

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135234.5 - phase: 0 /pseudo
(256 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
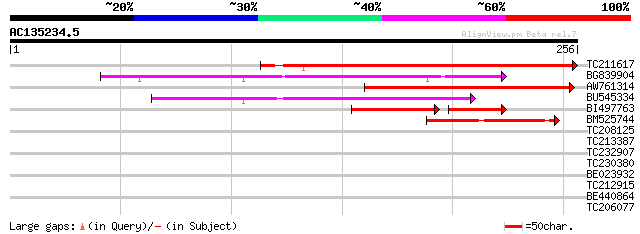
Score E
Sequences producing significant alignments: (bits) Value
TC211617 116 1e-26
BG839904 96 2e-20
AW761314 85 3e-17
BU545334 85 3e-17
BI497763 46 3e-10
BM525744 57 8e-09
TC208125 similar to UP|Q94AF0 (Q94AF0) AT4g29820/F27B13_60 (MRNA... 31 0.48
TC213387 30 1.1
TC232907 30 1.4
TC230380 similar to GB|AAO64866.1|29028974|BT005931 At4g29000 {A... 29 2.4
BE023932 SP|O48922|C98 Cytochrome P450 98A2 (EC 1.14.-.-). [Soyb... 29 2.4
TC212915 similar to UP|Q9SK61 (Q9SK61) Expressed protein, partia... 28 3.1
BE440864 27 7.0
TC206077 similar to UP|O80698 (O80698) F8K4.12, partial (31%) 27 9.1
>TC211617
Length = 534
Score = 116 bits (290), Expect = 1e-26
Identities = 62/144 (43%), Positives = 87/144 (60%), Gaps = 1/144 (0%)
Frame = -1
Query: 114 ERSGIYKPPKTRKKPNLE-GIGSMKCNCPFRLKGFFDKDTNDWWLAILCGMHNHDLEEKL 172
ERSG YK RKK + + KC CPF+++G W + ++CG+HNH+L + L
Sbjct: 528 ERSGKYK---CRKKEFVRRDTCTRKCGCPFKIRGKPMHGGEGWTVKLICGIHNHELAKTL 358
Query: 173 QGHLIAGRLSAEEKKKVIDMTKSLTVPRNILTNLKENNKESVTTIKQVYNVRTRWCKGER 232
H AGRL+ +EK + DMTKS PRNI LKE+N S TTIKQ+YN R+ + R
Sbjct: 357 VEHPYAGRLTDDEKNIIADMTKSNVKPRNIQLTLKEHNTNSCTTIKQIYNARSAYHSSIR 178
Query: 233 GDMTELQFLISKLVEHKYVYLHKM 256
GD TE+Q L+ L +Y++ H++
Sbjct: 177 GDDTEMQHLMRLLKRDQYIHWHRL 106
>BG839904
Length = 712
Score = 95.9 bits (237), Expect = 2e-20
Identities = 65/198 (32%), Positives = 98/198 (48%), Gaps = 15/198 (7%)
Frame = -3
Query: 42 MQARIGKIVEPKRVPI-GVVCEARDTASFFTTNARWREREELLTWVRRQGVKAGFGVCID 100
M+ + + E +PI G ++ D + F T + RE +L W R + GF + I
Sbjct: 590 MEDHVSNMYEVVNMPIDGAANKSDDCSDAFNTTEVFPSREAMLNWARDVAKENGFVLIIL 411
Query: 101 KSVL------------KRPYLTTQSERSGIYKPPKTRKKPNLEGIGSMKCNCPFRLKGFF 148
+S ++ ++ +RSG Y+ P + + + + G+ KC CPF+LKG
Sbjct: 410 RSETSTTCTRSTRCNQRKTFVIMGCDRSGKYRGPY-KNELSRKVSGTRKCECPFKLKGKA 234
Query: 149 DKDTNDWWLAILCGMHNHDLEEKLQGHLIAGRLSAEEKK--KVIDMTKSLTVPRNILTNL 206
K W + ++CG HNHDL E L GH AGRLSAEEK + D R+ + +
Sbjct: 233 LKKAEGWIVKVMCGCHNHDLXETLVGHPYAGRLSAEEKSFGRCFDKEHD-EAXRHSIDH* 57
Query: 207 KENNKESVTTIKQVYNVR 224
K +VTTIKQ+YN R
Sbjct: 56 KIITWVTVTTIKQIYNAR 3
>AW761314
Length = 331
Score = 85.1 bits (209), Expect = 3e-17
Identities = 40/95 (42%), Positives = 60/95 (63%)
Frame = +1
Query: 161 CGMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTKSLTVPRNILTNLKENNKESVTTIKQV 220
CG NH+L + L H A RL+ +E + +MTKS+ P+NIL LKE+N S T IKQ+
Sbjct: 46 CGSDNHELVKSLVRHPYASRLTKDENIIITNMTKSMVKPKNILLTLKEHNVNSYTAIKQI 225
Query: 221 YNVRTRWCKGERGDMTELQFLISKLVEHKYVYLHK 255
YN R+ +C RG T++Q L+ L + +Y++ H+
