

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135230.11 + phase: 0 /pseudo
(954 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
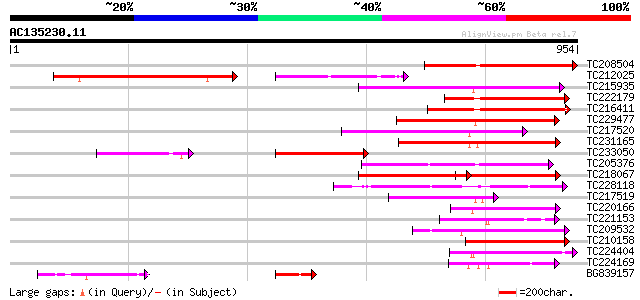
Score E
Sequences producing significant alignments: (bits) Value
TC208504 similar to UP|Q8H934 (Q8H934) Phototropin, partial (26%) 452 e-127
TC212025 similar to UP|Q8H934 (Q8H934) Phototropin, partial (27%) 445 e-125
TC215935 homologue to UP|Q39183 (Q39183) Serine/threonine protei... 341 1e-93
TC222179 similar to UP|Q7DMT0 (Q7DMT0) Protein kinase , partial ... 333 2e-91
TC216411 similar to UP|Q41384 (Q41384) Protein kinase (Fragment)... 315 8e-86
TC229477 similar to UP|Q9M1P3 (Q9M1P3) Protein kinase-like prote... 293 3e-79
TC217520 homologue to PIR|T00410|T00410 protein kinase homolog T... 275 9e-74
TC231165 similar to UP|Q6RC06 (Q6RC06) Serine/threonine protein ... 246 3e-65
TC233050 homologue to UP|Q41384 (Q41384) Protein kinase (Fragmen... 244 1e-64
TC205376 similar to UP|Q9MB45 (Q9MB45) IRE homolog 1 (Fragment),... 214 1e-55
TC218067 similar to UP|KPK1_PHAVU (P15792) Protein kinase PVPK-1... 202 6e-52
TC228118 similar to UP|KP19_ARATH (Q39030) Serine/threonine-prot... 196 3e-50
TC217519 homologue to PIR|T00410|T00410 protein kinase homolog T... 184 2e-46
TC220166 similar to UP|Q9LZS4 (Q9LZS4) Protein kinase-like prote... 182 8e-46
TC221153 similar to UP|Q9LSF1 (Q9LSF1) Protein kinases-like prot... 167 2e-41
TC209532 similar to UP|O23314 (O23314) Protein kinase, partial (... 166 3e-41
TC210158 weakly similar to UP|YPK2_YEAST (P18961) Serine/threoni... 165 8e-41
TC224404 homologue to UP|Q9M728 (Q9M728) Viroid symptom modulati... 163 3e-40
TC224169 similar to UP|Q9LT38 (Q9LT38) Protein kinase-like prote... 159 6e-39
BG839157 146 5e-35
>TC208504 similar to UP|Q8H934 (Q8H934) Phototropin, partial (26%)
Length = 1093
Score = 452 bits (1162), Expect = e-127
Identities = 220/257 (85%), Positives = 235/257 (90%)
Frame = +1
Query: 698 GELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLT 757
GELFLLLD+QPTKVLKEDAVRFYAAEV+I LEYLHCQGIIYRDLKPENVL+Q NGHVSLT
Sbjct: 1 GELFLLLDRQPTKVLKEDAVRFYAAEVVIVLEYLHCQGIIYRDLKPENVLLQSNGHVSLT 180
Query: 758 DFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYI 817
DFDLSCLTS KPQLIIPA KKK+KKK QK+Q++P FMAEPMRASNSFVGTEEYI
Sbjct: 181 DFDLSCLTSSKPQLIIPATNSKKKKKKK-----QKSQEVPMFMAEPMRASNSFVGTEEYI 345
Query: 818 APEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSP 877
APEIITGSGHTSAVDWWALGIL+YEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVS
Sbjct: 346 APEIITGSGHTSAVDWWALGILIYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSL 525
Query: 878 QAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPELDAPILLENDEK 937
Q KQLIYWLL RDPK+RLGS EGANEIK HPFF+ VNWAL+RCMKPPELDAP+L E +E+
Sbjct: 526 QGKQLIYWLLQRDPKDRLGSREGANEIKRHPFFRGVNWALVRCMKPPELDAPLLPETEEE 705
Query: 938 KEAKDIDPGLDDLQKNI 954
KEAKDI PGL+DLQ NI
Sbjct: 706 KEAKDIHPGLEDLQTNI 756
>TC212025 similar to UP|Q8H934 (Q8H934) Phototropin, partial (27%)
Length = 978
Score = 445 bits (1144), Expect = e-125
Identities = 236/325 (72%), Positives = 261/325 (79%), Gaps = 16/325 (4%)
Frame = +1
Query: 75 ETGAAAQRAAEWGLVLRTDAETGKPQGVGVRNSGDDEQNGK------FSGKRNSNNSGRV 128
E+G AAQRAAEWGLVLRTD ETGKPQGV VRNSG +E N S ++NS NS R
Sbjct: 1 ESGEAAQRAAEWGLVLRTDTETGKPQGVAVRNSGGEEPNAAKLAAAASSSRKNSQNSART 180
Query: 129 SGDSSDGGDPRG-FPRVSEDLKDALSAFQQTFVVSDATKPDYPIMYASAGFFNMTGYTSK 187
SGDSSDGG G PR+SED+ ALSAFQQTFVVSDATK DYPI+YASAGFF MTGYTSK
