

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
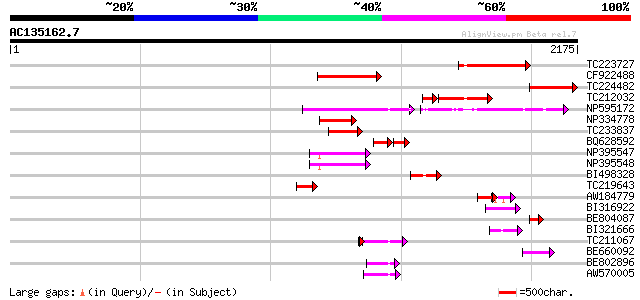
Query= AC135162.7 + phase: 0 /pseudo
(2175 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 372 e-102
CF922488 331 2e-90
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 275 2e-73
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 218 4e-71
NP595172 polyprotein [Glycine max] 228 2e-59
NP334778 reverse transcriptase [Glycine max] 219 8e-57
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 204 3e-52
BQ628592 106 2e-47
NP395547 reverse transcriptase [Glycine max] 188 3e-47
NP395548 reverse transcriptase [Glycine max] 178 3e-44
BI498328 122 2e-27
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 119 1e-26
AW184779 114 6e-25
BI316922 98 4e-20
BE804087 83 1e-15
BI321666 82 2e-15
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 63 1e-14
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 78 4e-14
BE802896 66 1e-10
AW570005 65 2e-10
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 372 bits (954), Expect = e-102
Identities = 171/274 (62%), Positives = 210/274 (76%)
Frame = +1
Query: 1723 NHWNDVPLIKVQRLERPSHVFAIGDVIDQAGENVVDYRPWYYDIKQFLLSREYPPGASKQ 1782
N D+P I+ +P+H + E D +PWY+DIK++++S+EY P +
Sbjct: 46 NAARDLPYIEFWCRGKPAHCCQV--------EEERDGKPWYFDIKRYVISKEYLPEIADN 201
Query: 1783 DKKTLRRLAGRFLLDGNILYKRNYDMVLLRCVDEHEAEQLMHDVHDGTFGTHATGHTMSR 1842
DK+TLRRLA F + G+ILYKRN+DM LRCVD EA ++ +VH+G+FGTHA GH M+R
Sbjct: 202 DKRTLRRLAAGFFMSGSILYKRNHDMKPLRCVDAREANHMIEEVHEGSFGTHANGHAMAR 381
Query: 1843 KLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNVMSSPWPFSMWGIDMIGRI 1902
K+LRAGYYW+ ME DC H RKCHKCQ +AD ++ PPH LNVMSSPWPFSMWGID+IG I
Sbjct: 382 KILRAGYYWLTMESDCCVHVRKCHKCQAFADNVNAPPHPLNVMSSPWPFSMWGIDVIGAI 561
Query: 1903 EPKASNGHRFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKIITDNGTN 1962
EPKASNGHRFILVAIDYFTKWVEAASYT+V + VV +FIK IICRYG+P KIITDNGTN
Sbjct: 562 EPKASNGHRFILVAIDYFTKWVEAASYTDVMRGVVVRFIKKEIICRYGLPRKIITDNGTN 741
Query: 1963 LNNNVVQALCEEFKIEHHNSSPYRPQMNGAVEAA 1996
LNN ++ +CEEFKI+HHN +PYRP+MN AVE A
Sbjct: 742 LNNKMMGEICEEFKIQHHNPTPYRPKMN*AVEVA 843
>CF922488
Length = 741
Score = 331 bits (849), Expect = 2e-90
Identities = 166/246 (67%), Positives = 192/246 (77%)
Frame = +3
Query: 1179 VPKKDGKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSP 1238
V K+DGKV MCVD+RDLN ASPKD FPLPHI+VLVDNT FSFMDGFSGYNQIK++P
Sbjct: 3 VLKEDGKV*MCVDYRDLN*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAP 182
Query: 1239 EDREKTSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSAD 1298
