DA RFD SFITQV+ NNY KSTATACC+
Sbjct: 4 KKVALVKVALLLFLLGFSATTADATRFDPRSFITQVL------PNNYYVKIKSTATACCD 165
Query: 65 TCLCSVTPELTQCRCADVGKT-CHSACKNCECGWSSP-LCTCYDITDFCYKPCN 116
CLC+ + QC C D +T CHS CKNC C P C C DIT+FCY CN
Sbjct: 166 LCLCTKSYP-PQCNCVDESETGCHSCCKNCICNKKFPRTCYCSDITNFCYDKCN 324
>TC214390 homologue to UP|Q9AVS2 (Q9AVS2) Trypsin inhibitor precursor,
partial (14%)
Length = 824
Score = 93.2 bits (230), Expect = 5e-20
Identities = 49/121 (40%), Positives = 70/121 (57%), Gaps = 5/121 (4%)
Frame = +2
Query: 6 KNKKAMVMKLALLVFLLGFTSTVVDAR-FDSTSFITQVISNGDSTTNNYDAKSTATACCN 64
K ++ + K+ L++FLLGFT+T VDA F+ S ITQ++ + +++ KS CC+
Sbjct: 188 KVREMELKKVGLVLFLLGFTATTVDATGFNPNSLITQLLPQIEGDSDDNYVKSNTEPCCD 367
Query: 65 TCLC--SVTPELTQCRCADVGKTCHSACKNCECG--WSSPLCTCYDITDFCYKPCNVWQQ 120
CLC S+ P+ C+C D + CHSACK C C W P C C+D TD CY C Q
Sbjct: 368 NCLCTNSIPPK---CQCTDWAEDCHSACKGCICQRIW-PPRCRCFDETDTCYDKCTSTSQ 535
Query: 121 Q 121
+
Sbjct: 536 E 538
>AW598622 weakly similar to PIR|S56648|S566 trypsin inhibitor precursor
(clone ATI21) - alfalfa, partial (80%)
Length = 316
Score = 90.1 bits (222), Expect = 4e-19
Identities = 53/105 (50%), Positives = 70/105 (66%), Gaps = 3/105 (2%)
Frame = +2
Query: 11 MVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTCLC-- 68
+++K+A ++FLLGFT+T VDARF + FITQ + N +S NN +ST ACCN+C C
Sbjct: 17 VLVKVATVLFLLGFTATDVDARFYPS*FITQFLPNAES--NNCYVESTTKACCNSCPCTK 190
Query: 69 SVTPELTQCRCADVGKTCHSACKNCECGWSSPLCTCY-DITDFCY 112
S+ P QCR +DVG+TC SACK C S P + Y DIT+F Y
Sbjct: 191 SIPP---QCRGSDVGETCDSACKT*MCTHSIPPQSHYSDITNFYY 316
>TC233788 similar to UP|Q40330 (Q40330) Serine proteinase inhibitor, partial
(57%)
Length = 456
Score = 87.0 bits (214), Expect = 4e-18
Identities = 47/111 (42%), Positives = 67/111 (60%), Gaps = 2/111 (1%)
Frame = +3
Query: 8 KKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTCL 67
KK +++K+ALL + FT++ VDAR S + + +I+ +NY+ KST +ACC+ C
Sbjct: 63 KKLVLVKVALLFLFICFTASNVDARSRSINPGSFIIAG-----DNYNLKSTTSACCDACA 227
Query: 68 CSVT-PELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFCYKPCN 116
C+ + P + C C D G+TCHSAC C C S P C C D T FCY C+
Sbjct: 228 CTKSIPPI--CHCHDFGETCHSACNLCICTASYPPQCRCLDQTTFCYDKCD 374
>BG043256 similar to PIR|S56647|S566 trypsin inhibitor precursor (clone
ATI18) - alfalfa, partial (51%)
Length = 196
Score = 74.7 bits (182), Expect = 2e-14
Identities = 35/64 (54%), Positives = 43/64 (66%), Gaps = 3/64 (4%)
Frame = +2
Query: 56 KSTATACCNT--CLCSVTPELTQCRCADVGKTCHSACKNCEC-GWSSPLCTCYDITDFCY 112
KST ACCN+ C S+ P QCRC D+G+TCHSACK C G + C C DIT+FCY
Sbjct: 14 KSTTKACCNSWPCTKSIPP---QCRCTDIGETCHSACKT*MCTGSMATQCHCNDITNFCY 184
Query: 113 KPCN 116
+PC+