Sbjct: 181 SGDSSDGGGGGGGIPRISEDVMGALSAFQQTFVVSDATKADYPILYASAGFFKMTGYTSK 360
Query: 188 EVIGRNCRFLQGADTDPQDVAKIREALEGGKSYCGRLLNYKKDGTPFWNLLTISPIKDDD 247
EVIGRNCRFLQGADTDP+DVAKIREAL+ GK YCGRLLNYKKDGTPFWNLLTISPIKD+D
Sbjct: 361 EVIGRNCRFLQGADTDPEDVAKIREALQAGKIYCGRLLNYKKDGTPFWNLLTISPIKDED 540
Query: 248 GNVLKLIGMLVEVNKHTEGSKEKNLRPNGLPESLIRYDARQKEKASSSVSELLQAMKRPR 307
G VLK IGM VEV+KHTEGSKEK LRPNGLPESLIRYDARQKEKA+SSV+ELLQAMKRPR
Sbjct: 541 GKVLKFIGMQVEVSKHTEGSKEKTLRPNGLPESLIRYDARQKEKATSSVTELLQAMKRPR 720
Query: 308 ALSESGQRPFIIKSGGCSEEDQEI--------EKVEHKSRRKSDSVASFRPKSQRK-SRS 358
ALSES RP I KSG S +++++ EK + RR S+S ASF KS+ +R
Sbjct: 721 ALSESASRPSIRKSGSRSSDEEKLEQEQEDDKEKAQKTLRRISESGASFGRKSEGSGNRI 900
Query: 359 SMERISELPENANKNSHRHSFMGLM 383
SMERISELPEN ++NS R SFMG +
Sbjct: 901 SMERISELPENKHRNSQRRSFMGYL 975
Score = 99.4 bits (246), Expect = 7e-21
Identities = 73/229 (31%), Positives = 119/229 (51%), Gaps = 6/229 (2%)
Frame = +1
Query: 448 FLELTEYSREEILGRNCRFLQGPETDPATVRKIREAIDNQTEVTVQLINYTRTGKKFWNL 507
F ++T Y+ +E++GRNCRFLQG +TDP V KIREA+ +L+NY + G FWNL
Sbjct: 331 FFKMTGYTSKEVIGRNCRFLQGADTDPEDVAKIREALQAGKIYCGRLLNYKKDGTPFWNL 510
Query: 508 FHLQPMRDHKGEVQYFIGVQLDGSQHVEPLHNCIKEDTAKEG---EQLVKQTA---ENVG 561
+ P++D G+V FIG+Q++ S+H E KE T + E L++ A E
Sbjct: 511 LTISPIKDEDGKVLKFIGMQVEVSKHTEG----SKEKTLRPNGLPESLIRYDARQKEKAT 678
Query: 562 EAVRELPDANQKPDDLWLNHSKVVHPKPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPL 621
+V EL A ++P L + S+ K + +D +++ E+ ++ + K R I
Sbjct: 679 SSVTELLQAMKRPRALSESASRPSIRKSGSRSSDE-EKLEQEQEDDKEKAQKTLRRI--- 846
Query: 622 GSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDML 670
++G+ + EG+G +M+ + + L NK HR R + L
Sbjct: 847 --SESGASFGRKSEGSGNRISMERISE---LPENK-HRNSQRRSFMGYL 975
>TC215935 homologue to UP|Q39183 (Q39183) Serine/threonine protein kinase
(Protein kinase 5) (AT5g47750/MCA23_7) , partial (82%)
Length = 2655
Score = 341 bits (874), Expect = 1e-93
Identities = 183/390 (46%), Positives = 234/390 (59%), Gaps = 44/390 (11%)
Frame = +3
Query: 588 KPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMD 647
KPH+ ++ W AIQ + + L HFR +K LG GD GSV+L EL GT YFAMK MD
Sbjct: 738 KPHKANDLRWEAIQAVRSRDGVLGLGHFRLLKRLGCGDIGSVYLSELSGTKCYFAMKVMD 917
Query: 648 KGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQ 707
KG + +R K+ RA TEREIL LDHPFLP LY F+T+ CL+ ++ PGG+L L +Q
Sbjct: 918 KGSLASRKKLLRAQTEREILQSLDHPFLPTLYTHFETEKFSCLVMEFCPGGDLHTLRQKQ 1097
Query: 708 PTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSC 767
P K E AV+FY AEVL+ALEYLH GI+YRDLKPENVL++ +GH+ L+DFDLS +
Sbjct: 1098 PGKHFPEQAVKFYVAEVLLALEYLHMLGIVYRDLKPENVLVRDDGHIMLSDFDLSLRCAV 1277
Query: 768 KPQLIIPANED-------------------------------------------KKKRKK 784
P L+ ++ D KK RK
Sbjct: 1278 SPTLVKTSSTDSEPLRKNAVYCVQPACIEPPSCIQPSCVAPTTCFSPRLFSSKSKKDRKP 1457
Query: 785 KKKKGQQKTQQIPTFMAEPMRA-SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEM 843
K + G Q +P +AEP A S SFVGT EY+APEII G GH SAVDWW GI LYE+
Sbjct: 1458 KNEIGNQ-VSPLPELIAEPTDARSMSFVGTHEYLAPEIIKGEGHGSAVDWWTFGIFLYEL 1634
Query: 844 LYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANE 903
L+G TPF+G + T N++ + L+FP++ VS A+ LI LL ++P++RL GA E
Sbjct: 1635 LFGKTPFKGSGNRATLFNVVGQPLRFPEAPVVSFAARDLIRGLLVKEPQHRLAYKRGATE 1814
Query: 904 IKSHPFFKNVNWALIRCMKPPELDAPILLE 933
IK HPFF+ VNWALIRC PPE+ + E
Sbjct: 1815 IKQHPFFEGVNWALIRCATPPEIPKAVEFE 1904
>TC222179 similar to UP|Q7DMT0 (Q7DMT0) Protein kinase , partial (44%)
Length = 882
Score = 333 bits (855), Expect = 2e-91