ED EKT+FIT WGTFCYK M FGL N GATYQR M LF DM+HKE+EVY+DDMIVKS
Sbjct: 183 EDMEKTTFITLWGTFCYKAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRT 362
Query: 1299 EEQHVEYLTKMFERLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMP 1358
EE+H+ L K+F RLRKY+LRLNP KC F V+S KLL FI S +GIEVD +KV+ I EM
Sbjct: 363 EEEHLVNLRKLFRRLRKYRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMA 542
Query: 1359 APQTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKNQPIVWNDECQEAFDSIKNYLL 1418
P TEKQV+GFLGRLNYI RFIS + ATC P+F LL KNQ + W+ +C AF+ IK L+
Sbjct: 543 KPHTEKQVQGFLGRLNYIVRFIS*LIATCEPLFILLCKNQFVKWDHDC*VAFERIKQCLI 722
Query: 1419 EPPILV 1424
P +LV
Sbjct: 723 NPHVLV 740
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 275 bits (702), Expect = 2e-73
Identities = 128/182 (70%), Positives = 157/182 (85%)
Frame = +1
Query: 1994 EAANKNIKRIVQKMVTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEAVLPLEVE 2053
EAANKNIK+I+QKM +YKDWHEMLP+ALHGYRT+VR+STGATPFSLVYGMEAVLP EVE
Sbjct: 1 EAANKNIKKIIQKMTVSYKDWHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEVE 180
Query: 2054 IPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAMARGQSYQARMKTAFDKKVHPREFK 2113
+PSLR++ E+ L E+EW Q+RYDQLNLIE KR+ AM+ G+ YQ RMK+AFDKKV R+F
Sbjct: 181 VPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKFH 360
Query: 2114 VGELVLKRRISQQPDPRGKWTPNYEGPYVVKKAFSGGALILTHMDGIELPNPVNADIVKK 2173
G+LVLK+ D RGKW PNYEGP+VVK+AFSGGAL+LT+MDG ELP+P+N+D+VK+
Sbjct: 361 EGDLVLKKMSHAVKDHRGKWAPNYEGPFVVKRAFSGGALVLTNMDGEELPSPMNSDVVKR 540
Query: 2174 YF 2175
Y+
Sbjct: 541 YY 546
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 218 bits (554), Expect(2) = 4e-71
Identities = 103/207 (49%), Positives = 144/207 (68%)
Frame = +2
Query: 1645 IEEAIDMRIKHLDIYGDSALIINQIKGEWETHHANLIPYRDYARRLLTYFTKVELHHIPR 1704
++ AID +K L +YGDSAL+I+Q++GE ET NLIPY+ Y + L +F ++ HH+
Sbjct: 206 VQAAIDSNVKLLKVYGDSALVIHQLRGECETRDPNLIPYQAYIKELAGFFDEISFHHVA* 385
Query: 1705 DENQMADALATLSSMFRVNHWNDVPLIKVQRLERPSHVFAIGDVIDQAGENVVDYRPWYY 1764
+ENQMADALATL SMF++ D+P I+ + RP+H + E D +PWY+
Sbjct: 386 EENQMADALATLVSMFQLTPHGDLPYIEFRCRGRPAHCCLV--------EEERDGKPWYF 541
Query: 1765 DIKQFLLSREYPPGASKQDKKTLRRLAGRFLLDGNILYKRNYDMVLLRCVDEHEAEQLMH 1824
DIK+++ S+EYP AS DK+ RRLA F + G+ILYKRN+DMVLL CV+ E E ++
Sbjct: 542 DIKRYVESKEYPLEASDNDKRRKRRLAAGFFMSGSILYKRNHDMVLLHCVNGKEVENMLG 721
Query: 1825 DVHDGTFGTHATGHTMSRKLLRAGYYW 1851
+VH+G+FGTH+ GH M+RK+LRAGYYW
Sbjct: 722 EVHEGSFGTHSNGHAMARKILRAGYYW 802
Score = 71.6 bits (174), Expect(2) = 4e-71
Identities = 35/60 (58%), Positives = 41/60 (68%), Gaps = 1/60 (1%)
