

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135162.11 + phase: 0 /pseudo
(385 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
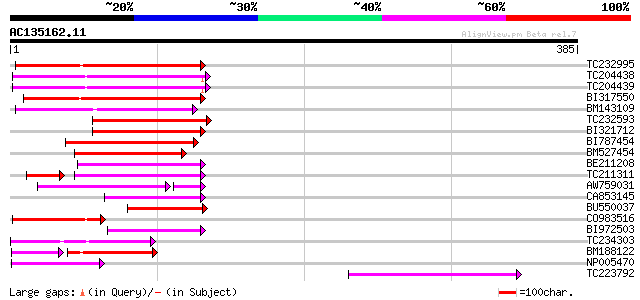
Score E
Sequences producing significant alignments: (bits) Value
TC232995 100 1e-21
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 95 6e-20
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 94 1e-19
BI317550 weakly similar to GP|9759590|dbj| polyprotein-like {Ara... 93 2e-19
BM143109 90 1e-18
TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (F... 77 2e-14
BI321712 75 5e-14
BI787454 weakly similar to GP|21434|emb|CA ORF4 {Solanum tuberos... 71 9e-13
BM527454 weakly similar to GP|27901709|gb| gag-pol polyprotein {... 69 3e-12
BE211208 64 1e-10
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 47 4e-10
AW759031 weakly similar to GP|22093573|db polyprotein {Oryza sat... 61 1e-09
CA853145 weakly similar to GP|27901709|gb| gag-pol polyprotein {... 59 3e-09
BU550037 57 2e-08
CO983516 54 9e-08
BI972503 weakly similar to GP|18378607|gb polyprotein {Oryza sat... 54 1e-07
TC234303 weakly similar to UP|Q8W153 (Q8W153) Polyprotein, parti... 52 5e-07
BM188122 weakly similar to PIR|I46200|I4620 retrovirus-related r... 44 3e-06
NP005470 reverse transcriptase 46 2e-05
TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polypro... 42 4e-04
>TC232995
Length = 1009
Score = 100 bits (249), Expect = 1e-21
Identities = 48/129 (37%), Positives = 85/129 (65%)
Frame = +2
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDL 64
+++L KALYGLKQ+ RAW +R+ NFL+++EF + K + +++K K ++++ + +YVDD+
Sbjct: 47 VYKLQKALYGLKQAPRAWYERLSNFLLEKEFSRGKVDTTLFIKR-KHNDILLVQIYVDDI 223
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
I +N + + F M +F M+ +G+L YFLG++ +T G+ +Q KY +++KRF
Sbjct: 224 IFGSTNDSLCKEFSLDMQSEFEMSMMGELKYFLGLQIKQTQ*GIFINQSKYCKELIKRFG 403
Query: 125 MLNCNPAST 133
M + ST
Sbjct: 404 MDSAKHMST 430
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 94.7 bits (234), Expect = 6e-20
Identities = 52/138 (37%), Positives = 81/138 (58%), Gaps = 4/138 (2%)
Frame = +1
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D ++RL KALYGLKQ+ RAW +R+ FL QQ + K + ++VK + NL+ +YVD
Sbjct: 3616 DHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAE-NLMIAQIYVD 3792
Query: 63 DLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKR 122
D++ + + F M +F M+ +G+LTYFLG++ + + + Q KYA +I+K+
Sbjct: 3793 DIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSKYAKNIVKK 3972
Query: 123 FNMLNCN----PASTLFK 136
F M N + PA T K
Sbjct: 3973 FGMENASHKRTPAPTHLK 4026