Identities = 167/212 (78%), Positives = 180/212 (84%), Gaps = 2/212 (0%)
Frame = +2
Query: 732 HCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQ 791
HCQGIIYRDLKPENVL+Q +GHVSLTDFDLSCLTSCKPQL++P +KKK Q
Sbjct: 2 HCQGIIYRDLKPENVLLQSSGHVSLTDFDLSCLTSCKPQLLVPVINEKKKA--------Q 157
Query: 792 KTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFR 851
K P FMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEM YGYTPFR
Sbjct: 158 KGPHAPIFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMFYGYTPFR 337
Query: 852 GKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFK 911
GKTRQ+TF NILHKDLKFPKSK VS AKQL+Y LL+RDPK+RLGS EGANEIK+HPFF+
Sbjct: 338 GKTRQRTFTNILHKDLKFPKSKQVSFSAKQLMYRLLNRDPKSRLGSREGANEIKNHPFFR 517
Query: 912 NVNWALIRCMKPPELDAPILLENDE--KKEAK 941
VNWAL+RC KPPELDAP LLE E +KEAK
Sbjct: 518 GVNWALVRCTKPPELDAP-LLETTEGGEKEAK 610
>TC216411 similar to UP|Q41384 (Q41384) Protein kinase (Fragment) , partial
(32%)
Length = 1044
Score = 315 bits (806), Expect = 8e-86
Identities = 157/242 (64%), Positives = 186/242 (75%), Gaps = 1/242 (0%)
Frame = +1
Query: 703 LLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLS 762
LLD+QP K+ KE++ RFYAAEV+I LEYLHC GIIYRDLKPEN+L+Q++GHV L DFDLS
Sbjct: 7 LLDKQPMKIFKEESARFYAAEVVIGLEYLHCLGIIYRDLKPENILLQKDGHVVLADFDLS 186
Query: 763 CLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEII 822
+TSCKPQ++ A K++ +++ PTF+AEP+ SNSFVGTEEYIAPEII
Sbjct: 187 FMTSCKPQVVKQAIPGKRR---------SRSEPPPTFVAEPVTQSNSFVGTEEYIAPEII 339
Query: 823 TGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQL 882
TG+GHTS +DWW LGILLYEMLYG TPFRGK RQKTF+NILHKDL FP S P S A+QL
Sbjct: 340 TGAGHTSGIDWWTLGILLYEMLYGRTPFRGKNRQKTFSNILHKDLTFPSSIPASLAARQL 519
Query: 883 IYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPELDAPI-LLENDEKKEAK 941
I LL RDP +R+GS GANEIK HPFF+ +NW LIR M PP LD P+ L+ ND AK
Sbjct: 520 INALLQRDPTSRIGSTTGANEIKQHPFFRGINWPLIRNMTPPPLDVPLKLIGND--PVAK 693
Query: 942 DI 943
DI
Sbjct: 694 DI 699
>TC229477 similar to UP|Q9M1P3 (Q9M1P3) Protein kinase-like protein, partial
(64%)
Length = 1182
Score = 293 bits (749), Expect = 3e-79
Identities = 152/307 (49%), Positives = 194/307 (62%), Gaps = 33/307 (10%)
Frame = +2
Query: 651 MLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTK 710
+ +R+K RA TEREIL+ LDHPFLP LYA+ +CL+T++ PGG+L +L +QP K
Sbjct: 8 LASRSKEGRAKTEREILESLDHPFLPTLYATIDAAKWLCLLTEFCPGGDLHILRQRQPHK 187
Query: 711 VLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCL---TSC 767
E AVRFYA+EVL+ALEYLH GI+YRDLKPENVL++ +GH+ LTDFDLS ++
Sbjct: 188 RFPEPAVRFYASEVLVALEYLHMMGIVYRDLKPENVLVRSDGHIMLTDFDLSLKCDDSTS 367
Query: 768 KPQLIIPANEDKKK-----------------------------RKKKKKKGQQKTQQIPT 798
PQ+I+ K K+K+K +Q P
Sbjct: 368 TPQIILDQKNTPHKDPRVEPSQTQFTSSSCILPNCIVPAVSCFHPKRKRKKKQTQHNGPE 547
Query: 799 FMAEPMRA-SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQK 857
F+AEP+ S SFVGT EY+APEI++G GH SAVDWW LGI ++E+ YG TPFRG +
Sbjct: 548 FVAEPIDVRSMSFVGTHEYLAPEIVSGEGHGSAVDWWTLGIFIFELFYGVTPFRGMDNEL 727
Query: 858 TFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWAL 917
T ANI+ + L+FPK V P AK LI LL +DP RLGS GA+ IK HPFF+ VNWAL
Sbjct: 728 TLANIVARALEFPKEPSVPPTAKDLISQLLVKDPSRRLGSTMGASSIKHHPFFQGVNWAL 907
Query: 918 IRCMKPP 924
+RC PP
Sbjct: 908 LRCTPPP 928
>TC217520 homologue to PIR|T00410|T00410 protein kinase homolog T13E15.16 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(41%)
Length = 1066
Score = 275 bits (702), Expect = 9e-74
Identities = 154/352 (43%), Positives = 203/352 (56%), Gaps = 39/352 (11%)
Frame = +1