Sbjct: 185 EPCS 196
>TC216981 UP|Q8RU22 (Q8RU22) Bowman-Birk type proteinase isoinhibitor C
precursor, complete
Length = 508
Score = 73.6 bits (179), Expect = 4e-14
Identities = 41/112 (36%), Positives = 58/112 (51%), Gaps = 2/112 (1%)
Frame = +1
Query: 6 KNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNT 65
KN ++ L++FL+G T+ AR + F + S+ D ++ CC+
Sbjct: 37 KNNMVVLKVCFLVLFLVGVTN----ARMELNLFKSDHSSSDDESSK---------PCCDL 177
Query: 66 CLCSVTPELTQCRCADVG-KTCHSACKNCECGWSSP-LCTCYDITDFCYKPC 115
C+C+ + QC CAD+ +CHSAC C C S P C C D TDFCYKPC
Sbjct: 178 CMCTASMP-PQCHCADIRLNSCHSACDRCACTRSMPGQCRCLDTTDFCYKPC 330
>TC225883 UP|Q8RU24 (Q8RU24) Bowman-Birk type proteinase isoinhibitor A1
precursor, complete
Length = 596
Score = 73.2 bits (178), Expect = 5e-14
Identities = 42/112 (37%), Positives = 62/112 (54%), Gaps = 2/112 (1%)
Frame = +2
Query: 6 KNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNT 65
KN ++ +L+FL+G T T + R + + D ++ D +S+ CC+
Sbjct: 38 KNNMVVLKVCLVLLFLVGGT-TSANLRLSKLGLLMK----SDHHQHSNDDESSKP-CCDQ 199
Query: 66 CLCSVTPELTQCRCADVG-KTCHSACKNCECGWSSPL-CTCYDITDFCYKPC 115
C C+ + QCRC+D+ +CHSACK+C C S P C C DITDFCY+PC
Sbjct: 200 CACTKSNP-PQCRCSDMRLNSCHSACKSCICALSYPAQCFCVDITDFCYEPC 352
Score = 30.8 bits (68), Expect = 0.30
Identities = 11/34 (32%), Positives = 19/34 (55%)
Frame = +2
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCE 94
+ C +C+C+++ QC C D+ C+ CK E
Sbjct: 266 SACKSCICALSYP-AQCFCVDITDFCYEPCKPSE 364
>TC214993 UP|Q8RU21 (Q8RU21) Bowman-Birk type proteinase isoinhibitor D
precursor, complete
Length = 563
Score = 73.2 bits (178), Expect = 5e-14
Identities = 40/114 (35%), Positives = 58/114 (50%), Gaps = 4/114 (3%)
Frame = +2
Query: 6 KNKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNT 65
KN ++ LL+FL+G T+ ++ F + +++YD + CC+
Sbjct: 89 KNNMVVLKVCLLLLFLVGVTAARMELSFFKSD-----------QSSSYDDDEYSKPCCDL 235
Query: 66 CLC--SVTPELTQCRCADVG-KTCHSACKNCECGWSSP-LCTCYDITDFCYKPC 115
C+C S+ P QC C D+ +CHS CK+C C S P C C D DFCYKPC
Sbjct: 236 CMCTRSMPP---QCSCEDIRLNSCHSDCKSCMCTRSQPGQCRCLDTNDFCYKPC 388
>TC221896 UP|Q8RU21 (Q8RU21) Bowman-Birk type proteinase isoinhibitor D
precursor, partial (95%)
Length = 449
Score = 72.0 bits (175), Expect = 1e-13
Identities = 42/113 (37%), Positives = 58/113 (51%), Gaps = 4/113 (3%)
Frame = +1
Query: 7 NKKAMVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC 66
N ++ LL+FL+G T+ AR SF+ +++YD + CC+ C
Sbjct: 1 NNMVVLKVCLLLLFLVGVTA----ARMCILSFLKS------DQSSSYDDDEYSKPCCDLC 150
Query: 67 LC--SVTPELTQCRCADVG-KTCHSACKNCECGWSSP-LCTCYDITDFCYKPC 115
+C S+ P QC C D+ +CHS CK+C C S P C C D DFCYKPC
Sbjct: 151 MCTRSMPP---QCSCEDIRLNSCHSDCKSCMCTRSQPGQCRCLDTNDFCYKPC 300