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 93.6 bits (231), Expect = 1e-19
Identities = 51/138 (36%), Positives = 81/138 (57%), Gaps = 4/138 (2%)
Frame = +1
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D ++RL KALYGLKQ+ RAW +R+ FL QQ + K + ++VK + NL+ +YVD
Sbjct: 3613 DHVYRLKKALYGLKQAPRAWYERLTEFLTQQGYRKGGIDKTLFVKQDAE-NLMIAQIYVD 3789
Query: 63 DLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKR 122
D++ + + F M +F M+ +G+LTYFLG++ + + + Q +YA +I+K+
Sbjct: 3790 DIVFGGMSNEMLRHFVQQMQSEFEMSLVGELTYFLGLQVKQMEDSIFLSQSRYAKNIVKK 3969
Query: 123 FNMLNCN----PASTLFK 136
F M N + PA T K
Sbjct: 3970 FGMENASHKRTPAPTHLK 4023
>BI317550 weakly similar to GP|9759590|dbj| polyprotein-like {Arabidopsis
thaliana}, partial (18%)
Length = 421
Score = 92.8 bits (229), Expect = 2e-19
Identities = 43/124 (34%), Positives = 77/124 (61%)
Frame = -2
Query: 10 KALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDLIVTDS 69
K+LYGLKQ+ R W +++ N L+++ +++ ++Y+++ +K + + +YVDD+I+
Sbjct: 420 KSLYGLKQASRKWYEKLTNLLLKEGYIQSISDYSLFTL-TKGNTFTALLVYVDDIILAGD 244
Query: 70 NLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFNMLNCN 129
++ E + K + F + +LGKL YFLG+E + G+ Q+KY D+LK +L C
Sbjct: 243 SIDEFDRIKNVLDLAFKIKNLGKLKYFLGLEVAHSRLGITISQRKYCLDLLKDSGLLGCK 64
Query: 130 PAST 133
PAST
Sbjct: 63 PAST 52
>BM143109
Length = 415
Score = 90.1 bits (222), Expect = 1e-18
Identities = 48/124 (38%), Positives = 74/124 (58%), Gaps = 1/124 (0%)
Frame = +1
Query: 5 IFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYV-KNSKDSNLVHICLYVDD 63
+F+L K LYGLKQ+LRAW + + FL+ + F K K + N+++ K D LV I YVDD
Sbjct: 43 VFKLKKVLYGLKQALRAWYELLSKFLLDKGFSKGKVDTNLFI*KKLNDILLVQI--YVDD 216
Query: 64 LIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRF 123
+I +N + + F M +F M+ + +L +FLG++ +T G+ Q KY D++ RF
Sbjct: 217 IIFGSTNDSLCKKFSQDMQNEFEMSMMRELNFFLGLQIKQTKNGIFISQSKYCKDLIHRF 396
Query: 124 NMLN 127
M N
Sbjct: 397 GMEN 408
>TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (Fragment),
partial (16%)
Length = 562
Score = 76.6 bits (187), Expect = 2e-14
Identities = 37/81 (45%), Positives = 52/81 (63%)
Frame = +1
Query: 57 ICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYA 116
+ LYVDDL+VT + +E FK MM+ F MT+LG +TYFLG+E ++ ++ Q+KYA
Sbjct: 199 VSLYVDDLLVTRDDARLVEEFKQEMMQAFEMTNLGLMTYFLGIEIKQSQNKVLICQRKYA 378
Query: 117 SDILKRFNMLNCNPASTLFKQ 137
+ILK+F M C ST Q
Sbjct: 379 KEILKKFQMEECKSVSTPMNQ 441
>BI321712
Length = 399
Score = 75.1 bits (183), Expect = 5e-14
Identities = 34/77 (44%), Positives = 50/77 (64%)
Frame = -3
Query: 57 ICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYA 116