Query: 558 ENVGEAVRELPDANQKPDDL-WLNHSKVVHPKPHRKDNDAWRAIQKIIENGEQISLKHFR 616
E+ +V D++ DD W N + + KPH+ ++ W+AI I + + H R
Sbjct: 13 ESAKTSVSRPSDSSGLSDDSNWSNITGSAN-KPHKGNDPRWKAILAIRLRDGILGMSHCR 189
Query: 617 PIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPFLP 676
+K LG GD GSV+L EL GT YFAMK MDK + +R K+ R TEREIL +LDHPFLP
Sbjct: 190 LLKRLGCGDIGSVYLSELSGTRCYFAMKVMDKASLASRKKLTRVQTEREILQLLDHPFLP 369
Query: 677 ALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGI 736
LY F+T CL+ +Y PGG+L L +QP K E A RFYAAEVL+ALEYLH G+
Sbjct: 370 TLYTHFETDRFSCLVMEYCPGGDLHTLRQRQPGKHFSEYAARFYAAEVLLALEYLHMLGV 549
Query: 737 IYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLI------------------------ 772
+YRDLKPENVL++ +GH+ L+DFDLS + P LI
Sbjct: 550 VYRDLKPENVLVRDDGHIMLSDFDLSLRCAVSPTLIRTSYDGDPSKRAGGAFCVQPACIE 729
Query: 773 -----------IPANEDKKKRKKKKKKGQQ--KTQQIPTFMAEPMRA-SNSFVGTEEYIA 818
IP +K +K +K + + +P +AEP +A S SFVGT EY+A
Sbjct: 730 PSSMCIQPACFIPRLFPQKNKKSRKPRADPGLPSSTLPELVAEPTQARSMSFVGTHEYLA 909
Query: 819 PEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFP 870
PEII G GH SAVDWW GI L+E+LYG TPF+G + T N++ + L+FP
Sbjct: 910 PEIIKGEGHGSAVDWWTFGIFLHELLYGKTPFKGSGNRATLFNVVGQQLRFP 1065
>TC231165 similar to UP|Q6RC06 (Q6RC06) Serine/threonine protein kinase,
partial (65%)
Length = 1129
Score = 246 bits (629), Expect = 3e-65
Identities = 131/296 (44%), Positives = 179/296 (60%), Gaps = 23/296 (7%)
Frame = +2
Query: 654 RNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLK 713
+ K RA ER+IL M+DHPFLP LYA F+ C++ +Y GG+L L P
Sbjct: 14 KKKAQRAEMERKILKMVDHPFLPTLYAEFKASNFSCIVMEYCSGGDLHSLQHNHPNNRFS 193
Query: 714 EDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQL-- 771
+ +FYAAEVL+ALEYLH GIIYRDLKPENVL++ +GH+ L+DFDLS + P +
Sbjct: 194 LSSTKFYAAEVLVALEYLHMLGIIYRDLKPENVLVRSDGHIMLSDFDLSLCSDAIPAVES 373
Query: 772 ----IIPANEDKKKRKKK-------------KKKGQQKTQQIPTFMAEPMRA-SNSFVGT 813
+ PA + ++ + + Q Q F+AEP+ A S SFVGT
Sbjct: 374 PDCSLDPAFAPALRYTRQYSTPFSCLSNRVFRSRKVQTLQPNRLFVAEPVGARSCSFVGT 553
Query: 814 EEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSK 873
EY++PE+ +G+ H +AVDWW+ GI +YEM+YG TPF G + + T +I+ K L FP S
Sbjct: 554 HEYVSPEVASGNSHGNAVDWWSFGIFIYEMVYGRTPFAGSSNEATLRSIIKKPLAFPTST 733
Query: 874 PVSP---QAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPEL 926
P S A+ LI LL++DP RLGS G+ ++K HPFF +N ALIR + PPE+
Sbjct: 734 PSSTLEMHARDLISGLLNKDPNRRLGSKRGSADVKKHPFFAGLNLALIRTVTPPEV 901
>TC233050 homologue to UP|Q41384 (Q41384) Protein kinase (Fragment) , partial
(27%)
Length = 928
Score = 244 bits (623), Expect = 1e-64
Identities = 115/156 (73%), Positives = 133/156 (84%)
Frame = +1
Query: 448 FLELTEYSREEILGRNCRFLQGPETDPATVRKIREAIDNQTEVTVQLINYTRTGKKFWNL 507
FLELTEY+REEILGRNCRFLQGPETD ATV +IR+AI Q E+TVQLINYT++GKKFWNL
Sbjct: 337 FLELTEYTREEILGRNCRFLQGPETDQATVSRIRDAIREQREITVQLINYTKSGKKFWNL 516
Query: 508 FHLQPMRDHKGEVQYFIGVQLDGSQHVEPLHNCIKEDTAKEGEQLVKQTAENVGEAVREL 567
FHLQPMRD KGE+QYFIGVQLDGS HVEPL N + E T ++ +LVK TAENV EAVREL
Sbjct: 517 FHLQPMRDQKGELQYFIGVQLDGSDHVEPLKNRLSETTEQQSAKLVKATAENVDEAVREL 696
Query: 568 PDANQKPDDLWLNHSKVVHPKPHRKDNDAWRAIQKI 603
PDAN +P+DLW HS+ V P+PH+KDN +W AIQK+
Sbjct: 697 PDANLRPEDLWAIHSQPVFPRPHKKDNPSWIAIQKV 804
Score = 113 bits (282), Expect = 5e-25
Identities = 62/167 (37%), Positives = 91/167 (54%), Gaps = 4/167 (2%)
Frame = +1
Query: 147 DLKDALSAFQQTFVVSDATKPDYPIMYASAGFFNMTGYTSKEVIGRNCRFLQGADTDPQD 206
DL L ++ FV+SD PD PI++AS F +T YT +E++GRNCRFLQG +TD