Frame = +1
Query: 1583 LINEGPDPD-SKWGLVFDGAVNAYGKGIGAVIVSPQGHHIPFTTRILFECTNNMAEYEAC 1641
L E D D KW + FD A N G G+GA++VSP IPFTTR+ F+CTNNMAEYEAC
Sbjct: 16 LFEEKLDEDRDKWIVWFDRASNVLGHGVGAILVSPDNQCIPFTTRLGFDCTNNMAEYEAC 195
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 228 bits (581), Expect = 2e-59
Identities = 144/430 (33%), Positives = 225/430 (51%), Gaps = 1/430 (0%)
Frame = +1
Query: 1124 EHRIPTKPECPPVRQKLRRTHPDMALKIKNEVQKQIDAGFLMTVEYPEWVANIVPVPKKD 1183
+H IP K PV+ + R +I+ +Q+ + G + P + I+ V KKD
Sbjct: 1756 DHAIPLKQGSGPVKVRPYRYPHTQKDQIEKMIQEMLVQGIIQPSNSP-FSLPILLVKKKD 1932
Query: 1184 GKVRMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSPEDREK 1243
G R C D+R LN + KD+FP+P +D L+D ++ FS +D SGY+QI + PEDREK
Sbjct: 1933 GSWRFCTDYRALNAITVKDSFPMPTVDELLDELHGAQYFSKLDLRSGYHQILVQPEDREK 2112
Query: 1244 TSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSADEEQHV 1303
T+F T G + + VMPFGL NA AT+Q M +F + K V V+ DD+++ SA + H+
Sbjct: 2113 TAFRTHHGHYEWLVMPFGLTNAPATFQCLMNKIFQFALRKFVLVFFDDILIYSASWKDHL 2292
Query: 1304 EYLTKMFERLRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMPAPQTE 1363
++L + + L++++L +KC+FG LG VS G+ ++ KV+A+ + P P
Sbjct: 2293 KHLESVLQTLKQHQLFARLSKCSFGDTEVDYLGHKVSGLGVSMENTKVQAVLDWPTPNNV 2472
Query: 1364 KQVRGFLGRLNYISRFISHMTATCGPIFKLLRKNQPIVWNDECQEAFDSIKNYLLEPPIL 1423
KQ+RGFLG Y RFI GP+ LL+K+ +WN+E + AF +K + E P+L
Sbjct: 2473 KQLRGFLGLTGYYRRFIKSYANIAGPLTDLLQKDS-FLWNNEAEAAFVKLKKAMTEAPVL 2649
Query: 1424 VPPVEGRPLIMYLAVFDESMGCVLGQQDETGKKEHAIYYLSKKFTDCETRYTMLEKTCCA 1483
P +P I+ +G VLGQ H I Y SKK + + + A
Sbjct: 2650 SLPDFSQPFILETDASGIGVGAVLGQNG------HPIAYFSKKLAPRMQKQSAYTRELLA 2811
Query: 1484 LAWATKRLRHYLVNHTTWLISRMDPIKYIFEKAAVTGKIARWQMLLSEYDIVFKTQ-KAI 1542
+ A + RHYL+ + + + +K + +++ T + W YD FK + K
Sbjct: 2812 ITEALSKFRHYLLGNKFIIRTDQRSLKSLMDQSLQTPEQQAWLHKFLGYD--FKIEYKPG 2985
Query: 1543 KGSILADHLA 1552
K + AD L+
Sbjct: 2986 KDNQAADALS 3015
Score = 95.1 bits (235), Expect = 3e-19
Identities = 147/585 (25%), Positives = 240/585 (40%), Gaps = 15/585 (2%)
Frame = +1
Query: 1574 LKSKDCEEPLINEGPDPDSKWGLVFDGAVNAYGKGIGAVIVSPQGHHIPFTTRILFECTN 1633
LK E P+++ PD + + +A G G+GAV+ GH I + ++ L
Sbjct: 2617 LKKAMTEAPVLSL---PDFSQPFILE--TDASGIGVGAVL-GQNGHPIAYFSKKLAPRMQ 2778
Query: 1634 NMAEYEACIFGIEEAIDMRIKHLDIYGDSALIINQIKGEWETHHANLIPYRDYA--RRLL 1691
+ Y + I EA+ + +H + G+ +I + +L A + L
Sbjct: 2779 KQSAYTRELLAITEALS-KFRHY-LLGNKFIIRTDQRSLKSLMDQSLQTPEQQAWLHKFL 2952
Query: 1692 TYFTKVELHHIPRDENQMADALATLSSMFRVNHWNDVPLIKVQRLERPSHVFAIGDVIDQ 1751
Y K+E + P +NQ ADAL S MF + W++ I ++ L I D
