

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC135160.3 - phase: 0
(239 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
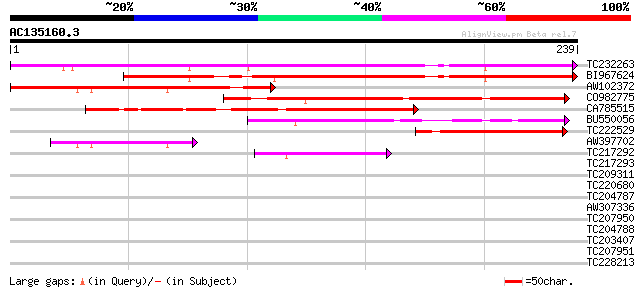
TC232263 174 3e-44
BI967624 159 1e-39
AW102372 109 1e-24
CO982775 101 3e-22
CA785515 99 2e-21
BU550056 76 1e-14
TC222529 homologue to UP|Q8I3F3 (Q8I3F3) Early transcribed membr... 70 9e-13
AW397702 57 1e-08
TC217292 weakly similar to UP|Q84WK9 (Q84WK9) At4g33985, partial... 42 2e-04
TC217293 39 0.002
TC209311 similar to GB|AAT41814.1|48310405|BT014831 At3g62070 {A... 34 0.052
TC220680 similar to UP|Q9SLT9 (Q9SLT9) ZCF37 protein (T30E16.15)... 32 0.34
TC204787 similar to UP|Q84WK9 (Q84WK9) At4g33985, partial (61%) 31 0.57
AW307336 weakly similar to GP|22202765|dbj P0039H02.8 {Oryza sat... 31 0.57
TC207950 similar to GB|AAT06459.1|46931310|BT012640 At3g57910 {A... 30 1.3
TC204788 similar to UP|Q84WK9 (Q84WK9) At4g33985, partial (61%) 30 1.3
TC203407 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-pr... 29 1.7
TC207951 similar to GB|AAT06459.1|46931310|BT012640 At3g57910 {A... 29 2.2
TC228213 homologue to UP|Q6KA74 (Q6KA74) Ankyrin repeat protein-... 28 2.8
>TC232263
Length = 897
Score = 174 bits (441), Expect = 3e-44
Identities = 111/254 (43%), Positives = 150/254 (58%), Gaps = 15/254 (5%)
Frame = +2
Query: 1 MSAEQILTLLDSFWFETTILT-NKNP--------SKVEEILPQDTNLLLVPTPNLDVRFY 51
M AE +L L DS WFE IL N +P ++ +L ++ L R
Sbjct: 17 MDAEAVLNLFDSCWFEVNILRRNSSPHTSITSYTENLDHVLKEEKEPKLPLIRTSHTRSM 196
Query: 52 SEQNLSIIGSVFSDSPSPNSVLT-SSKLRTIPSEREFGEFSTGTSSENH---HQKEDVAY 107
S+Q S DS SP+SVL KL+TI S +E + + H +
Sbjct: 197 SDQLSMTTTSFNQDSLSPDSVLLLHPKLQTILSGKEATDSEPDENPRTQVRPHPEVLCPK 376
Query: 108 TKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGREN 167
KK S S R+RR+S+SLS+LEF+ELKGFMDLGFVFSEEDKDS L S+IPGLQRL ++
Sbjct: 377 KNKKISSSRTRKRRESKSLSDLEFEELKGFMDLGFVFSEEDKDSSLASIIPGLQRLRKKE 556
Query: 168 DDAEEGEDEEHKKIDENVLSDNKPYLSEAWDV--FDQRERKNPLVNWRVPDKGSEIDMKD 225
++ EE ED DE +S +PYLSEAW+V +D+R+++N LVNW++P +E DMK+
Sbjct: 557 EEEEENED-----CDE--ISVPRPYLSEAWEVQEYDRRKKENSLVNWKMPAINNETDMKE 715
Query: 226 NLKFWAHAVASIAR 239
+L++WAH VAS R
Sbjct: 716 SLRWWAHTVASTVR 757
>BI967624
Length = 693
Score = 159 bits (401), Expect = 1e-39
Identities = 99/196 (50%), Positives = 132/196 (66%), Gaps = 5/196 (2%)
Frame = -2
Query: 49 RFYSEQNLSIIGSVFSDSPSPNSVLT-SSKLRTIPSEREFGEFSTGTSSENHHQKEDVAY 107
R S+Q I S DS SP+SVL KL+ I S +E T SE+ ++V
Sbjct: 665 RSMSDQLSMITTSFNHDSLSPDSVLLLHPKLQIILSGKE------ATDSES----DEVLC 516
