

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
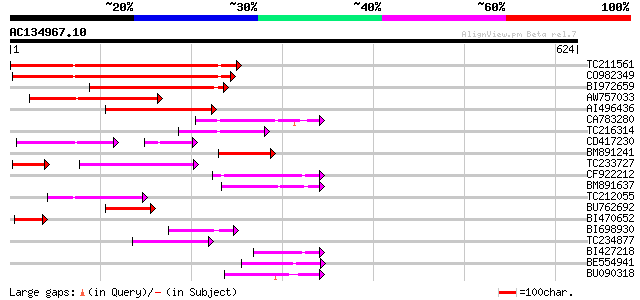
Query= AC134967.10 + phase: 0 /pseudo
(624 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 245 5e-65
CO982349 201 1e-51
BI972659 132 4e-31
AW757033 124 1e-28
AI496436 109 3e-24
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 91 2e-18
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 89 8e-18
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 70 2e-17
BM891241 80 4e-15
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 53 9e-15
CF922212 77 3e-14
BM891637 69 6e-12
TC212055 69 8e-12
BU762692 59 8e-09
BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase ... 55 1e-07
BI698930 53 5e-07
TC234877 50 3e-06
BI427218 49 9e-06
BE554941 48 1e-05
BU090318 weakly similar to GP|9743345|gb| F21J9.30 {Arabidopsis ... 44 2e-04
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 245 bits (625), Expect = 5e-65
Identities = 121/255 (47%), Positives = 166/255 (64%)
Frame = +1
Query: 1 WRKWIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEA 60
W KWI+ECV +A+ SVLVNGSPT EF +RGLRQGDPL+PFLF + AEGLN +M+ VE
Sbjct: 157 WIKWIEECVKSASISVLVNGSPTAEFLPQRGLRQGDPLTPFLFNIVAEGLNGLMRRAVEE 336
Query: 61 QLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSM 120
L+ Y +GA N + +S LQ+ADDT+ G + NV A++ L FE +S L++NF KS
Sbjct: 337 NLYKPYLVGA-NGVPISILQYADDTIFFGEAAKENVEAIKVILRSFELVSNLRINFAKSC 513
Query: 121 LVGVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWNS 180
+ W EAA+ L C + PF+YL +PIG N RR W+ I+ + + +LS W
Sbjct: 514 FGVFGVTDQWKQEAANYLHCSLLAFPFIYLGIPIGANSRRCHMWDSIIRKCERKLSKWK* 693
Query: 181 RFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISW 240
R +SFGGR+ L+KSVLTS+P+Y SFF+ P ++ + + F GGG D +KISW
Sbjct: 694 RHISFGGRVTLIKSVLTSIPIYFFSFFRVPQSVVDKLVKI*RTFLWGGG---PDQKKISW 864
Query: 241 LSWQTVCLKKECGGL 255
+ W+ +CL KE GG+
Sbjct: 865 IRWEKLCLPKERGGI 909
>CO982349
Length = 795
Score = 201 bits (510), Expect = 1e-51
Identities = 99/245 (40%), Positives = 154/245 (62%)
Frame = +3
Query: 4 WIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLF 63
WI+ C+ +A+ S+LVN SP +EF +RGLRQGDPL+P LF + AEGL +M+ V+ + F
Sbjct: 69 WIRGCLHSASVSILVNESPINEFSPQRGLRQGDPLAPLLFNIVAEGLTGLMREAVDRKRF 248
Query: 64 SGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLVG 123
+ + +G N VS LQ+ADDT+ + N++ ++ L FE SGLK+NF +S
Sbjct: 249 NSFLVGK-NKEPVSILQYADDTIFFEEATMENMKVIKTILRCFELASGLKINFAESRFGA 425
Query: 124 VNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWNSRFL 183
+ W EAA L+C + +PF YL +PI NPR +PI+ + +++L+ W R +
Sbjct: 426 IWKPDHWCKEAAEFLNCSMLSMPFSYLGIPIEANPRCREI*DPIIRKCETKLARWKQRHI 605
Query: 184 SFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISWLSW 243
S GGR+ L+ ++LT+LP+Y SFF+AP+ ++ + ++ F GGG R+I+W+ W
Sbjct: 606 SLGGRVTLINAILTALPIYFFSFFRAPSKVVQKLVNIQRKFLWGGG---SXQRRIAWVKW 776
Query: 244 QTVCL 248
+TVCL
Sbjct: 777 ETVCL 791
>BI972659
Length = 453
Score = 132 bits (333), Expect = 4e-31
Identities = 64/154 (41%), Positives = 97/154 (62%)
Frame = +1
Query: 88 LGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLVGVNIGSSWLYEAASVLSCKVGIVPF 147
+G SW N+ L++ L FE +SGL++N+ KS V +W ++AA +L+C+ +PF