Sbjct: 244 DLATTLERIEKNFVISDPRLPDNPIIFASDSFLELTEYTREEILGRNCRFLQGPETDQAT 423
Query: 207 VAKIREALEGGKSYCGRLLNYKKDGTPFWNLLTISPIKDDDGNVLKLIGMLVEVNKHTEG 266
V++IR+A+ + +L+NY K G FWNL + P++D G + IG+ ++ + H E
Sbjct: 424 VSRIRDAIREQREITVQLINYTKSGKKFWNLFHLQPMRDQKGELQYFIGVQLDGSDHVEP 603
Query: 267 SKEKNLRPNGLPESLIRYDAR----QKEKASSSVSELLQAMKRPRAL 309
K N L E+ + A+ E +V EL A RP L
Sbjct: 604 LK------NRLSETTEQQSAKLVKATAENVDEAVRELPDANLRPEDL 726
>TC205376 similar to UP|Q9MB45 (Q9MB45) IRE homolog 1 (Fragment), partial
(45%)
Length = 2377
Score = 214 bits (545), Expect = 1e-55
Identities = 123/328 (37%), Positives = 189/328 (57%), Gaps = 5/328 (1%)
Frame = +2
Query: 593 DNDAWRAIQK--IIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGV 650
++D R+++ I + ++ S+ F IKP+ G G V L + TG FA+K + K
Sbjct: 215 EDDVVRSLRTSPIHSSRDRTSIDDFEIIKPISRGAFGRVFLAKKRTTGDLFAIKVLKKAD 394
Query: 651 MLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTK 710
M+ +N V ER+IL + +PF+ + SF + ++ L+ +Y GG+L+ LL +
Sbjct: 395 MIRKNAVESILAERDILITVRNPFVVRFFYSFTCRENLYLVMEYLNGGDLYSLL--RNLG 568
Query: 711 VLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSC--LTSCK 768
L E+ R Y AEV++ALEYLH +++RDLKP+N+LI +GH+ LTDF LS L +
Sbjct: 569 CLDEEVARVYIAEVVLALEYLHSLRVVHRDLKPDNLLIAHDGHIKLTDFGLSKVGLINST 748
Query: 769 PQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHT 828
L PA + + +T + R S VGT +Y+APEI+ G+GH
Sbjct: 749 DDLSGPAVNGTSLLE------EDETDVFTSADQRERREKRSAVGTPDYLAPEILLGTGHG 910
Query: 829 SAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPK-SKPVSPQAKQLIYWLL 887
DWW++G++L+E+L G PF + Q F NIL++ + +P + +SPQA+ LI LL
Sbjct: 911 FTADWWSVGVILFELLVGIPPFNAEHPQTIFDNILNRKIPWPAVPEEMSPQAQDLIDRLL 1090
Query: 888 HRDPKNRLGSLEGANEIKSHPFFKNVNW 915
DP RLGS +GA+E+K H FFK++NW
Sbjct: 1091TEDPNQRLGS-KGASEVKQHVFFKDINW 1171
>TC218067 similar to UP|KPK1_PHAVU (P15792) Protein kinase PVPK-1 , partial
(77%)
Length = 1540
Score = 202 bits (514), Expect = 6e-52
Identities = 99/189 (52%), Positives = 129/189 (67%)
Frame = +3
Query: 588 KPHRKDNDAWRAIQKIIENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMD 647
KPH+ ++ W AIQ I + ++HFR +K LG GD GSV+L EL GT FAMK M+
Sbjct: 261 KPHKANDIRWEAIQAIRVRDGVLEMRHFRLLKKLGCGDIGSVYLAELSGTRTCFAMKVMN 440
Query: 648 KGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQ 707
K + +R K+ R+ TEREIL LDHPFLP LY F+T+T CL+ ++ PGG+L L +Q
Sbjct: 441 KTELASRKKLVRSQTEREILQSLDHPFLPTLYTHFETETFSCLVMEFCPGGDLHALRQRQ 620
Query: 708 PTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSC 767
P K E A RFY AEVL+ALEYLH G+IYRDLKPENVL++ +GH+ L+DFDLS +
Sbjct: 621 PGKYFSEIAARFYVAEVLLALEYLHMLGVIYRDLKPENVLVREDGHIMLSDFDLSLRCAV 800
Query: 768 KPQLIIPAN 776
P L+ +N
Sbjct: 801 SPTLVKSSN 827
Score = 175 bits (443), Expect = 1e-43
Identities = 92/177 (51%), Positives = 110/177 (61%), Gaps = 1/177 (0%)
Frame = +1
Query: 751 NGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKTQQIPTFMAEPMRA-SNS 809
N HVSL F PAN KKK+ K K Q + +P +AEP A S S
Sbjct: 913 NLHVSLRAFS-------------PANPRKKKKSKPKNDVQNQVTPLPELIAEPTNARSMS 1053
Query: 810 FVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKF 869
FVGT EY+APEII G GH SAVDWW GI LYE+L+G TPF+G + T N++ + LKF
Sbjct: 1054 FVGTHEYLAPEIIKGEGHGSAVDWWTFGIFLYELLFGRTPFKGSANRATLFNVVGQPLKF 1233
Query: 870 PKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPEL 926
P+S VS A+ LI LL ++P+NRL GA EIK HPFF NVNWALIRC PPE+
Sbjct: 1234 PESPTVSFAARDLIRGLLVKEPQNRLAYRRGATEIKQHPFFHNVNWALIRCANPPEV 1404
>TC228118 similar to UP|KP19_ARATH (Q39030) Serine/threonine-protein kinase