Query: 108 TKK--KQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQRLGR 165
KK K S + R+RR+S+SLS+LEF+ELKGFMDLGFVFSEEDKDS L S+IPGLQRLG+
Sbjct: 515 PKKNKKISSNGTRKRRESKSLSDLEFEELKGFMDLGFVFSEEDKDSSLASIIPGLQRLGK 336
Query: 166 ENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDV--FDQRERKNPLVNWRVPDKGSEIDM 223
+ ++ EE +D DE +S +PYLSEAW+V D+R+++NPLVNW++P +E DM
Sbjct: 335 KEEEKEEDDD-----CDE--ISVPRPYLSEAWEVQECDRRKKENPLVNWKMPAINNETDM 177
Query: 224 KDNLKFWAHAVASIAR 239
K++L++WAH VAS R
Sbjct: 176 KESLRWWAHTVASTVR 129
>AW102372
Length = 410
Score = 109 bits (272), Expect = 1e-24
Identities = 65/123 (52%), Positives = 81/123 (65%), Gaps = 11/123 (8%)
Frame = +1
Query: 1 MSAEQILTLLDSFWFETTILTNKNPSK---------VEEILP-QDTNLLLVPTPNLDVRF 50
M+AE +L LLDSFWFE +IL++K P VEE+LP D LLLVPTP L+VR
Sbjct: 40 MAAEHVLRLLDSFWFEASILSSKIPPTNPISHNHKVVEEVLPLDDAKLLLVPTPTLEVRS 219
Query: 51 YSEQNLSIIGSVFSD-SPSPNSVLTSSKLRTIPSEREFGEFSTGTSSENHHQKEDVAYTK 109
YS+QNL V D SPSPNSVLT +LRTIPSERE EFS+G +H KE ++ +
Sbjct: 220 YSDQNLDSTTCVLCDYSPSPNSVLTPQRLRTIPSEREIREFSSG-----NHGKEGMSNKR 384
Query: 110 KKQ 112
K++
Sbjct: 385 KQK 393
>CO982775
Length = 613
Score = 101 bits (252), Expect = 3e-22
Identities = 60/149 (40%), Positives = 90/149 (60%), Gaps = 3/149 (2%)
Frame = -1
Query: 91 STGTSSENHHQKEDVAYTKKKQSQSYRRRRRKS---RSLSELEFKELKGFMDLGFVFSEE 147
S+ S++N H+K +K + + R+ +KS R+L ELE E+KGFMDLGF F +E
Sbjct: 565 SSSPSTQNRHRK------LRKPATTCARKLQKSISCRTLGELELDEVKGFMDLGFTFKKE 404
Query: 148 DKDSKLVSLIPGLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKN 207
+++S+IPGLQRLG DA E + H + +E +PYLSEAW + + +
Sbjct: 403 CLSPRMMSVIPGLQRLGVM--DATETVEGNHIEAEEQKRGIMRPYLSEAWPI---KRPDS 239
Query: 208 PLVNWRVPDKGSEIDMKDNLKFWAHAVAS 236
PL+N ++P + S +MK +L+FWA VAS
Sbjct: 238 PLLNLKIPKRCSSANMKKHLRFWAKTVAS 152
>CA785515
Length = 409
Score = 99.0 bits (245), Expect = 2e-21
Identities = 65/140 (46%), Positives = 90/140 (63%)
Frame = -1
Query: 33 PQDTNLLLVPTPNLDVRFYSEQNLSIIGSVFSDSPSPNSVLTSSKLRTIPSEREFGEFST 92
P + LL + + + R S+Q LS + DS SP+SV S KL+TI S ++
Sbjct: 385 PSEPTLLRIQSGHS--RSMSDQ-LSSMTCFKDDSLSPDSVF-SPKLQTILSGKDV----- 233
Query: 93 GTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSK 152
T +E H + + KK+ RR++R S+SLS+LEF+ELKGFMDLGFVFSEEDKDS
Sbjct: 232 -TDAEAAHVQLQLLVLPKKRE---RRKKRPSKSLSDLEFEELKGFMDLGFVFSEEDKDSS 65
Query: 153 LVSLIPGLQRLGRENDDAEE 172
L S+IPGLQRLG+ +++ E+
Sbjct: 64 LASIIPGLQRLGKSDEEEED 5
>BU550056
Length = 614
Score = 76.3 bits (186), Expect = 1e-14