+CLYVDDLI T +N + E FK M +F MTD+G + Y+LG+E + +G+ Q+ YA
Sbjct: 385 LCLYVDDLIFTGNNPSMFEEFKKDMSNEFEMTDMGLMAYYLGIEVKQEDKGIFITQEGYA 206
Query: 117 SDILKRFNMLNCNPAST 133
++LK+F M + NP T
Sbjct: 205 KEVLKKFKMDDANPVGT 155
>BI787454 weakly similar to GP|21434|emb|CA ORF4 {Solanum tuberosum}, partial
(21%)
Length = 421
Score = 70.9 bits (172), Expect = 9e-13
Identities = 32/90 (35%), Positives = 55/90 (60%)
Frame = +2
Query: 39 KAEYNVYVKNSKDSNLVHICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLG 98
+A+++V+ ++ V++ +YVDD+++T + +I K H+ F DL L YFLG
Sbjct: 11 EADHSVFYCHTSPGKCVYLMVYVDDIMITKKDATKIVQLKEHLFNHFQTKDLRYLKYFLG 190
Query: 99 MEFTKTSEGLVKHQKKYASDILKRFNMLNC 128
+E ++ +G+V Q+KYA DIL+ M NC
Sbjct: 191 IEVAQSGDGVVISQRKYALDILEETGMQNC 280
>BM527454 weakly similar to GP|27901709|gb| gag-pol polyprotein {Vitis
vinifera}, partial (19%)
Length = 437
Score = 69.3 bits (168), Expect = 3e-12
Identities = 29/76 (38%), Positives = 49/76 (64%)
Frame = +2
Query: 45 YVKNSKDSNLVHICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKT 104
++ + S V++ +YVDD+++T ++ +I KGH+ F DLGK YFLG+E ++
Sbjct: 5 FLLSYSSSRCVYLMVYVDDIVITGNDQGKIAQLKGHLFSHFQTKDLGKFEYFLGIEVAQS 184
Query: 105 SEGLVKHQKKYASDIL 120
+G++ Q+KYA DIL
Sbjct: 185 KDGIIISQRKYALDIL 232
>BE211208
Length = 413
Score = 63.5 bits (153), Expect = 1e-10
Identities = 32/88 (36%), Positives = 52/88 (58%), Gaps = 1/88 (1%)
Frame = +2
Query: 47 KNSKDSNLVHICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSE 106
+ KD NLV++ +YVDD+I+T + I++ H+ F++ LG+L YFLG+E T
Sbjct: 2 RGKKDRNLVYLLVYVDDIIITGRSNYLIQSLVHHLNSNFSLKQLGQLDYFLGIEVHHTPT 181
Query: 107 G-LVKHQKKYASDILKRFNMLNCNPAST 133
G ++ Q KY D+L + +M P S+
Sbjct: 182 GSVLLTQSKYICDLLHKTDMAEAKPISS 265
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 47.4 bits (111), Expect(2) = 4e-10
Identities = 27/89 (30%), Positives = 43/89 (47%)
Frame = +3
Query: 45 YVKNSKDSNLVHICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKT 104
Y + K + I +YVDD+I ++ + F M F + G+L + LG++ +
Sbjct: 459 YSERLKKETFLIIHIYVDDIIFGATSKRMCKEFFELMKDGFETSMKGELKFLLGLQIIQK 638
Query: 105 SEGLVKHQKKYASDILKRFNMLNCNPAST 133
G+ HQ+KY LKRF M P +T
Sbjct: 639 VYGIFIHQEKYTKSHLKRFRMDEAKPMAT 725
Score = 34.7 bits (78), Expect(2) = 4e-10
Identities = 14/26 (53%), Positives = 20/26 (76%)
Frame = +2
Query: 12 LYGLKQSLRAWNKRIVNFLVQQEFVK 37
+YGLKQ+LRAW +R+ +FLV F +
Sbjct: 362 VYGLKQALRAWYERLSSFLVSNGFTR 439
>AW759031 weakly similar to GP|22093573|db polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (5%)
Length = 430
Score = 60.8 bits (146), Expect(2) = 1e-09
Identities = 28/90 (31%), Positives = 51/90 (56%)
Frame = +3