AtPK19 (Ribosomal-protein S6 kinase homolog) , partial
(65%)
Length = 1305
Score = 196 bits (499), Expect = 3e-50
Identities = 133/400 (33%), Positives = 199/400 (49%), Gaps = 6/400 (1%)
Frame = +1
Query: 545 TAKEGEQLVKQTAENVGEAVRELPDANQKPDDLWLNHSKVVHPKPHRKDNDAWRAIQKII 604
T E E ++ GEA++++ +++ + L KD D + KI
Sbjct: 16 TIHETEDSLELVEHVNGEAIKDIKESSFVEESL--------------KDEDG--NLMKI- 144
Query: 605 ENGEQISLKHFRPIKPLGSGDTGSVHLVELEGTGQYFAMKAMDKGVMLNRNKVHRACTER 664
++S+ F +K +G G V+ V + T + +AMK M K ++ +N ER
Sbjct: 145 ---HRVSIDDFEILKVVGQGAFAKVYQVRKKDTSEIYAMKVMRKDKIMEKNHAEYMKAER 315
Query: 665 EILDMLDHPFLPALYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEV 724
+I ++HPF+ L SFQTK + L+ D+ GG LF L Q + +ED R Y AE+
Sbjct: 316 DIWTKIEHPFVVQLRYSFQTKYRLYLVLDFVNGGHLFFQLYHQ--GLFREDLARIYTAEI 489
Query: 725 LIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKRKK 784
+ A+ +LH GI++RDLKPEN+L+ +GHV LTDF L+
Sbjct: 490 VSAVSHLHANGIMHRDLKPENILLDADGHVMLTDFGLA---------------------- 603
Query: 785 KKKKGQQKTQQIPTFMAEPMRASNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEML 844
K+ ++ T+ SNS GT EY+APEII G GH A DWW++G+LL+E L
Sbjct: 604 --KQFEESTR------------SNSMCGTLEYMAPEIILGKGHDKAADWWSVGVLLFETL 741
Query: 845 YGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLG-SLEGANE 903
G PF G R K I+ +K P +S +A L+ LL ++ RLG G E
Sbjct: 742 TGKPPFCGGNRDKIQQKIVKDKIKLPAF--LSSEAHSLLKGLLQKEQARRLGCGPRGVEE 915
Query: 904 IKSHPFFKNVNWALI--RCMKP---PELDAPILLENDEKK 938
IKSH +FK +NW + R ++P PE+ + N EK+
Sbjct: 916 IKSHKWFKPINWRKLEAREIQPSFRPEVAGVHCVANFEKR 1035
>TC217519 homologue to PIR|T00410|T00410 protein kinase homolog T13E15.16 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(26%)
Length = 704
Score = 184 bits (466), Expect = 2e-46
Identities = 104/222 (46%), Positives = 129/222 (57%), Gaps = 37/222 (16%)
Frame = +1
Query: 638 GQYFAMKAMDKGVMLNRNKVHRACTEREILDMLDHPFLPALYASFQTKTHVCLITDYYPG 697
G +FAMK MDK + +RNK+ RA TEREIL +LDHPFLP LY F+T CL+ +Y PG
Sbjct: 37 GAFFAMKVMDKASLASRNKLTRAQTEREILQLLDHPFLPTLYTHFETDRFCCLVMEYCPG 216
Query: 698 GELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLT 757
G+L L +QP K E A RFYAAEVL+ALEYLH G++YRDLKPENVL++ +GH+ L+
Sbjct: 217 GDLHTLRQRQPGKHFSEYAARFYAAEVLLALEYLHMLGVVYRDLKPENVLVRDDGHIMLS 396
Query: 758 DFDLSCLTSCKPQLIIPANEDKKKR------------------------------KKKKK 787
DFDLS + P LI + D KR +K KK
Sbjct: 397 DFDLSLRCAVSPTLIRNFDSDPSKRGGGAFCVQPACIEPSSVCIQPSCFMPRLFAQKNKK 576
Query: 788 KGQQKTQ------QIPTFMAEPMRA-SNSFVGTEEYIAPEII 822
K + +P +AEP A S SFVGT EY+APEII
Sbjct: 577 SRTPKAEPGMPSSTLPELVAEPTTARSMSFVGTHEYLAPEII 702
>TC220166 similar to UP|Q9LZS4 (Q9LZS4) Protein kinase-like protein, partial
(19%)
Length = 1017
Score = 182 bits (461), Expect = 8e-46
Identities = 96/222 (43%), Positives = 129/222 (57%), Gaps = 37/222 (16%)
Frame = +2
Query: 742 KPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANE------------------------ 777
KPEN+L++ +GH+ LTDFDLS P L+ +++
Sbjct: 2 KPENILVREDGHIMLTDFDLSLRCDVSPTLLKSSSDVDPAKISGPCAQSSCIEPFCIEPA 181
Query: 778 ------------DKKKRKKKKKKGQQKTQQIPTFMAEPMRA-SNSFVGTEEYIAPEIITG 824
K +K K + + +P +AEP A SNSFVGT EY+APEII G
Sbjct: 182 CQVPCFSPRILPPAAKARKLKTDLAAQLRSLPQLVAEPTDARSNSFVGTHEYLAPEIIKG 361
Query: 825 SGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIY 884
GH +AVDWW G+ LYE+LYG TPF+G ++T AN++ + L+FP + VS Q + LI
Sbjct: 362 EGHGAAVDWWTFGVFLYELLYGRTPFKGSNNEETLANVVLQGLRFPDTPFVSIQGRDLIR 541
Query: 885 WLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPEL 926
LL ++P+NRLGS +GA EIK HPFF+ +NWALIRC PPEL