Identities = 53/140 (37%), Positives = 73/140 (51%), Gaps = 4/140 (2%)
Frame = -1
Query: 101 QKEDVAYTKKKQSQSYRRR----RRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSL 156
+K +V K+ RRR + RSLS+LEF+E++GF DLGF F +E L S+
Sbjct: 590 RKTEVESFNKEGIMEMRRRYLNQKTMRRSLSDLEFEEVQGFKDLGFSFEKETLSPSLASI 411
Query: 157 IPGLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPD 216
+PGLQ ++ D+ EE + + +PYLSEAW V + + NW
Sbjct: 410 LPGLQE--KKRDETEEDK------------AARRPYLSEAWLV---QSCAPAIPNW--TS 288
Query: 217 KGSEIDMKDNLKFWAHAVAS 236
S DMK +KFWA AVAS
Sbjct: 287 HKSSGDMKVQIKFWARAVAS 228
>TC222529 homologue to UP|Q8I3F3 (Q8I3F3) Early transcribed membrane protein,
partial (6%)
Length = 196
Score = 70.1 bits (170), Expect = 9e-13
Identities = 30/64 (46%), Positives = 47/64 (72%)
Frame = +1
Query: 172 EGEDEEHKKIDENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKGSEIDMKDNLKFWA 231
EG++EE D S +PYLSEAW + ++R+++NPL+NW++P +EID+KD+L++WA
Sbjct: 13 EGDEEEE---DSEGSSVQRPYLSEAWKIQERRKKENPLMNWKIPALNNEIDIKDSLRWWA 183
Query: 232 HAVA 235
VA
Sbjct: 184 QTVA 195
>AW397702
Length = 221
Score = 56.6 bits (135), Expect = 1e-08
Identities = 37/73 (50%), Positives = 44/73 (59%), Gaps = 11/73 (15%)
Frame = -2
Query: 18 TILTNKNPSK---------VEEILP-QDTNLLLVPTPNLDVRFYSEQNLSIIGSVFSD-S 66
+IL+NK S VEE+LP D LLLVPTP L+VR YS+QNL + D S
Sbjct: 220 SILSNKTLSNSISHDHNKVVEEVLPLDDAKLLLVPTPTLEVRSYSDQNLDSTTCLLCDYS 41
Query: 67 PSPNSVLTSSKLR 79
SPNSVLT +LR
Sbjct: 40 QSPNSVLTPQRLR 2
>TC217292 weakly similar to UP|Q84WK9 (Q84WK9) At4g33985, partial (50%)
Length = 831
Score = 42.0 bits (97), Expect = 2e-04
Identities = 24/60 (40%), Positives = 35/60 (58%), Gaps = 2/60 (3%)
Frame = +1
Query: 104 DVAYTKKKQSQS--YRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLIPGLQ 161
D A+ ++K + RR+ R S+SLSE + ELK +LGF F + D KL + IP L+
Sbjct: 145 DEAWQRRKDNNHPVNRRQHRLSKSLSEDDLDELKACFELGFGFDSPEIDPKLSNTIPALE 324
>TC217293
Length = 751
Score = 38.9 bits (89), Expect = 0.002
Identities = 23/64 (35%), Positives = 35/64 (53%), Gaps = 6/64 (9%)
Frame = +2
Query: 104 DVAYTKKKQSQ------SYRRRRRKSRSLSELEFKELKGFMDLGFVFSEEDKDSKLVSLI 157
D A+ ++K + ++R R S+SLSE + ELK +LGF F + D KL + I
Sbjct: 5 DEAWQRRKDNSHHISGDNHRCSHRLSKSLSEDDLDELKACFELGFGFDSPEIDPKLSNTI 184
Query: 158 PGLQ 161
P L+
Sbjct: 185 PALE 196
>TC209311 similar to GB|AAT41814.1|48310405|BT014831 At3g62070 {Arabidopsis
thaliana;} , partial (30%)
Length = 814
Score = 34.3 bits (77), Expect = 0.052
Identities = 44/163 (26%), Positives = 68/163 (40%), Gaps = 10/163 (6%)
Frame = +1
