

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
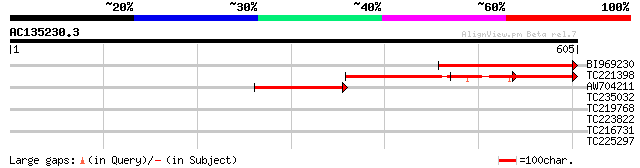
Query= AC135230.3 + phase: 0 /pseudo
(605 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
BI969230 244 6e-65
TC221398 174 8e-44
AW704211 162 4e-40
TC235032 weakly similar to UP|Q7XDX0 (Q7XDX0) Contains similarit... 37 0.019
TC219768 similar to GB|AAD17339.1|4325339|F15P23 F15P23.1 {Arabi... 35 0.096
TC223822 32 0.62
TC216731 homologue to GB|AAL10483.1|15983773|AY056792 AT4g29060/... 29 5.3
TC225297 similar to UP|Q6VEJ5 (Q6VEJ5) Alanine aminotransferase,... 29 6.9
>BI969230
Length = 781
Score = 244 bits (624), Expect = 6e-65
Identities = 119/148 (80%), Positives = 126/148 (84%)
Frame = -1
Query: 458 PHLYFANDDLEYQTKFPSERLGMGAFRIALESIFNRTHHHSLEYTCFGKPHPSVFKNAEI 517
P FANDDLEYQTKF ERLGMGA R+ LESIFNR H SLEYT FGKPHPSVFKNAEI
Sbjct: 781 PXXIFANDDLEYQTKFXXERLGMGAXRMXLESIFNRIHPQSLEYTSFGKPHPSVFKNAEI 602
Query: 518 VLQKHVPRVDEDSYDINHKNAQPFQTLYMIGDNPAVDIRGARQTGHPWFSILTRTGVFKG 577
+L+K V + +D YDINH N F+TLYMIGDNPAVDI+GARQTGHPWFSILTRTGVFKG
Sbjct: 601 LLEKLVASLYDDFYDINHDNTSYFKTLYMIGDNPAVDIKGARQTGHPWFSILTRTGVFKG 422
Query: 578 KENHDKFPADLVVDTVEEAVDYILAKEC 605
KENHDKF ADLVVDTVEEAVDYIL KEC
Sbjct: 421 KENHDKFSADLVVDTVEEAVDYILTKEC 338
>TC221398
Length = 891
Score = 174 bits (442), Expect = 8e-44
Identities = 102/194 (52%), Positives = 118/194 (60%), Gaps = 10/194 (5%)
Frame = +2
Query: 359 GEPALVMSEYGFKNVISIDAYASRFENIDPLAPYKKWTTKLATTQNPKFDESGPQIDVFS 418
GEPA +M+EYGFK +SID YAS FENIDPLAPYKKWTTKLA T N KF+ES P+ DVFS
Sbjct: 2 GEPAAMMTEYGFKYALSIDEYASCFENIDPLAPYKKWTTKLAATLNAKFNESVPRNDVFS 181
Query: 419 ERVQAAFIVSDPVDWSRDVQVLCDILKTGGLPGRNVGTQPHLYFANDDLEYQTKFPSERL 478
+RVQAAF+VSDPVDWSRD+Q C + P L Y T
Sbjct: 182 KRVQAAFVVSDPVDWSRDIQGGCFFSMDKSCRKIQIIWSPFKI-----LYYYTY*QGPTK 346
Query: 479 GMGAFRIAL----------ESIFNRTHHHSLEYTCFGKPHPSVFKNAEIVLQKHVPRVDE 528
G F + + NRTH SL YT FGKPH SVFKNAEIVL+K V + +
Sbjct: 347 GRLNFLLNVWEWELSGLH*NPSSNRTHPCSLRYTYFGKPHSSVFKNAEIVLEKLVASLYD 526
Query: 529 DSYDINHKNAQPFQ 542
D YDINH +A F+
Sbjct: 527 DFYDINHDSASYFK 568
Score = 144 bits (364), Expect = 9e-35
Identities = 89/142 (62%), Positives = 96/142 (66%), Gaps = 7/142 (4%)
Frame = +1
Query: 471 TKFPSERLGMGAFRIALESIFNRTHHHSLEYTCFGKPHPSVFKNAEIVLQKHVPRVDEDS 530
TKFPSERLGMGAFRIALESIF LE F ++ I +Q+ E S
Sbjct: 352 TKFPSERLGMGAFRIALESIFK*DSPLFLEVYLF-------WQTTFICIQECRNCARETS 510
Query: 531 YD-----INHKN--AQPFQTLYMIGDNPAVDIRGARQTGHPWFSILTRTGVFKGKENHDK 583
+ +K+ F LYMIGDNPAVDIRGARQTGHPWFSILTRT VFKGKENHDK
Sbjct: 511 GVTL**FL*YKS**RFIF*KLYMIGDNPAVDIRGARQTGHPWFSILTRTVVFKGKENHDK 690
Query: 584 FPADLVVDTVEEAVDYILAKEC 605
F ADLVVDTVEEAVDYILA EC
Sbjct: 691 FSADLVVDTVEEAVDYILANEC 756
>AW704211
Length = 401
Score = 162 bits (410), Expect = 4e-40
Identities = 78/99 (78%), Positives = 89/99 (89%)
Frame = +2
Query: 262 RPSFGIAFDIDGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGGIPEAKRASEL 321
RP+FGIAFDIDGV+LLGN+PVGGSP AL++LY+ DG LK PYVFLTNGGG PEAKRA EL
Sbjct: 104 RPTFGIAFDIDGVLLLGNSPVGGSPGALKRLYDADGKLKIPYVFLTNGGGYPEAKRAFEL 283