>AW620893 weakly similar to PIR|S56648|S566 trypsin inhibitor precursor
(clone ATI21) - alfalfa, partial (53%)
Length = 313
Score = 69.3 bits (168), Expect = 8e-13
Identities = 43/101 (42%), Positives = 58/101 (56%), Gaps = 3/101 (2%)
Frame = +2
Query: 14 KLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTC--LCSVT 71
K+A ++FL G T+ V ARFD S ++ + D T NY KSTA+ACC +C S+
Sbjct: 26 KVAAVLFLWGCTAAVGGARFDPRSCVSGGVHCAD--TGNYYVKSTASACCGSCAGTKSIR 199
Query: 72 PELTQCRCADVGKTCHSACKNCECGWS-SPLCTCYDITDFC 111
P Q RC D+G+T HSAC+ C S +P C D+ FC
Sbjct: 200 P---QWRC*DIGRTWHSACQTGICTRSIAPQGHCADVARFC 313
>AI941171 homologue to GP|19698251|dbj Bowman-Birk type proteinase
isoinhibitor D {Glycine soja}, partial (81%)
Length = 267
Score = 60.1 bits (144), Expect = 5e-10
Identities = 27/68 (39%), Positives = 39/68 (56%), Gaps = 2/68 (2%)
Frame = +1
Query: 50 TNNYDAKSTATACCNTCLCSVTPELTQCRCADVG-KTCHSACKNCECGWSSPL-CTCYDI 107
+++YD + CC C+C+ + +QC C D+ +CHS CK+ C S + C C D
Sbjct: 52 SSSYDDDEYSKPCCELCMCTRSMP-SQCGCEDIRLNSCHSDCKS*SCTRSQAVQCRCLDT 228
Query: 108 TDFCYKPC 115
DFCYKPC
Sbjct: 229 NDFCYKPC 252
>NP005212 Bowman-Birk protease inhibitor
Length = 129
Score = 52.8 bits (125), Expect = 7e-08
Identities = 21/32 (65%), Positives = 24/32 (74%), Gaps = 1/32 (3%)
Frame = +1
Query: 85 TCHSACKNCECGWSSPL-CTCYDITDFCYKPC 115
+CHSACK+C C S P C C DITDFCY+PC
Sbjct: 4 SCHSACKSCICALSYPAQCFCVDITDFCYEPC 99
Score = 30.8 bits (68), Expect = 0.30
Identities = 11/34 (32%), Positives = 19/34 (55%)
Frame = +1
Query: 61 ACCNTCLCSVTPELTQCRCADVGKTCHSACKNCE 94
+ C +C+C+++ QC C D+ C+ CK E
Sbjct: 13 SACKSCICALSYP-AQCFCVDITDFCYEPCKPSE 111
>AW472023 similar to GP|19698245|db Bowman-Birk type proteinase isoinhibitor
A1 {Glycine soja}, partial (93%)
Length = 331
Score = 45.1 bits (105), Expect = 2e-05
Identities = 36/110 (32%), Positives = 55/110 (49%), Gaps = 3/110 (2%)
Frame = +2
Query: 11 MVMKLAL-LVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTCLCS 69
+V+K+ L L+FL+G T T + R + + D ++ D +S+ CC+ C C
Sbjct: 2 VVLKVFLVLLFLVGGT-TSANLRLSKLGLLMK----SDHHQHSNDDESSKP-CCDQCAC- 160
Query: 70 VTPELTQCRCADVG-KTCHSACKNCECGWSSPL-CTCYDITDFCYKPCNV 117
+ C D+ + HSAC + C S P C C DITDF Y+PC +
Sbjct: 161 IKSNPPYCLW*DLMLNSSHSACISLICALSYPAQCLCDDITDFYYEPCKL 310
>AW460078 similar to GP|19698251|dbj Bowman-Birk type proteinase isoinhibitor
D {Glycine soja}, partial (91%)
Length = 298
Score = 45.1 bits (105), Expect = 2e-05
Identities = 33/109 (30%), Positives = 56/109 (51%), Gaps = 5/109 (4%)
Frame = +2
Query: 11 MVMKLALLV-FLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTCLC- 68
+V+K+ LL+ FL+G T+ ++ F + +++YD + + CCN +