Query: 83 SEREFGEFSTGTSSENHHQKEDVAYTKKKQSQ----SYRRRRRKSRSLSELEFKELKGFM 138
SE E G + S+ +K+ Y ++ S RRRR+S F++ K
Sbjct: 178 SENELGSCYSLPSTPPRMRKQLKDYASDNGNEITNKSDPRRRRRSFRRERWGFEKTKIRK 357
Query: 139 DLGFVFSEEDKDSKLVSLIP----GLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLS 194
+ G D D V LI G++ L + ++ + D + E +L + LS
Sbjct: 358 EEGL---SGDSDQSGVMLITRPKGGIRSLCMDLEEVKACRDLGFELEHERML-EIPSRLS 525
Query: 195 EAWDVFDQRERKN-PLVNWRVPDKGSEI-DMKDNLKFWAHAVA 235
+ D N P+ NWR+ G + D+K LK WA AVA
Sbjct: 526 FSNSTLDTSSGSNSPIANWRISAPGDDPRDVKARLKVWAQAVA 654
>TC220680 similar to UP|Q9SLT9 (Q9SLT9) ZCF37 protein (T30E16.15), partial
(33%)
Length = 1227
Score = 31.6 bits (70), Expect = 0.34
Identities = 34/150 (22%), Positives = 65/150 (42%), Gaps = 19/150 (12%)
Frame = +3
Query: 99 HHQKEDV------AYTKKKQSQSYRRRRRKSRS-------LSELEFKELKGF-----MDL 140
HHQ+EDV Y+++K+S++ + +R L++L+ + K + D+
Sbjct: 312 HHQEEDVPCSSPKKYSRRKESRNINKNPYSNRGLDKFSALLADLDERRQKVYSQMSPQDI 491
Query: 141 GFV-FSEEDKDSKLVSLIPGLQRLGRENDDAEEGEDEEHKKIDENVLSDNKPYLSEAWDV 199
FV F+ D +P + ++ + N D ++ + EE K S+ SE +
Sbjct: 492 SFVRFAYSSND----DFVPIVVKVKKNNKDQKKHKSEELKARHITSFSEQLEKSSEETTL 659
Query: 200 FDQRERKNPLVNWRVPDKGSEIDMKDNLKF 229
+++++ N L E K NL F
Sbjct: 660 EERKQKLNKL----------EAHKKKNLSF 719
>TC204787 similar to UP|Q84WK9 (Q84WK9) At4g33985, partial (61%)
Length = 977
Score = 30.8 bits (68), Expect = 0.57
Identities = 29/125 (23%), Positives = 52/125 (41%), Gaps = 4/125 (3%)
Frame = +2
Query: 119 RRRKSRSLSELEFKELKGFMDLGFVFS---EEDKDSKLVSLIPGLQRLGRENDDAEEGED 175
+ R+S+S+++ + ELK ++LGF F E + D +L +P L N
Sbjct: 272 KNRRSKSVTDEDVDELKACIELGFGFDSSPENELDQRLSDTLPALGLYYAVN-------- 427
Query: 176 EEHKKIDENVLSDNKPYLSEAWDVFDQRERKNPLVNWRVPDKGSEID-MKDNLKFWAHAV 234
K+ + ++++ P S A D + + + G +K L+ WA V
Sbjct: 428 ---KRYNNSLVTKTTPSSSSAASDCDSSPSPHGSPHSAIFTTGDNPQTVKTRLRQWAQVV 598
Query: 235 ASIAR 239
A R
Sbjct: 599 ACAVR 613
>AW307336 weakly similar to GP|22202765|dbj P0039H02.8 {Oryza sativa
(japonica cultivar-group)}, partial (18%)
Length = 371
Score = 30.8 bits (68), Expect = 0.57
Identities = 16/59 (27%), Positives = 35/59 (59%), Gaps = 4/59 (6%)
Frame = +1
Query: 109 KKKQSQSYRRRRRKSRSLSELEFKELKGFMDLGFVFSEE----DKDSKLVSLIPGLQRL 163
KKK+ + +++++K + + E K+ K +++GF ++ + K K+ + IP LQ+L
Sbjct: 76 KKKKKKKKKKKKKKKKKKKKKEKKKKKKNLEIGFFYNCKKG*TTKKGKMGNKIPQLQKL 252
>TC207950 similar to GB|AAT06459.1|46931310|BT012640 At3g57910 {Arabidopsis
thaliana;} , partial (72%)
Length = 1259
Score = 29.6 bits (65), Expect = 1.3