Sbjct: 1 VGEASWDNIIVLKSMLRGFEMVSGLRINYAKSQFGIVGFQPNWAHDAAQLLNCRQLDIPF 180
Query: 148 LYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWNSRFLSFGGRLVLLKSVLTSLPVYALSFF 207
YL MPI WEP++N+ K++LS WN ++LS GGR+ L+KSVL +LP+Y LSFF
Sbjct: 181 HYLGMPIAVKASSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFF 360
Query: 208 KAPTGIISSIDSLFINFFLGGGGGSEDNRKISWL 241
K P I+ + SL F GG++ + +ISW+
Sbjct: 361 KIPQRIVDKLVSLQRTFM---WGGNQHHNRISWV 453
>AW757033
Length = 441
Score = 124 bits (311), Expect = 1e-28
Identities = 65/146 (44%), Positives = 89/146 (60%)
Frame = -2
Query: 23 TDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLFSGYSIGAANPIVVSHLQFA 82
T EF RGLRQGDPL+P LF +AAEGL +M+ V + +++G VS LQ+A
Sbjct: 440 TREFSPHRGLRQGDPLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKK-PVSILQYA 264
Query: 83 DDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLVGVNIGSSWLYEAASVLSCKV 142
DDT+ G + NVR ++ L FE SGLK+NF KS +++ W EAA ++C +
Sbjct: 263 DDTIFFGEATMENVRVIKTILRGFELASGLKINFAKSRFGAISVPD*WRKEAAEFMNCSL 84
Query: 143 GIVPFLYLDMPIGGNPRRLCFWEPIV 168
+PF YL +PIG NPRR W+PI+
Sbjct: 83 LSLPFSYLGIPIGANPRRRETWDPII 6
>AI496436
Length = 414
Score = 109 bits (273), Expect = 3e-24
Identities = 52/122 (42%), Positives = 74/122 (60%)
Frame = +1
Query: 106 FEAMSGLKVNFHKSMLVGVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRRLCFWE 165
FE SGLK+NF KS + W EAA L+ V +PF+YL +PIG N R WE
Sbjct: 16 FELSSGLKINFAKSQFGTICKSDIWRKEAADYLNSSVLSMPFVYLGIPIGANSRHSDVWE 195
Query: 166 PIVNRIKSRLSGWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFF 225
P+V + + +L+ W +++SFGGR+ L+ SVLT+LP+Y LSFF+ P ++ D F
Sbjct: 196 PVVRKCERKLASWKQKYVSFGGRVTLINSVLTALPIYLLSFFRIPDKVVDKADYNSA*IF 375
Query: 226 LG 227
+G
Sbjct: 376 MG 381
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 90.9 bits (224), Expect = 2e-18
Identities = 56/148 (37%), Positives = 81/148 (53%), Gaps = 6/148 (4%)
Frame = +2
Query: 205 SFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISWLSWQTVCLKKECGGLGVRKLKEFN 264
SFF P II+ ++SL F GG D+RKI+W++W+TVCL K GGLG++ L+ FN
Sbjct: 5 SFFSLP*KIIAKLESLQRRFLWGG---EADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFN 175
Query: 265 LALLGKWCWRMLVDRGGLWYRVLVARYGELGGR-LEVG--GRSVSCWWSEV---RRLRDG 318
LLGKW W + + W +VL ++YG G R LE G G S WW ++ ++L+
Sbjct: 176 TTLLGKWRWDLFYIQQEPWAKVLQSKYG--GWRALEEGSSGSKDSAWWKDLIKTQQLQRN 349
Query: 319 VGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
+ + + +VG G FW D
Sbjct: 350 IP------LKRETIWKVGGGDRIKFWED 415
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 88.6 bits (218), Expect = 8e-18
Identities = 41/101 (40%), Positives = 61/101 (59%)
Frame = +2
Query: 186 GGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSEDNRKISWLSWQT 245
GGR+ L+KSVL++LP+ LSFFK P I+ + +L F GG ++ + +I W+ W
Sbjct: 5 GGRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGG---TQHHNRIPWVKWAD 175
Query: 246 VCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRV 286
+C K GGLG++ L FN AL G+W W + + LW R+
Sbjct: 176 ICNPKIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWARL 298
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 70.5 bits (171), Expect(2) = 2e-17
Identities = 39/112 (34%), Positives = 60/112 (52%)
