

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135231.7 + phase: 0 /pseudo
(943 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
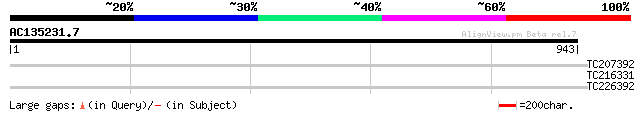
TC207392 similar to UP|BNK_DROME (P40794) Bottleneck protein, pa... 30 5.0
TC216331 similar to UP|O81446 (O81446) Calcineurin B-like protei... 30 6.5
TC226392 29 8.5
>TC207392 similar to UP|BNK_DROME (P40794) Bottleneck protein, partial (7%)
Length = 1544
Score = 30.0 bits (66), Expect = 5.0
Identities = 18/47 (38%), Positives = 24/47 (50%)
Frame = +2
Query: 667 YACFLCLSFFYTGLE*NRSFGVRCNTPFLPLPQLRKLYFICWRLCRL 713
Y LCLS F TGLE R F + +L L L+ L ++ + L L
Sbjct: 962 YFTILCLSSFVTGLEPLRFFYSTSRSTYLSLQVLQTLAYVLYLLLLL 1102
>TC216331 similar to UP|O81446 (O81446) Calcineurin B-like protein 2
(AT5g55990/MDA7_3), partial (35%)
Length = 530
Score = 29.6 bits (65), Expect = 6.5
Identities = 20/83 (24%), Positives = 39/83 (46%), Gaps = 2/83 (2%)
Frame = +2
Query: 765 LYLDRIINQQRPPPLNNIESRGSLPWPEDTNVTSMRPSLLTSTVQVLVFVFAMQKVR--L 822
LYL I+ P +++++S SLP + ++ S + +L+F + +
Sbjct: 20 LYLSSTISLSSPSQIHSLQSSISLPQIPPNSNQTLHYSKFQNFFTLLLFPLLLPDLLPPR 199
Query: 823 FWLRL*HTLVLFRWKWVKL*GCT 845
FW L+ + WKW+K+ C+
Sbjct: 200 FWYL---GLITWSWKWLKVMECS 259
>TC226392
Length = 2066
Score = 29.3 bits (64), Expect = 8.5
Identities = 18/63 (28%), Positives = 29/63 (45%), Gaps = 4/63 (6%)
Frame = +3
Query: 667 YACFLCLSFFYTGLE*----NRSFGVRCNTPFLPLPQLRKLYFICWRLCRLN*NNICHLY 722
+ C L S+F TG+ N + V T PL L L ICW +C++ ++ Y
Sbjct: 1758 FKCLLRESWFTTGIGVHYLNNFDYQVTG*TEVKPLSLLVVLIVICW*VCKIRVTSLVFFY 1937
Query: 723 YGV 725
+ +
Sbjct: 1938 FSI 1946
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.365 0.165 0.642
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 50,960,862
Number of Sequences: 63676
Number of extensions: 862006
Number of successful extensions: 11943
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 4855
Number of HSP's successfully gapped in prelim test: 524
Number of HSP's that attempted gapping in prelim test: 6879
Number of HSP's gapped (non-prelim): 5909
length of query: 943
length of database: 12,639,632
effective HSP length: 106
effective length of query: 837
effective length of database: 5,889,976
effective search space: 4929909912
effective search space used: 4929909912
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135231.7