Identities = 27/111 (24%), Positives = 45/111 (40%), Gaps = 1/111 (0%)
Frame = +1
Query: 79 RTIPSEREFGEFSTGTSSENHHQKEDVAYTKKKQSQSYRRRRRKSRSLSELEFKELKGFM 138
R P E G E+ H+++ +K + R+RRK L + K
Sbjct: 355 RAEPVGIEIRRSRAGVGLEDPHKEK-----RKNEGIMMDRKRRKEEDLMKEFGSRQKSQW 519
Query: 139 DLGFVFSEEDKDSKLVSLIPGLQRLG-RENDDAEEGEDEEHKKIDENVLSD 188
V +K + + + + ++NDD EGE+EE ++I E L D
Sbjct: 520 QSRRVIINYNKAKAALDQLENREIVEPQKNDDVAEGEEEEEEEITEEDLHD 672
>TC204788 similar to UP|Q84WK9 (Q84WK9) At4g33985, partial (61%)
Length = 1517
Score = 29.6 bits (65), Expect = 1.3
Identities = 15/45 (33%), Positives = 27/45 (59%), Gaps = 3/45 (6%)
Frame = +3
Query: 119 RRRKSRSLSELEFKELKGFMDLGFVFS---EEDKDSKLVSLIPGL 160
+ R+S+S+++ + ELK ++LGF F E + D +L +P L
Sbjct: 270 KNRRSKSVTDEDVDELKACIELGFGFDSSPEVELDQRLSDTLPAL 404
>TC203407 weakly similar to UP|Q9ZT16 (Q9ZT16) Arabinogalactan-protein
(AtAGP4) (AT5g10430/F12B17_220), partial (50%)
Length = 966
Score = 29.3 bits (64), Expect = 1.7
Identities = 14/32 (43%), Positives = 19/32 (58%)
Frame = -3
Query: 95 SSENHHQKEDVAYTKKKQSQSYRRRRRKSRSL 126
S+E HQ+ KKK ++ RRRRRK + L
Sbjct: 829 SNEQTHQQRTKTRRKKKTHKTRRRRRRKKKPL 734
>TC207951 similar to GB|AAT06459.1|46931310|BT012640 At3g57910 {Arabidopsis
thaliana;} , partial (23%)
Length = 707
Score = 28.9 bits (63), Expect = 2.2
Identities = 12/24 (50%), Positives = 17/24 (70%)
Frame = +3
Query: 165 RENDDAEEGEDEEHKKIDENVLSD 188
++NDD EGE+EE ++I E L D
Sbjct: 57 QKNDDVAEGEEEEEEEITEEDLHD 128
>TC228213 homologue to UP|Q6KA74 (Q6KA74) Ankyrin repeat protein-like,
partial (12%)
Length = 530
Score = 28.5 bits (62), Expect = 2.8
Identities = 15/35 (42%), Positives = 24/35 (67%), Gaps = 3/35 (8%)
Frame = +3
Query: 102 KEDVAYTKKKQSQSYRRRRRK---SRSLSELEFKE 133
KED A TKK + ++ RR+++K + S +E EFK+
Sbjct: 252 KEDPAETKKGKDKNIRRKKKKGGINESKNESEFKK 356
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.130 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,937,398
Number of Sequences: 63676
Number of extensions: 130051
Number of successful extensions: 747
Number of sequences better than 10.0: 63
Number of HSP's better than 10.0 without gapping: 719
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 732
length of query: 239
length of database: 12,639,632
effective HSP length: 94
effective length of query: 145
effective length of database: 6,654,088
effective search space: 964842760
effective search space used: 964842760
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 57 (26.6 bits)
Medicago: description of AC135160.3


