

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135233.1 + phase: 0
(87 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
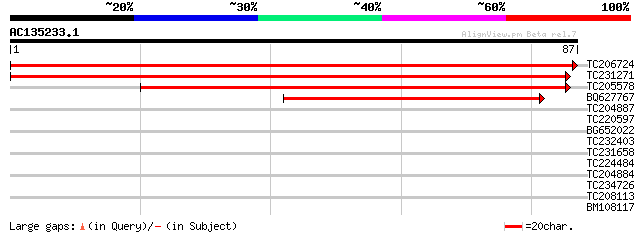
Score E
Sequences producing significant alignments: (bits) Value
TC206724 similar to UP|Q9FHY6 (Q9FHY6) GAMM1 protein-like, parti... 153 1e-38
TC231271 weakly similar to UP|Q94A80 (Q94A80) AT5g41970/MJC20_7,... 128 6e-31
TC205578 similar to UP|Q94A65 (Q94A65) AT4g27700/T29A15_190, par... 98 8e-22
BQ627767 similar to GP|15146292|gb| AT5g41970/MJC20_7 {Arabidops... 71 8e-14
TC204887 similar to UP|Q9ZTX0 (Q9ZTX0) Similarity to SCAMP37, pa... 27 2.3
TC220597 weakly similar to UP|WR32_ARATH (P59583) Probable WRKY ... 27 2.3
BG652022 26 3.0
TC232403 26 4.0
TC231658 26 4.0
TC224484 similar to UP|Q38848 (Q38848) LRP1, partial (20%) 25 6.7
TC204884 similar to UP|Q9ZTX0 (Q9ZTX0) Similarity to SCAMP37, co... 25 6.7
TC234726 weakly similar to GB|AAN31089.1|23506155|AY149935 At5g0... 25 6.7
TC208113 25 8.8
BM108117 homologue to GP|20515818|gb| Subtilisin-like serine pro... 25 8.8
>TC206724 similar to UP|Q9FHY6 (Q9FHY6) GAMM1 protein-like, partial (93%)
Length = 1529
Score = 153 bits (387), Expect = 1e-38
Identities = 72/87 (82%), Positives = 81/87 (92%)
Frame = +3
Query: 1 MKIDPPIKYVLYEDERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGIPGCV 60
+K DP +KYVLY+DERSK WRVQAV+VSPD F+SRKALPSQWRGLRD+ LSK+SGIPGCV
Sbjct: 888 LKNDPAVKYVLYQDERSKHWRVQAVAVSPDSFQSRKALPSQWRGLRDEELSKKSGIPGCV 1067
Query: 61 FVHMSGFIGGNQTFEGALAMAKAALKM 87
FVH+SGFIGGNQ F+GALAMAKAALKM
Sbjct: 1068FVHISGFIGGNQNFDGALAMAKAALKM 1148
>TC231271 weakly similar to UP|Q94A80 (Q94A80) AT5g41970/MJC20_7, partial
(37%)
Length = 583
Score = 128 bits (321), Expect = 6e-31
Identities = 59/86 (68%), Positives = 73/86 (84%)
Frame = +1
Query: 1 MKIDPPIKYVLYEDERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGIPGCV 60
MKI P IKYVLY D+RS+ WR+QAV++SP +FESRK LP WRGL +D LS+ +GIPGC
Sbjct: 160 MKISPSIKYVLYPDDRSENWRLQAVAISPAKFESRKPLPYLWRGLENDNLSEVAGIPGCT 339
Query: 61 FVHMSGFIGGNQTFEGALAMAKAALK 86
FVHMSGFIGGN+++ GALAMA+A+LK
Sbjct: 340 FVHMSGFIGGNRSYGGALAMARASLK 417
>TC205578 similar to UP|Q94A65 (Q94A65) AT4g27700/T29A15_190, partial (78%)
Length = 1454
Score = 97.8 bits (242), Expect = 8e-22
Identities = 45/66 (68%), Positives = 57/66 (86%)
Frame = -2
Query: 21 RVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGIPGCVFVHMSGFIGGNQTFEGALAM 80
R+QAV++SP +FESRK LP WRGL +D LS+ +GIPGC FVHMSGFIGGN+++ GALAM
Sbjct: 1444 RLQAVAISPAKFESRKPLPYLWRGLENDNLSEVAGIPGCTFVHMSGFIGGNRSYGGALAM 1265
Query: 81 AKAALK 86
A+A+LK
Sbjct: 1264 ARASLK 1247
>BQ627767 similar to GP|15146292|gb| AT5g41970/MJC20_7 {Arabidopsis
thaliana}, partial (10%)
Length = 167
Score = 71.2 bits (173), Expect = 8e-14
Identities = 32/40 (80%), Positives = 37/40 (92%)
Frame = +1
Query: 43 RGLRDDILSKESGIPGCVFVHMSGFIGGNQTFEGALAMAK 82
+GLRD+ LSKESGIPGCVFVHMSGFIGGN+ F+GALAM +
Sbjct: 1 QGLRDEELSKESGIPGCVFVHMSGFIGGNKNFDGALAMGE 120
>TC204887 similar to UP|Q9ZTX0 (Q9ZTX0) Similarity to SCAMP37, partial (54%)
Length = 673
Score = 26.6 bits (57), Expect = 2.3
Identities = 10/28 (35%), Positives = 14/28 (49%)
Frame = +2
Query: 38 LPSQWRGLRDDILSKESGIPGCVFVHMS 65
LP QW G D+ ++ GC +H S
Sbjct: 413 LPRQWEGR*DETGGRKGSTEGCNMIHQS 496
>TC220597 weakly similar to UP|WR32_ARATH (P59583) Probable WRKY