Query: 20 RAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVDDLIVTDSNLAEIEAFKG 79
RAW R + L+Q + +A+++V+ +S +++ +Y+DD+++T + I K
Sbjct: 6 RAWFGRFSSELLQFCMTRYEADHSVFSLHSPSGLCIYLVVYLDDIVITGDDFDGIHRLKS 185
Query: 80 HMMKKFNMTDLGKLTYFLGMEFTKTSEGLV 109
H+ KF DLG YFLG+E ++S G +
Sbjct: 186 HLHNKFQTKDLGPPNYFLGIEVAQSSSGFL 275
Score = 19.6 bits (39), Expect(2) = 1e-09
Identities = 9/22 (40%), Positives = 13/22 (58%)
Frame = +2
Query: 112 QKKYASDILKRFNMLNCNPAST 133
Q+KY DIL +ML+ + T
Sbjct: 281 QRKYVLDILIETSMLDFRHSDT 346
>CA853145 weakly similar to GP|27901709|gb| gag-pol polyprotein {Vitis
vinifera}, partial (29%)
Length = 563
Score = 59.3 bits (142), Expect = 3e-09
Identities = 28/69 (40%), Positives = 39/69 (55%)
Frame = +3
Query: 65 IVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFN 124
++T + I K H+ + F DLG+L YFLG+E ++ G+ Q+KYA DILK
Sbjct: 12 VITRDDHRGIRDLKSHLFQHFQTKDLGQLKYFLGIEVAQSKTGIAICQRKYALDILKDTG 191
Query: 125 MLNCNPAST 133
M NC P T
Sbjct: 192 MQNCKPVDT 218
>BU550037
Length = 728
Score = 56.6 bits (135), Expect = 2e-08
Identities = 25/54 (46%), Positives = 38/54 (70%)
Frame = -3
Query: 81 MMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFNMLNCNPASTL 134
M +F MTDLG++ YFLGM ++ +G+ QKKYA +IL++F+M C P +T+
Sbjct: 363 MESEFEMTDLGQMKYFLGM*IFQSEDGIFISQKKYAWEILRKFHMERCKPIATV 202
>CO983516
Length = 724
Score = 54.3 bits (129), Expect = 9e-08
Identities = 27/63 (42%), Positives = 41/63 (64%)
Frame = +2
Query: 3 DMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYVD 62
D ++RL KALYGLKQ+ RAW +R+ L QQ + K + ++VK + NL+ +YVD
Sbjct: 533 DHVYRLKKALYGLKQAPRAWYERLTELLTQQGYRKGGIDKTLFVKQDAE-NLMIAQIYVD 709
Query: 63 DLI 65
D++
Sbjct: 710 DIV 718
>BI972503 weakly similar to GP|18378607|gb polyprotein {Oryza sativa
(japonica cultivar-group)}, partial (5%)
Length = 327
Score = 53.9 bits (128), Expect = 1e-07
Identities = 26/67 (38%), Positives = 37/67 (54%)
Frame = +2
Query: 67 TDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGMEFTKTSEGLVKHQKKYASDILKRFNML 126
T ++ +I K H+ F DLG L YFLG+E ++ +V Q+KYA DIL+ M
Sbjct: 2 TGNDATKIVQLKEHLFSHFQTKDLGYLKYFLGIEVAQSGVDVVISQRKYALDILEETGMQ 181
Query: 127 NCNPAST 133
NC P +
Sbjct: 182 NCRPVDS 202
>TC234303 weakly similar to UP|Q8W153 (Q8W153) Polyprotein, partial (10%)
Length = 558
Score = 52.0 bits (123), Expect = 5e-07
Identities = 32/99 (32%), Positives = 56/99 (56%)
Frame = +1
Query: 1 NEDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLY 60
+ +M+ +L ++ YGLKQS RAW + + +A+++V+ +S +++ +Y
Sbjct: 271 SSNMVCQLCRSFYGLKQSPRAWPFLYCGAAIWYD--SHEADHSVFYCHSPQG-CIYLIVY 441