Sbjct: 2953 GYDFKIE--YKPGKDNQAADAL---SRMFMLA-WSEPHSIFLEELRAR----LISDP--- 3093
Query: 1752 AGENVVDYRPWYYDIKQFLLSREYPPGASKQDKKTLRRLAGRFLLDGNILYKRNYDMVLL 1811
+KQ L Y GA A + + +LY + D V++
Sbjct: 3094 -------------HLKQ--LMETYKQGAD----------ASHYTVREGLLYWK--DRVVI 3192
Query: 1812 RCVDEHEAEQLMHDVHDGTFGTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIY 1871
+E +++ + H G HA G T + L+A +YW M+ D + +KC CQ
Sbjct: 3193 PA-EEEIVNKILQEYHSSPIGGHA-GITRTLARLKAQFYWPKMQEDVKAYIQKCLICQQA 3366
Query: 1872 ADKIHVPPHALNVMSSPWPFSMW---GIDMIGRIEPKASNGHRFILVAIDYFTKWVEAAS 1928
+P L + P P +W +D I + S G I+V ID TK+
Sbjct: 3367 KSNNTLPAGLLQPL--PIPQQVWEDVAMDFITGLPN--SFGLSVIMVVIDRLTKYAHFIP 3534
Query: 1929 Y-TNVTKQVVAKFIKNNIICRYGVPSKIITDNGTNLNNNVVQALCEEFKIEHHN---SSP 1984
+ +VVA+ ++I+ +G+P I++D + Q L FK++ SS
Sbjct: 3535 LKADYNSKVVAEAFMSHIVKLHGIPRSIVSDRDRVFTSTFWQHL---FKLQGTTLAMSSA 3705
Query: 1985 YRPQMNGAVEAANKNIKRIVQKMVTTY-KDWHEMLPYALHGYRTTVRSSTGATPFSLVYG 2043
Y PQ +G E NK ++ ++ + K W + LP+A Y T S G TPF +YG
Sbjct: 3706 YHPQSDGQSEVLNKCLEMYLRCFTYEHPKGWVKALPWAEFWYNTAYHMSLGMTPFRALYG 3885
Query: 2044 MEAVLPLEVEIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRMDAMARGQSYQARMKTAF 2103
E P+L + AE + D+ L+ + +++ + R Q MK
Sbjct: 3886 REP--------PTLTRQACSIDDPAEVREQLTDRDALLAKLKIN-LTRAQQV---MKRQA 4029
Query: 2104 DKKVHPREFKVGELVL-----KRRISQQPDPRGKWTPNYEGPYVV 2143
DKK F++G+ VL R+ S K + Y GP+ V
Sbjct: 4030 DKKRLDVSFQIGDEVLVKLQPYRQHSAVLRKNQKLSMRYFGPFKV 4164
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 219 bits (559), Expect = 8e-57
Identities = 101/143 (70%), Positives = 117/143 (81%)
Frame = +3
Query: 1187 RMCVDFRDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSPEDREKTSF 1246
RMCVD+RDLN+ASPKDNFPLPHID+L+ N A +FSFMDGFSGYNQIKM+PED EKT+F
Sbjct: 3 RMCVDYRDLNRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTF 182
Query: 1247 ITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSADEEQHVEYL 1306
IT WGTFCYKVM FGL N GATY R M LF DM+HKE+E YVD+MI KS EE+H+ L
Sbjct: 183 ITLWGTFCYKVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNL 362
Query: 1307 TKMFERLRKYKLRLNPNKCTFGV 1329
+F +LRKY+LRLNP KC FG+
Sbjct: 363 QNLFGQLRKYRLRLNPRKCVFGL 431
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 204 bits (520), Expect = 3e-52
Identities = 96/133 (72%), Positives = 113/133 (84%)
Frame = +2
Query: 1222 FSFMDGFSGYNQIKMSPEDREKTSFITPWGTFCYKVMPFGLINAGATYQRGMTTLFHDMI 1281
FSFMDGFSGYNQI M+ ED EKT+F+T WGTF Y+VM FGL N GATYQR M LFHDM+
Sbjct: 2 FSFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMM 181
Query: 1282 HKEVEVYVDDMIVKSADEEQHVEYLTKMFERLRKYKLRLNPNKCTFGVRSGKLLGFIVSQ 1341