Sbjct: 542 GLLVKEPENRLGSEKGAAEIKQHPFFEGLNWALIRCAIPPEL 667
>TC221153 similar to UP|Q9LSF1 (Q9LSF1) Protein kinases-like protein, partial
(38%)
Length = 977
Score = 167 bits (424), Expect = 2e-41
Identities = 95/228 (41%), Positives = 129/228 (55%), Gaps = 26/228 (11%)
Frame = +1
Query: 723 EVLIALEYLHCQGIIYRDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPANEDKKKR 782
E+++ALEYLH GI+YRDLKPENV+IQ NGH+ L DFDLS + K + N
Sbjct: 4 ELVLALEYLHNLGIVYRDLKPENVMIQDNGHIMLVDFDLSKKLNPKSPHSLSQNSSPSPN 183
Query: 783 KKKKKKGQQKTQQIPTFM----------AEP---------------MRASNSFVGTEEYI 817
K K+ +Q+ + +F +EP + SNSFVGTEEY+
Sbjct: 184 SKTKQTRKQRLMRFYSFCNSGILPCDSDSEPPLSSVNSVRHTESDLVEKSNSFVGTEEYV 363
Query: 818 APEIITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSP 877
APEI++G GH +VDWW+ G++LYEMLYG TPF+G R++TF IL K+ P+
Sbjct: 364 APEIVSGKGHGFSVDWWSYGVVLYEMLYGTTPFKGSNRKETFYRILMKE---PELTGEKT 534
Query: 878 QAKQLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNW-ALIRCMKPP 924
+ LI LL ++P R+ NEIK H FFK V W ++ +PP
Sbjct: 535 ALRDLIGKLLEKNPDRRI----QVNEIKGHNFFKGVKWDTVLHIARPP 666
>TC209532 similar to UP|O23314 (O23314) Protein kinase, partial (64%)
Length = 1021
Score = 166 bits (421), Expect = 3e-41
Identities = 102/277 (36%), Positives = 155/277 (55%), Gaps = 12/277 (4%)
Frame = +1
Query: 678 LYASFQTKTHVCLITDYYPGGELFLLLDQQPTKVLKEDAVRFYAAEVLIALEYLHCQGII 737
LY SFQ + ++ LI +Y PGG++ LL ++ +L ED RFY E ++A+E +H I
Sbjct: 34 LYCSFQDEEYLYLIMEYLPGGDMMTLLMRKD--ILTEDEARFYVGETVLAIESIHKHNYI 207
Query: 738 YRDLKPENVLIQRNGHVSLTDF----DLSCLTSCKPQLIIPANEDKKKRKKKKKKGQQKT 793
+RD+KP+N+L+ RNGH+ L+DF L C + + N + + ++T
Sbjct: 208 HRDIKPDNLLLDRNGHMKLSDFGLCKPLDCSNLQEKDFSVGMNRSGALQSDGRPAAPKRT 387
Query: 794 Q--QIPTFMAEPMRA-SNSFVGTEEYIAPEIITGSGHTSAVDWWALGILLYEMLYGYTPF 850
Q Q+ + P + S VGT +YIAPE++ G+ DWW+LG ++YEML GY PF
Sbjct: 388 QQEQLQHWQKNPTKCLPYSTVGTPDYIAPEVLLKKGYGVECDWWSLGAIMYEMLVGYPPF 567
Query: 851 RGKTRQKTFANIL--HKDLKFPKSKPVSPQAKQLIYWLLHRDPKNRLGSLEGANEIKSHP 908
T I+ LKFP+ +S +AK LI LL + + RLG+ +GA+EIK+HP
Sbjct: 568 YSHEPMWTCRKIVTWRTTLKFPEEAKLSAEAKDLICRLL-CNVEQRLGT-KGADEIKAHP 741
Query: 909 FFKNVNWALIRCMKP---PELDAPILLENDEKKEAKD 942
+FK V W + M+ PE++ + +N EK E D
Sbjct: 742 WFKGVEWDKLYQMQAAFIPEVNDELDTQNFEKFEEGD 852
>TC210158 weakly similar to UP|YPK2_YEAST (P18961) Serine/threonine-protein
kinase YPK2/YKR2 , partial (5%)
Length = 823
Score = 165 bits (418), Expect = 8e-41
Identities = 84/178 (47%), Positives = 112/178 (62%), Gaps = 3/178 (1%)
Frame = +2
Query: 768 KPQLIIPANEDKKKRKKKKKKGQQ--KTQQIPTFMAEPMRA-SNSFVGTEEYIAPEIITG 824
+P +P +K +K +K KG + +P +AEP A S SFVGT EY+APEII G
Sbjct: 41 QPSCFMPRLFAQKNKKSRKPKGDPGLPSSTLPELVAEPTTARSMSFVGTHEYLAPEIIKG 220
Query: 825 SGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIY 884
GH SAVDWW GI L+E+LYG TPF+G + T N++ + L+FP+S S ++ LI
Sbjct: 221 EGHGSAVDWWTFGIFLHELLYGKTPFKGSGNRATLFNVVGQQLRFPESPATSYASRDLIR 400
Query: 885 WLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPELDAPILLENDEKKEAKD 942
LL ++P++RLG GA EIK HPFF+ VNWALIRC PPE+ P+ + K E D
Sbjct: 401 GLLVKEPQHRLGVKRGATEIKQHPFFEGVNWALIRCSTPPEVPRPVECDLPAKFEPVD 574
>TC224404 homologue to UP|Q9M728 (Q9M728) Viroid symptom modulation protein,
partial (40%)
Length = 829
Score = 163 bits (413), Expect = 3e-40
Identities = 100/254 (39%), Positives = 132/254 (51%), Gaps = 40/254 (15%)
Frame = +1
Query: 741 LKPENVLIQRNGHVSLTDFDLSCLTSCKPQLIIPA-----------NED----------- 778
LKPEN+L++ GH+ L+DFDLS S P L+ + N D
Sbjct: 1 LKPENLLVRDEGHIMLSDFDLSLRCSVNPTLVKSSSAHASNSSSGSNNDVGSILTDDQAV 180