Query: 322 SELLGLNVSASQVLQGHSPFRQLVNRFEDKLIVAAGKGE 360
S+LLG+NV+ SQVLQGHSPF+QLV RFE+ LIVA GKGE
Sbjct: 284 SKLLGINVTPSQVLQGHSPFKQLVKRFENDLIVAVGKGE 400
>TC235032 weakly similar to UP|Q7XDX0 (Q7XDX0) Contains similarity to
ribosomal protein, partial (24%)
Length = 689
Score = 37.4 bits (85), Expect = 0.019
Identities = 15/34 (44%), Positives = 22/34 (64%)
Frame = +3
Query: 171 IQHVSETRGFPWMIGSIDCMHWEWKNYPKAWEGK 204
+ + E RGF M+GSID +HW+ K AW+G+
Sbjct: 21 LMQICEIRGFQSMLGSIDYLHWK*KTCLVAWKGQ 122
>TC219768 similar to GB|AAD17339.1|4325339|F15P23 F15P23.1 {Arabidopsis
thaliana;} , partial (26%)
Length = 982
Score = 35.0 bits (79), Expect = 0.096
Identities = 23/61 (37%), Positives = 35/61 (56%), Gaps = 2/61 (3%)
Frame = -3
Query: 209 NHVRARSRVASRITSRSSEA--HVDKV*NTETNLISLIISLLLLCYPILINDLLIRPSFG 266
NH+ +R + +S+S++ HV KV + LIISLL++CYP L ++ SFG
Sbjct: 356 NHIHHPNR*IYKCSSKSTQVSIHV*KV-AVSPSFS*LIISLLIICYPFLPQFIIF*ESFG 180
Query: 267 I 267
I
Sbjct: 179 I 177
>TC223822
Length = 1054
Score = 32.3 bits (72), Expect = 0.62
Identities = 29/112 (25%), Positives = 43/112 (37%), Gaps = 13/112 (11%)
Frame = +3
Query: 485 IALESIFNRTHHHSLEYTCFGKPHPSVFKNAEIVLQKHVPR--------VDEDSYDINHK 536
+A F HH Y +P P+ + + H+P+ E I
Sbjct: 144 LATNPAFRVIHHRIKHYYQEPRPVPTHPNSPSLTPSTHLPQNMMKAKNIKGEGEIPIKFD 323
Query: 537 NAQPFQTLYMIGD-----NPAVDIRGARQTGHPWFSILTRTGVFKGKENHDK 583
N+ P T + G +P V I A+ PW +R GVFKG+ N+ K
Sbjct: 324 NSIPTSTPMVHGQVSKVASPQVGI-SAKVEPEPWHVQGSRVGVFKGEHNNGK 476
>TC216731 homologue to GB|AAL10483.1|15983773|AY056792 AT4g29060/F19B15_90
{Arabidopsis thaliana;} , partial (13%)
Length = 723
Score = 29.3 bits (64), Expect = 5.3
Identities = 14/35 (40%), Positives = 18/35 (51%)
Frame = -1
Query: 488 ESIFNRTHHHSLEYTCFGKPHPSVFKNAEIVLQKH 522
ESI N T + L TC K HP +F + + V H
Sbjct: 705 ESIINETSNSKLNKTCNSKMHPLIFFSIQTVEN*H 601
>TC225297 similar to UP|Q6VEJ5 (Q6VEJ5) Alanine aminotransferase, partial
(98%)
Length = 1833
Score = 28.9 bits (63), Expect = 6.9
Identities = 25/98 (25%), Positives = 45/98 (45%), Gaps = 2/98 (2%)
Frame = +1
Query: 272 DGVILLGNTPVGGSPAALRKLYNYDGTLKFPYVFLTNGGG--IPEAKRASELSELLGLNV 329
DG++ P+ P + + G L Y+ G G IPE K+ E ++ G+NV
Sbjct: 640 DGILC----PIPQYPLYSASIDLHGGCLVPYYLDEATGWGLEIPELKKQLEAAKSKGINV 807
Query: 330 SASQVLQGHSPFRQLVNRFEDKLIVAAGKGEPALVMSE 367
A V+ +P Q++ + IV K E +++++
Sbjct: 808 RALVVINPGNPTGQVLGEANQRDIVEFCKQEGLVLLAD 921
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.139 0.418
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,674,156
Number of Sequences: 63676
Number of extensions: 350298
Number of successful extensions: 1650
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 1641
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1650
length of query: 605
length of database: 12,639,632
effective HSP length: 103
effective length of query: 502
effective length of database: 6,081,004
effective search space: 3052664008
effective search space used: 3052664008
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 62 (28.5 bits)
Medicago: description of AC135230.3