HKE+EVYVDDMI KS E +H+ L K+F RL+KY+L+LNP KCTFGV+SGKLLGFIVSQ
Sbjct: 182 HKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVSQ 361
Query: 1342 KGIEVDPDKVRAI 1354
KGIE+DP+KV+A+
Sbjct: 362 KGIEIDPEKVKAL 400
>BQ628592
Length = 423
Score = 106 bits (264), Expect(2) = 2e-47
Identities = 48/76 (63%), Positives = 58/76 (76%)
Frame = -1
Query: 1394 LRKNQPIVWNDECQEAFDSIKNYLLEPPILVPPVEGRPLIMYLAVFDESMGCVLGQQDET 1453
L KNQ I+WN QEAF+ IK L P +L+PPV GRP ++Y+ + DESMGCVL Q D++
Sbjct: 423 LPKNQAILWNSNYQEAFEKIKQSLANPSVLMPPVTGRPFLLYMTMLDESMGCVLVQHDDS 244
Query: 1454 GKKEHAIYYLSKKFTD 1469
GKKE AIYYLSKKFTD
Sbjct: 243 GKKEQAIYYLSKKFTD 196
Score = 104 bits (259), Expect(2) = 2e-47
Identities = 43/61 (70%), Positives = 56/61 (91%)
Frame = -3
Query: 1474 YTMLEKTCCALAWATKRLRHYLVNHTTWLISRMDPIKYIFEKAAVTGKIARWQMLLSEYD 1533
Y+MLE+TCC L WA+ RLR Y+++HTTWLIS+MDP+KYIFEK A+TG+IARWQ+LLSE++
Sbjct: 184 YSMLERTCCTLVWASHRLRQYMLSHTTWLISKMDPVKYIFEKPALTGRIARWQVLLSEFN 5
Query: 1534 I 1534
I
Sbjct: 4 I 2
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 188 bits (477), Expect = 3e-47
Identities = 96/252 (38%), Positives = 144/252 (57%), Gaps = 18/252 (7%)
Frame = +1
Query: 1151 IKNEVQKQIDAGFLMTVEYPEWVANIVPVPKKDGKV------------------RMCVDF 1192
++ EV K ++AG + + WV+ + VPKK G RMC+D+
Sbjct: 1 VRKEVFKLLEAGLIYPISDSSWVSPVQVVPKKGGMTVVKNDRNELIPTRRVTRWRMCIDY 180
Query: 1193 RDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSPEDREKTSFITPWGT 1252
R LN+A+ KD++PLP +D ++ A+ + F+DG+SGYNQI + P+D+EKT+F P+
Sbjct: 181 RKLNEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSV 360
Query: 1253 FCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSADEEQHVEYLTKMFER 1312
F Y+ MPFGL NA T+QR M +F DM+ K +EV++DD A + L K+ +R
Sbjct: 361 FAYRRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQR 540
Query: 1313 LRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMPAPQTEKQVRGFLGR 1372
K L LN KC F V+ G +LG +S++GIEV +K+ I ++P P K + FLG
Sbjct: 541 CEKSNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGH 720
Query: 1373 LNYISRFISHMT 1384
+ + RFI T
Sbjct: 721 VGFYRRFIKDFT 756
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 178 bits (451), Expect = 3e-44
Identities = 90/252 (35%), Positives = 142/252 (55%), Gaps = 18/252 (7%)
Frame = +1
Query: 1151 IKNEVQKQIDAGFLMTVEYPEWVANIVPVPKKDGKV------------------RMCVDF 1192
++ EV K ++ G + + WV+ ++ V KK+G ++C+D+
Sbjct: 1 VRKEVLKLLEVGLIYPISDSAWVSPVLVVSKKEGMTVIRNEKNDLIPTRTVTSWKLCIDY 180
Query: 1193 RDLNKASPKDNFPLPHIDVLVDNTAQSKVFSFMDGFSGYNQIKMSPEDREKTSFITPWGT 1252