Sbjct: 2 VVLKVCLLLLFLVGVTAAHMELIFFKSD-----------QSSSYDNNNYSKPCCNLYMYT 148
Query: 69 -SVTPELTQCRCADVG-KTCHSACKNCECGWSSPLCTCY-DITDFCYKP 114
S+ P QC C ++ +CHS CK+C C S P+ Y + +F YKP
Sbjct: 149 HSMPP---QCICKNIKLNSCHSYCKSCMCTHSHPIHYHYLNTNNFYYKP 286
>BF597231 similar to GP|19698251|db Bowman-Birk type proteinase isoinhibitor
D {Glycine soja}, partial (93%)
Length = 304
Score = 43.5 bits (101), Expect = 5e-05
Identities = 33/109 (30%), Positives = 54/109 (49%), Gaps = 4/109 (3%)
Frame = +2
Query: 11 MVMKLALLVFLLGFTSTVVDARFDSTSFITQVISNGDSTTNNYDAKSTATACCNTCLC-- 68
+++K+ LL+ +L + V AR + + F + +++YD +A C + C+C
Sbjct: 2 VMLKVPLLMIIL---AAVTVARMELSFFKSD-------HSSSYDDDESAIPCWDLCMCTR 151
Query: 69 SVTPELTQCRCADVGK-TCHSACKNCECGWSSP-LCTCYDITDFCYKPC 115
S+ P CR D+ +CHS ++C C S P C C D YKPC
Sbjct: 152 SLPP---PCR*EDIRLCSCHSDGQSCMCTRSQPGQCRCLDTN*LRYKPC 289
>AW472596 similar to GP|19698249|dbj Bowman-Birk type proteinase isoinhibitor
C {Glycine soja}, partial (57%)
Length = 190
Score = 43.1 bits (100), Expect = 6e-05
Identities = 20/45 (44%), Positives = 27/45 (59%), Gaps = 2/45 (4%)
Frame = +2
Query: 62 CCNTCLCSVTPELTQCRCADVG-KTCHSACKNCECGWS-SPLCTC 104
CC+ C+C+ + QC CAD+ +C SAC C+C S S LC C
Sbjct: 59 CCDLCMCTESMP-PQCHCADITLNSCDSACDYCDCSHSMSGLCRC 190
>TC214143 UP|Q39832 (Q39832) Ribulose-1,5-bisphosphate carboxylase small
subunit precursor (Ribulose-1,5-bisphosphate carboxylase
small subunit rbcS1), complete
Length = 1249
Score = 32.3 bits (72), Expect = 0.10
Identities = 15/40 (37%), Positives = 20/40 (49%)
Frame = +1
Query: 38 FITQVISNGDSTTNNYDAKSTATACCNTCLCSVTPELTQC 77
F+ +S STT + D AT C +CLC V L +C
Sbjct: 817 FVCSTVSCTVSTTGHQDTTMDATGPCGSCLCLVALMLLRC 936
>TC207896 GB|BAA13184.1|1498170|D86929 uricase {Glycine max;} , complete
Length = 1348
Score = 32.0 bits (71), Expect = 0.14
Identities = 16/40 (40%), Positives = 20/40 (50%), Gaps = 2/40 (5%)
Frame = +2
Query: 62 CCNTC--LCSVTPELTQCRCADVGKTCHSACKNCECGWSS 99
C + C LC + PE C C GKT + C CGWS+
Sbjct: 311 CYSAC*ALCIILPEGYWCYCEYCGKTMGA----CHCGWST 418
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.131 0.419
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,768,401
Number of Sequences: 63676
Number of extensions: 198668
Number of successful extensions: 2185
Number of sequences better than 10.0: 131
Number of HSP's better than 10.0 without gapping: 2057
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2117
length of query: 164
length of database: 12,639,632
effective HSP length: 90
effective length of query: 74
effective length of database: 6,908,792
effective search space: 511250608
effective search space used: 511250608
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC135311.2