Sbjct: 226 YNARSAYCSSIRGSDTKMQQLMKLLEQDQYIHWHR 330
>BU545334
Length = 556
Score = 85.1 bits (209), Expect = 3e-17
Identities = 53/152 (34%), Positives = 79/152 (51%), Gaps = 6/152 (3%)
Frame = -1
Query: 65 DTASFFTTNARWREREELLTWVRRQGVKAGFGVCIDKSVL------KRPYLTTQSERSGI 118
D + F T + R+ +L R + GF I +S + ++ ER+G
Sbjct: 454 DCSYAFNTXQVFGTRDYVLQ*XRSVAHENGFAAVIMRSDTHTGSRGRSSFVLIGCERNGQ 275
Query: 119 YKPPKTRKKPNLEGIGSMKCNCPFRLKGFFDKDTNDWWLAILCGMHNHDLEEKLQGHLIA 178
YK K + IGS K CPF+L+G W + ++CG+HNH+L + L H A
Sbjct: 274 YKC--RNKXXVIRAIGSRKYGCPFKLRGKPVIGGQGWMMKLICGIHNHELAKSLVEHPYA 101
Query: 179 GRLSAEEKKKVIDMTKSLTVPRNILTNLKENN 210
GRL+ +EK ++D+TKS+ PRNIL LKE+N
Sbjct: 100 GRLTKDEKTIIVDITKSMVKPRNILLILKEHN 5
>BI497763
Length = 421
Score = 45.8 bits (107), Expect(2) = 3e-10
Identities = 18/40 (45%), Positives = 27/40 (67%)
Frame = +2
Query: 155 WWLAILCGMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTK 194
W + +LCG HNH+L + L H GRL+ +EK +++MTK
Sbjct: 47 WMVKLLCGSHNHELTKSLVEHPYVGRLTKDEKTIIVNMTK 166
Score = 35.8 bits (81), Expect(2) = 3e-10
Identities = 16/26 (61%), Positives = 18/26 (68%)
Frame = +3
Query: 199 PRNILTNLKENNKESVTTIKQVYNVR 224
PRNIL LKE N TTI+Q+YN R
Sbjct: 162 PRNILLMLKEYNANGCTTIRQIYNAR 239
>BM525744
Length = 411
Score = 57.0 bits (136), Expect = 8e-09
Identities = 29/60 (48%), Positives = 41/60 (68%)
Frame = +3
Query: 189 VIDMTKSLTVPRNILTNLKENNKESVTTIKQVYNVRTRWCKGERGDMTELQFLISKLVEH 248
VIDMTK + PRNIL LK++N + TTI+ +YN R + ++G TE+Q L+ KL+EH
Sbjct: 9 VIDMTKKMVEPRNILLTLKDHNND--TTIRHIYNARQAYRSSQKGPRTEMQHLL-KLLEH 179
>TC208125 similar to UP|Q94AF0 (Q94AF0) AT4g29820/F27B13_60 (MRNA cleavage
factor subunit-like protein), partial (86%)
Length = 1141
Score = 31.2 bits (69), Expect = 0.48
Identities = 32/134 (23%), Positives = 54/134 (39%), Gaps = 15/134 (11%)
Frame = +1
Query: 93 AGFGVCIDKSVL----KRPYLTTQSERSGIYKPPKTRKKPNLEGIGSMKCNCPFRLKGFF 148
+G C++ +L K P+L R+ IYK P R +P +K +L
Sbjct: 331 SGIRTCVEAVLLVELFKHPHLLLLQIRNSIYKLPGGRLRPGESDTDGLKRKLARKLSVNE 510
Query: 149 DKDTNDWWLAILCGM-HNHDLEEKLQGHLIAGRLSAEEKKKV----IDMTKSLTVPRNI- 202
D D ++W + GM D E + + +E KV + ++ VP+N+
Sbjct: 511 DGDGSEWEVGECLGMWWRPDFETLMYPFIPPNVKKPKECTKVFLVKLPESRKFIVPKNMR 690
Query: 203 -----LTNLKENNK 211
L + EN+K
Sbjct: 691 VLAVPLCQVHENHK 732
>TC213387
Length = 861
Score = 30.0 bits (66), Expect = 1.1
Identities = 27/108 (25%), Positives = 42/108 (38%), Gaps = 10/108 (9%)
Frame = +2
Query: 144 LKGFFDKDTNDWWLAILCGMHNHDLEEKLQGHLIAGRLSAEEKKKVIDMTKSLTVPRNIL 203
LKG D N W+L + C + D + +Q H + G L+ KK V+ + R ++
Sbjct: 146 LKGSVDHLINLWFLVLQCQLQYKDFQ--VQAHSLMGILA---KKAVLQEDLRMKA*RGLI 310
Query: 204 TNLKENNKESVTTIKQVY----------NVRTRWCKGERGDMTELQFL 241