Query: 61 VDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGM 99
VDD+ +T S+ I K + +F DLGKL YFLG+
Sbjct: 442 VDDIGITGSDQHGIT*LK*XLCCQFQTKDLGKLRYFLGI 558
>BM188122 weakly similar to PIR|I46200|I4620 retrovirus-related reverse
transcriptase homolog - pine (Pinus coulteri)
retrotransposon copia-like, partial (53%)
Length = 420
Score = 44.3 bits (103), Expect(2) = 3e-06
Identities = 24/61 (39%), Positives = 39/61 (63%)
Frame = +1
Query: 40 AEYNVYVKNSKDSNLVHICLYVDDLIVTDSNLAEIEAFKGHMMKKFNMTDLGKLTYFLGM 99
+E +YVK + NL+ + +++DD+ VT +N + FK M+ F MT+LG ++Y LGM
Sbjct: 148 SEAILYVKE-ECGNLLVVSVHLDDMFVTVTNEILVGEFKAEML*LFEMTNLGFVSYILGM 324
Query: 100 E 100
E
Sbjct: 325 E 327
Score = 24.6 bits (52), Expect(2) = 3e-06
Identities = 13/35 (37%), Positives = 18/35 (51%)
Frame = +2
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFV 36
E+ ++ L KAL GL Q+ AW +L FV
Sbjct: 32 EEKVY*LKKALNGLNQAPTAWYNMFDEYLQGLGFV 136
>NP005470 reverse transcriptase
Length = 267
Score = 46.2 bits (108), Expect = 2e-05
Identities = 22/63 (34%), Positives = 36/63 (56%)
Frame = +1
Query: 2 EDMIFRLHKALYGLKQSLRAWNKRIVNFLVQQEFVKCKAEYNVYVKNSKDSNLVHICLYV 61
E + +L ++LYGLKQS R W +F+ Q F + + VY +D ++++ LYV
Sbjct: 79 ERYVSQLQRSLYGLKQSPRQWYMSFDSFITNQGFKRSLYDCCVYHNKVEDGLMIYLLLYV 258
Query: 62 DDL 64
DD+
Sbjct: 259 DDM 267
>TC223792 weakly similar to UP|Q9FH39 (Q9FH39) Copia-type polyprotein,
partial (7%)
Length = 804
Score = 42.4 bits (98), Expect = 4e-04
Identities = 43/117 (36%), Positives = 54/117 (45%)
Frame = +2
Query: 231 LPTQMQIGVEICKIGKALHATCSNF*MLLYHGVQKSNLY*YFQLVNLSTL*GV*QHVKQY 290
L TQ IG I KAL CS MLL+ ++ + * + LV LS VKQ+
Sbjct: 227 LATQTLIGKGIQNRRKALEDICSCIMMLLWLRALRNRM**PYPLVKLSMWQHHLVPVKQF 406
Query: 291 G*IMC*RN*ILKSASLLHCL*TINLP*T*PKIMCCMEEASILKQSFIFSENK*TKRL 347
G* + RN* +L C T +L P CM EA+ L FI SE K K +
Sbjct: 407 G**IFLRN*S*GKGNLSTC*LTTSLLLIWPNTPPCMVEANTLSLGFIISEIKFQKEM 577
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.366 0.163 0.562
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,029,434
Number of Sequences: 63676
Number of extensions: 184394
Number of successful extensions: 2127
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 1592
Number of HSP's successfully gapped in prelim test: 51
Number of HSP's that attempted gapping in prelim test: 517
Number of HSP's gapped (non-prelim): 1660
length of query: 385
length of database: 12,639,632
effective HSP length: 99
effective length of query: 286
effective length of database: 6,335,708
effective search space: 1812012488
effective search space used: 1812012488
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC135162.11