transcription factor 32 (WRKY DNA-binding protein 32),
partial (13%)
Length = 894
Score = 26.6 bits (57), Expect = 2.3
Identities = 20/57 (35%), Positives = 28/57 (49%)
Frame = +3
Query: 10 VLYEDERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGIPGCVFVHMSG 66
V+ EDE ++Q RV S+S ESR + PS + L S +P + HM G
Sbjct: 87 VMEEDEETQQ-RVTDSSLST---ESRTSTPSDGAPPNSETLVHRSSLPHDLATHMQG 245
>BG652022
Length = 424
Score = 26.2 bits (56), Expect = 3.0
Identities = 17/46 (36%), Positives = 23/46 (49%)
Frame = -1
Query: 15 ERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGIPGCV 60
E SK ++++ S SP+ GLRD +L E GIPG V
Sbjct: 244 EESKVCKLESSSTSPEDIA*---------GLRDTLLYLEPGIPGLV 134
>TC232403
Length = 1252
Score = 25.8 bits (55), Expect = 4.0
Identities = 16/38 (42%), Positives = 19/38 (49%)
Frame = -1
Query: 31 RFESRKALPSQWRGLRDDILSKESGIPGCVFVHMSGFI 68
+ E RK PSQ R R ESG P C+ + GFI
Sbjct: 571 QLEHRKG-PSQGRNFR------ESGAPSCIEIPSPGFI 479
>TC231658
Length = 582
Score = 25.8 bits (55), Expect = 4.0
Identities = 16/39 (41%), Positives = 20/39 (51%)
Frame = -1
Query: 25 VSVSPDRFESRKALPSQWRGLRDDILSKESGIPGCVFVH 63
+S S +F R A+PS WR SKES CV+ H
Sbjct: 153 ISSSNSQFHERFAIPSGWRR*HLP-ASKESRHA*CVYTH 40
>TC224484 similar to UP|Q38848 (Q38848) LRP1, partial (20%)
Length = 574
Score = 25.0 bits (53), Expect = 6.7
Identities = 11/29 (37%), Positives = 16/29 (54%)
Frame = -3
Query: 45 LRDDILSKESGIPGCVFVHMSGFIGGNQT 73
L D + + G P +H+SG +GGN T
Sbjct: 398 LYDQGVEDKEGYPNLSELHLSGGVGGNVT 312
>TC204884 similar to UP|Q9ZTX0 (Q9ZTX0) Similarity to SCAMP37, complete
Length = 1343
Score = 25.0 bits (53), Expect = 6.7
Identities = 11/28 (39%), Positives = 15/28 (53%)
Frame = +1
Query: 38 LPSQWRGLRDDILSKESGIPGCVFVHMS 65
LP QW G D+ S++ GC +H S
Sbjct: 1000 LPWQW*GC*DETGSRQGSNEGCNMMHQS 1083
>TC234726 weakly similar to GB|AAN31089.1|23506155|AY149935
At5g03760/F17C15_180 {Arabidopsis thaliana;} , partial
(10%)
Length = 694
Score = 25.0 bits (53), Expect = 6.7
Identities = 14/45 (31%), Positives = 21/45 (46%)
Frame = +3
Query: 4 DPPIKYVLYEDERSKQWRVQAVSVSPDRFESRKALPSQWRGLRDD 48
DPP++ L + V + DRF S LP ++G +DD
Sbjct: 420 DPPLELGLASFASHSSLNFKTVPM--DRFSSSTILPEAFQGAKDD 548
>TC208113
Length = 1343
Score = 24.6 bits (52), Expect = 8.8
Identities = 12/41 (29%), Positives = 22/41 (53%)
Frame = +1
Query: 16 RSKQWRVQAVSVSPDRFESRKALPSQWRGLRDDILSKESGI 56
++K+ +QAVS + ++ L + R DIL ++GI
Sbjct: 208 QAKEADIQAVSELKHALDKKETLLMELRSANSDILENKNGI 330
>BM108117 homologue to GP|20515818|gb| Subtilisin-like serine proteases
{Thermoanaerobacter tengcongensis}, partial (2%)
Length = 496
Score = 24.6 bits (52), Expect = 8.8
Identities = 9/18 (50%), Positives = 12/18 (66%)
Frame = -1
Query: 29 PDRFESRKALPSQWRGLR 46
P R+ +R+ LPS WR R
Sbjct: 145 PTRWRARRRLPSSWRQAR 92
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,721,937
Number of Sequences: 63676
Number of extensions: 40105
Number of successful extensions: 170
Number of sequences better than 10.0: 28
Number of HSP's better than 10.0 without gapping: 170
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 170
length of query: 87
length of database: 12,639,632
effective HSP length: 63
effective length of query: 24
effective length of database: 8,628,044
effective search space: 207073056
effective search space used: 207073056
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC135233.1