+ N I Q Y + + W KG + L FL
Sbjct: 311 SMSMRGNLAKKEAILQYYLRLRILTCILSSKV*WLKGVLDHLINLWFL 454
>TC232907
Length = 762
Score = 29.6 bits (65), Expect = 1.4
Identities = 13/30 (43%), Positives = 18/30 (59%)
Frame = +3
Query: 107 PYLTTQSERSGIYKPPKTRKKPNLEGIGSM 136
P TQS++ GI +PP T KK L +G +
Sbjct: 531 PGALTQSQKRGIGRPPGTGKKQQLASLGEL 620
>TC230380 similar to GB|AAO64866.1|29028974|BT005931 At4g29000 {Arabidopsis
thaliana;} , partial (52%)
Length = 1463
Score = 28.9 bits (63), Expect = 2.4
Identities = 11/22 (50%), Positives = 14/22 (63%)
Frame = +3
Query: 69 FFTTNARWREREELLTWVRRQG 90
FFTTN + ++R L W RQG
Sbjct: 534 FFTTNIKEKKRSRTLIWANRQG 599
>BE023932 SP|O48922|C98 Cytochrome P450 98A2 (EC 1.14.-.-). [Soybean]
{Glycine max}, partial (21%)
Length = 417
Score = 28.9 bits (63), Expect = 2.4
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = -1
Query: 23 GSGDVKPTINLKPFKVQIPMQARIGKIVEP 52
G GD++ + PF+ Q+P A +G V P
Sbjct: 309 GKGDLRSLLQSSPFQAQLPCLAPLGPFVAP 220
>TC212915 similar to UP|Q9SK61 (Q9SK61) Expressed protein, partial (8%)
Length = 739
Score = 28.5 bits (62), Expect = 3.1
Identities = 22/76 (28%), Positives = 34/76 (43%), Gaps = 2/76 (2%)
Frame = -3
Query: 177 IAGRLSAEEKKKVIDMTKSLTVPRNILTNLKENNKESVTTIKQVYNVRTRWCKGERGDM- 235
IAG +A + D T N TN+ NN++ T + + YN+R+R E+ ++
Sbjct: 635 IAGAQTAATQFSKKDQNSRSTAWNNAPTNMNANNED--TEVARSYNLRSRGRSTEKDNIS 462
Query: 236 -TELQFLISKLVEHKY 250
TEL L Y
Sbjct: 461 STELNMKFGDLTLPSY 414
>BE440864
Length = 376
Score = 27.3 bits (59), Expect = 7.0
Identities = 12/44 (27%), Positives = 20/44 (45%)
Frame = -1
Query: 42 MQARIGKIVEPKRVPIGVVCEARDTASFFTTNARWREREELLTW 85
M+ +G+ +PK VC R +S T RW ++ + W
Sbjct: 250 MKLLVGQTEKPKGAMRWFVCGQRKRSSMVTEKCRWHDKRKQ*IW 119
>TC206077 similar to UP|O80698 (O80698) F8K4.12, partial (31%)
Length = 780
Score = 26.9 bits (58), Expect = 9.1
Identities = 12/30 (40%), Positives = 15/30 (50%)
Frame = +1
Query: 60 VCEARDTASFFTTNARWREREELLTWVRRQ 89
V R + + F RWR R E TW RR+
Sbjct: 361 VARRRGSCTSFARVKRWRRRRECGTWRRRR 450
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.136 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,608,637
Number of Sequences: 63676
Number of extensions: 157151
Number of successful extensions: 875
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 869
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 872
length of query: 256
length of database: 12,639,632
effective HSP length: 95
effective length of query: 161
effective length of database: 6,590,412
effective search space: 1061056332
effective search space used: 1061056332
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 58 (26.9 bits)
Medicago: description of AC135234.5