R LN+A+ KD+FPLP +D +++ A + F+D + GYNQI + P+D+EK +F P+G
Sbjct: 181 RKLNEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGV 360
Query: 1253 FCYKVMPFGLINAGATYQRGMTTLFHDMIHKEVEVYVDDMIVKSADEEQHVEYLTKMFER 1312
F Y+ +PFGL NA T+Q M +F D++ K +EV++DD V E ++ L + +R
Sbjct: 361 FAYRRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQR 540
Query: 1313 LRKYKLRLNPNKCTFGVRSGKLLGFIVSQKGIEVDPDKVRAIREMPAPQTEKQVRGFLGR 1372
+ L LN KC F VR G +LG +S +GIEVD K+ I ++P P K +R FLG+
Sbjct: 541 CVETNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQ 720
Query: 1373 LNYISRFISHMT 1384
+ RFI T
Sbjct: 721 ARFYRRFIKDFT 756
>BI498328
Length = 335
Score = 122 bits (305), Expect = 2e-27
Identities = 63/117 (53%), Positives = 78/117 (65%)
Frame = +1
Query: 1538 TQKAIKGSILADHLAYQPLDDYQPIEFDFPDEEIMYLKSKDCEEPLINEGPDPDSKWGLV 1597
TQKA+KGS LAD+LA PL Y+P+ +FPDE+IM L + IN KW +
Sbjct: 4 TQKAVKGSALADYLAQ*PLQGYRPMHPEFPDEDIMALFEEKRTHEDIN-------KWIVC 162
Query: 1598 FDGAVNAYGKGIGAVIVSPQGHHIPFTTRILFECTNNMAEYEACIFGIEEAIDMRIK 1654
FDGA NA G G+GAV+VSP IPFT R+ F+CTNNMAEYEAC G++ AID +K
Sbjct: 163 FDGASNALGHGVGAVLVSPDDQCIPFTARLGFDCTNNMAEYEACALGVQAAIDFDVK 333
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein (Fragment),
partial (8%)
Length = 1320
Score = 119 bits (299), Expect = 1e-26
Identities = 53/80 (66%), Positives = 63/80 (78%)
Frame = +3
Query: 1101 LREYPDIFAWSYEDMPGLDPMIVEHRIPTKPECPPVRQKLRRTHPDMALKIKNEVQKQID 1160
LR+Y DIFAWSY+DMPGL IV+HR+P PEC PV+QKLRR P+ +LKIK EV+K D
Sbjct: 1080 LRDYQDIFAWSYQDMPGLSSDIVQHRLPLNPECSPVKQKLRRMKPETSLKIKEEVKK*FD 1259
Query: 1161 AGFLMTVEYPEWVANIVPVP 1180
AGFL YP+WVANIVP+P
Sbjct: 1260 AGFLAVARYPKWVANIVPIP 1319
>AW184779
Length = 432
Score = 114 bits (284), Expect = 6e-25
Identities = 49/74 (66%), Positives = 59/74 (79%)
Frame = +1
Query: 1796 LDGNILYKRNYDMVLLRCVDEHEAEQLMHDVHDGTFGTHATGHTMSRKLLRAGYYWMAME 1855
L NILYKRN+DMVLLRCVD EAEQ++ +VH+G+FGTHA H M++K+LR GYYW+ ME
Sbjct: 1 LSRNILYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTME 180
Query: 1856 HDCYQHARKCHKCQ 1869
DC H KCHKCQ
Sbjct: 181 SDCCIHVWKCHKCQ 222
Score = 84.7 bits (208), Expect = 4e-16
Identities = 49/114 (42%), Positives = 59/114 (50%), Gaps = 22/114 (19%)
Frame = +3
Query: 1848 GYYWMAMEHDCY-------------QHARKCHKCQIYADKIHVPPHALNVMSSPWPF--- 1891
G W +H C+ R H C + VP +M +P P+
Sbjct: 99 GILWNTCQHTCHGPEDSESGVLLAHYGERLLHPC---VEMP*VPVRPSLIMLTPHPYL*M 269
Query: 1892 ------SMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYTNVTKQVVAK 1939
MWGID+IG IEPKASNGH FILVAIDYFTKWVEA SY +VT+ VV +
Sbjct: 270 SWQHLGHMWGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVSYASVTRSVVIR 431