Query: 779 ----------------KKKRKKKKKKG-QQKTQQIPTFMAEPMRA-SNSFVGTEEYIAPE 820
KK RK K G ++P MAEP S SFVGT EY+APE
Sbjct: 181 QSTTQVSSFFPRILPSKKNRKAKSDFGILVGGGRLPELMAEPTNVRSMSFVGTHEYLAPE 360
Query: 821 IITGSGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAK 880
II G GH SAVDWW GI LYE+L+G TPF+G + T N++ + L+FP++ VS A+
Sbjct: 361 IIKGEGHGSAVDWWTFGIFLYELLHGTTPFKGSGYKATLFNVVGQPLRFPETPQVSAVAR 540
Query: 881 QLIYWLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALIRCMKPPELDAPILLENDEKKEA 940
LI LL ++P+ R+ GA EIK HPFF+ +NWAL+R PP + I + K +
Sbjct: 541 DLIRGLLVKEPQKRIAYKRGATEIKQHPFFEGMNWALVRSATPPHIPEVI---DFSKYAS 711
Query: 941 KDIDPGLDDLQKNI 954
KD P D +I
Sbjct: 712 KDTAPPPDKKMADI 753
>TC224169 similar to UP|Q9LT38 (Q9LT38) Protein kinase-like protein, partial
(44%)
Length = 907
Score = 159 bits (402), Expect = 6e-39
Identities = 93/221 (42%), Positives = 123/221 (55%), Gaps = 35/221 (15%)
Frame = +2
Query: 739 RDLKPENVLIQRNGHVSLTDFDLSCLTSCKPQ-------LIIPANEDKKKRKKKK----- 786
RDLKPENVL+Q GH++LTDFDLS + KP+ + +P + + R+K +
Sbjct: 2 RDLKPENVLVQNTGHITLTDFDLSRKLNPKPKPNPQVPSIPLPNSNVPEPRRKHRRNFSR 181
Query: 787 ---------------KKGQQKTQQI---PTFMAEPM----RASNSFVGTEEYIAPEIITG 824
K G +K + P +P SNSFVGTEEY++PE++ G
Sbjct: 182 WISLFPPDGTHHNNNKNGLKKAKSARVSPVSRRKPSFSNGERSNSFVGTEEYVSPEVVRG 361
Query: 825 SGHTSAVDWWALGILLYEMLYGYTPFRGKTRQKTFANILHKDLKFPKSKPVSPQAKQLIY 884
GH AVDWWALGIL+YEMLYG TPF+GK R++TF N++ K F + LI
Sbjct: 362 DGHEFAVDWWALGILIYEMLYGTTPFKGKNRKETFRNVITKPPVFVGKRTA---LTDLIE 532
Query: 885 WLLHRDPKNRLGSLEGANEIKSHPFFKNVNWALI-RCMKPP 924
LL +DP RLG GA EIK H FF+ V W L+ ++PP
Sbjct: 533 KLLEKDPTKRLGYTRGAVEIKEHEFFRGVRWELLTEVVRPP 655
>BG839157
Length = 740
Score = 146 bits (368), Expect = 5e-35
Identities = 88/196 (44%), Positives = 118/196 (59%), Gaps = 8/196 (4%)
Frame = +2
Query: 48 TETEEQHKDFISTDEVTNTSWMAIKEGETGAAAQRAAEWGLVLRTDAETGKPQGVGVRNS 107
T+ + +D S + + S + E A+R AEWGLV+ + +
Sbjct: 143 TKDKSTSEDNYSRNHLNEKSSSIVTEANI---AERTAEWGLVVNSR---------NFKAL 286
Query: 108 GDDEQNGKFSGKRNSNNSGR------VSGDSSDGGDPRG--FPRVSEDLKDALSAFQQTF 159
G + +G F G R+ N S R SG+S+ G + FPRVS++LK+AL+ QQTF
Sbjct: 287 GGENTSGSFDGDRSRNLSDRFVEPTRTSGESNYGSESSSGVFPRVSQELKEALATLQQTF 466
Query: 160 VVSDATKPDYPIMYASAGFFNMTGYTSKEVIGRNCRFLQGADTDPQDVAKIREALEGGKS 219
VVSDATKPD PIMYAS+GFF MTGY+SKE+IGRNCRFLQG +TD +VAKIR+ALE +
Sbjct: 467 VVSDATKPDCPIMYASSGFFTMTGYSSKEIIGRNCRFLQGPETDKNEVAKIRDALE-MEE 643
Query: 220 YCGRLLNYKKDGTPFW 235
+ NY+K + FW
Sbjct: 644 XLWQTXNYRKMDS-FW 688
Score = 61.6 bits (148), Expect = 2e-09
Identities = 35/69 (50%), Positives = 43/69 (61%)
Frame = +2
Query: 448 FLELTEYSREEILGRNCRFLQGPETDPATVRKIREAIDNQTEVTVQLINYTRTGKKFWNL 507
F +T YS +EI+GRNCRFLQGPETD V KIR+A++ + E Q NY R FW +
Sbjct: 521 FFTMTGYSSKEIIGRNCRFLQGPETDKNEVAKIRDALEME-EXLWQTXNY-RKMDSFW-I 691
Query: 508 FHLQPMRDH 516
H P R H
Sbjct: 692 SHXXPSR*H 718
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 39,573,586
Number of Sequences: 63676
Number of extensions: 549622
Number of successful extensions: 9865
Number of sequences better than 10.0: 578
Number of HSP's better than 10.0 without gapping: 4694
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 7366
length of query: 954
length of database: 12,639,632
effective HSP length: 106
effective length of query: 848
effective length of database: 5,889,976
effective search space: 4994699648
effective search space used: 4994699648
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC135230.11