>BI316922
Length = 405
Score = 97.8 bits (242), Expect = 4e-20
Identities = 45/135 (33%), Positives = 80/135 (58%)
Frame = +3
Query: 1825 DVHDGTFGTHATGHTMSRKLLRAGYYWMAMEHDCYQHARKCHKCQIYADKIHVPPHALNV 1884
++H G G H+ M+ ++LR GYY M C ++ +KC +C + + H+ L+
Sbjct: 3 EMHRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHN 182
Query: 1885 MSSPWPFSMWGIDMIGRIEPKASNGHRFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNN 1944
+ +PWPF++ G+D++ R P + +++LV ID FTKW+E ++ V KF+ N
Sbjct: 183 IVAPWPFAI*GVDIL-RPFPLSKRQVKYLLVGIDQFTKWIETEHIAIISIANVRKFV*RN 359
Query: 1945 IICRYGVPSKIITDN 1959
I+C +G+P+ +I+DN
Sbjct: 360 IVC*FGIPNTLISDN 404
>BE804087
Length = 160
Score = 82.8 bits (203), Expect = 1e-15
Identities = 38/53 (71%), Positives = 44/53 (82%)
Frame = +2
Query: 1994 EAANKNIKRIVQKMVTTYKDWHEMLPYALHGYRTTVRSSTGATPFSLVYGMEA 2046
EAANKNIK+ + KM +YKDWHEM +ALH YRT VR+STGATP+SLVYG EA
Sbjct: 2 EAANKNIKKNI*KMTVSYKDWHEMFSFALHMYRTLVRTSTGATPYSLVYGKEA 160
>BI321666
Length = 430
Score = 82.4 bits (202), Expect = 2e-15
Identities = 45/129 (34%), Positives = 71/129 (54%), Gaps = 3/129 (2%)
Frame = +2
Query: 1840 MSRKLLRAGYYWMAMEHDCYQHARKCHKCQ---IYADKIHVPPHALNVMSSPWPFSMWGI 1896
+S +L++ +Y ++ D Y HA+ C+KCQ + + +P H + + F WGI
Sbjct: 2 ISTNVLQSRFYLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEI---FDYWGI 172
Query: 1897 DMIGRIEPKASNGHRFILVAIDYFTKWVEAASYTNVTKQVVAKFIKNNIICRYGVPSKII 1956
D +G P SN +ILV +DY +KWVEA + ++V KF+K I R GVP +I
Sbjct: 173 DFVGPFPPSFSN--EYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLI 346
Query: 1957 TDNGTNLNN 1965
+ G++L N
Sbjct: 347 DNGGSHLCN 373
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 63.2 bits (152), Expect(2) = 1e-14
Identities = 42/171 (24%), Positives = 74/171 (42%)
Frame = +1
Query: 1355 REMPAPQTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKNQPIVWNDECQEAFDSIK 1414
R P ++ +R F G ++ RF+ + + P+ +L++KN W ++ ++AF +K
Sbjct: 88 RMAPTLKSVGDIRSFHGLASFYRRFVPNFSTVASPLNELVKKNMAFTWGEKQEQAFALLK 267
Query: 1415 NYLLEPPILVPPVEGRPLIMYLAVFDESMGCVLGQQDETGKKEHAIYYLSKKFTDCETRY 1474
L + P+L P + + + VL Q H I Y S+K Y
Sbjct: 268 EKLTKAPVLALPDFSKTFELECDASGVGVRAVLLQGG------HPIAYFSEKLHSATLNY 429
Query: 1475 TMLEKTCCALAWATKRLRHYLVNHTTWLISRMDPIKYIFEKAAVTGKIARW 1525
+K AL A + H+LV + S +KYI K+ + + A+W
Sbjct: 430 PTYDKELYALIRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKW 582
Score = 36.6 bits (83), Expect(2) = 1e-14
Identities = 12/25 (48%), Positives = 21/25 (84%)
Frame = +2
Query: 1336 GFIVSQKGIEVDPDKVRAIREMPAP 1360
GF+V + G+++DP+K++AI+E P P
Sbjct: 29 GFVVGRNGVQMDPEKIKAIQEWPPP 103
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 78.2 bits (191), Expect = 4e-14
Identities = 41/122 (33%), Positives = 71/122 (57%), Gaps = 1/122 (0%)
Frame = -3
Query: 1968 VQALCEEFKIEHHNSSPYRPQMNGAVEAANKNIKRIVQKMVT-TYKDWHEMLPYALHGYR 2026
+ AL +++ + H S+PY PQ NG E +N+ IKRI++K+V + KDW L AL +R
Sbjct: 376 MHALLKKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHR 197
Query: 2027 TTVRSSTGATPFSLVYGMEAVLPLEVEIPSLRVIMEAKLSEAEWCQSRYDQLNLIEEKRM 2086
T ++ G +P+ +V+G LP+E+E + + S + + R QL+ ++E R+
Sbjct: 196 TAYKAPIGMSPYRVVFGKACHLPVEIEHKAYWAVKTCNFSMDQAGEERKLQLSELDEIRL 17
Query: 2087 DA 2088
+A
Sbjct: 16 EA 11
>BE802896
Length = 416
Score = 66.2 bits (160), Expect = 1e-10
Identities = 39/128 (30%), Positives = 61/128 (47%)
Frame = -2
Query: 1369 FLGRLNYISRFISHMTATCGPIFKLLRKNQPIVWNDECQEAFDSIKNYLLEPPILVPPVE 1428
FLG + RFI P+ LL+K +ND+C+ AFD +K L+ PI+ P
Sbjct: 412 FLGHAGFYRRFIRDFRKVALPLSNLLQKEVEFDFNDKCK*AFDCLKRALITTPIIQAPDW 233
Query: 1429 GRPLIMYLAVFDESMGCVLGQQDETGKKEHAIYYLSKKFTDCETRYTMLEKTCCALAWAT 1488
P + + ++G VL Q+ + K IYY S+ + YT EK A+ +A
Sbjct: 232 TAPFELMCDASNYALGVVLAQKID--KLPRVIYYSSRTLDAAQANYTTTEKELLAIVFAL 59
Query: 1489 KRLRHYLV 1496
++ YL+
Sbjct: 58 EKFHSYLL 35
>AW570005
Length = 413
Score = 65.5 bits (158), Expect = 2e-10
Identities = 42/141 (29%), Positives = 62/141 (43%)
Frame = -2
Query: 1358 PAPQTEKQVRGFLGRLNYISRFISHMTATCGPIFKLLRKNQPIVWNDECQEAFDSIKNYL 1417
P P+T + +RGFL + RFI A P+ LL K+ VW+ E AF ++KN +
Sbjct: 412 PPPRTARSLRGFLRLTGFYRRFIKGYAAMAAPLSHLLTKDS-FVWSPEADVAFQALKNVV 236
Query: 1418 LEPPILVPPVEGRPLIMYLAVFDESMGCVLGQQDETGKKEHAIYYLSKKFTDCETRYTML 1477
+L P +P + MG VL Q+ H I + SK+F R +
Sbjct: 235 TNTLVLALPDFTKPFTVETDASGSDMGAVLSQEG------HPIAFFSKEFCPKLVRSSTY 74
Query: 1478 EKTCCALAWATKRLRHYLVNH 1498
A+ K+ R YL+ H
Sbjct: 73 VHELAAITNVVKKWRQYLLGH 11
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.340 0.148 0.492
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 101,573,193
Number of Sequences: 63676
Number of extensions: 1548610
Number of successful extensions: 12434
Number of sequences better than 10.0: 57
Number of HSP's better than 10.0 without gapping: 11880
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 12411
length of query: 2175
length of database: 12,639,632
effective HSP length: 112
effective length of query: 2063
effective length of database: 5,507,920
effective search space: 11362838960
effective search space used: 11362838960
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.9 bits)
S2: 67 (30.4 bits)
Medicago: description of AC135162.7