Frame = -3
Query: 8 CVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLFLLAAEGLNIVMKSLVEAQLFSGYS 67
C+ST T S+L NG D F G+RQ DP++P+LF+L E L+ ++ ++ +L
Sbjct: 668 CMSTTTMSLLWNGEVLDHFSPNIGVRQRDPIAPYLFVLCLERLSHLINLVMNKKL*RSIQ 489
Query: 68 IGAANPIVVSHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKS 119
+ +++SHL F DD +L + V + L LF SG KVN K+
Sbjct: 488 VNRGG-LLISHLAFVDDLILFVEANMDQVDVINLTLELFSDSSGAKVNVDKT 336
Score = 37.0 bits (84), Expect(2) = 2e-17
Identities = 23/59 (38%), Positives = 33/59 (54%), Gaps = 1/59 (1%)
Frame = -2
Query: 149 YLDMPI-GGNPRRLCFWEPIVNRIKSRLSGWNSRFLSFGGRLVLLKSVLTSLPVYALSF 206
YL +PI NP F + I++++ R S W S LS GRL L K V+ +LP + + F
Sbjct: 243 YLGVPIFHKNPTSHTF-DFIIDKVNKRFSRWKSNLLSMVGRLTLTK*VV*ALPSHIMQF 70
>BM891241
Length = 407
Score = 79.7 bits (195), Expect = 4e-15
Identities = 30/63 (47%), Positives = 45/63 (70%)
Frame = -2
Query: 230 GGSEDNRKISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVA 289
GG D++KI W+ W+ VCL K GGLG++ + +FN+ALLGKW W + D+ LW R++ +
Sbjct: 406 GGDHDHKKIPWVKWEVVCLPKTKGGLGIKDISKFNIALLGKWIWALASDQQQLWARIINS 227
Query: 290 RYG 292
+YG
Sbjct: 226 KYG 218
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 53.1 bits (126), Expect(2) = 9e-15
Identities = 37/132 (28%), Positives = 64/132 (48%), Gaps = 1/132 (0%)
Frame = -2
Query: 77 SHLQFADDTLLLGSKSWANVRALRAALVLFEAMSGLKVNFHKSMLVGVNIGSSWLYEAAS 136
SH+ ADD ++ + NV L L++ G ++ HKS + +I L +
Sbjct: 451 SHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIFIGSIPGQRLRVISD 272
Query: 137 VLSCKVGIVPFLYLDMPI-GGNPRRLCFWEPIVNRIKSRLSGWNSRFLSFGGRLVLLKSV 195
+L +PF Y +PI G P R + + ++IK +L+ W LS GR+ L+ SV
Sbjct: 271 LLQYFHSDIPFNYFGVPIFKGKPTRRHLY-CVADKIKCKLASWKGSILSIMGRIQLVNSV 95
Query: 196 LTSLPVYALSFF 207
+ + +Y+ S +
Sbjct: 94 IHGMLLYSFSVY 59
Score = 45.4 bits (106), Expect(2) = 9e-15
Identities = 20/40 (50%), Positives = 27/40 (67%)
Frame = -3
Query: 4 WIKECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPFLF 43
WI+ + +A S+ VNG P F +RG+RQGDPLSP L+
Sbjct: 663 WIRVILLSAKLSISVNGEPVGFFSCQRGVRQGDPLSPSLY 544
>CF922212
Length = 445
Score = 76.6 bits (187), Expect = 3e-14
Identities = 48/125 (38%), Positives = 66/125 (52%), Gaps = 2/125 (1%)
Frame = -2
Query: 224 FFLGGGGGSEDNRKISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLW 283
+FL GGG + +KI+W++W +VCL KE GLG R L++FN ALLGK W + +G L
Sbjct: 399 WFLWGGGLGQ--KKIAWVNWDSVCLPKEKEGLGNRDLRKFNYALLGKRRWNLFHHQGELG 226
Query: 284 YRVLVARYGELGGRLEVGG-RSVSCWWSEVRRLRDGVGDGGSGWFGDQV-LRRVGDGVVT 341
RVL ++Y E +S S WW E+ + DG WF + R+G
Sbjct: 225 ARVLDSKYKRWRNLDEERRVKSESFWWQEISFITHSTEDG--SWFEKRTKTGRLGCRAKV 52
Query: 342 SFWSD 346
FW D
Sbjct: 51 RFWED 37
>BM891637
Length = 427
Score = 68.9 bits (167), Expect = 6e-12
Identities = 38/115 (33%), Positives = 60/115 (52%), Gaps = 2/115 (1%)
Frame = -1
Query: 234 DNRKISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGE 293
+ +KI+W+SW+ C + GGLG++ +K N ALL KW W M LW R+L+++Y
Sbjct: 424 EGKKIAWISWRQCCASGDVGGLGIKDIKILNNALLIKWKWLMFHQPHQLWNRILISKYKG 245
Query: 294 LGGRLEVGGRS--VSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
G L+ G + S WW+++R + + +Q +VG G FW D
Sbjct: 244 WRG-LDQGPQKYYFSPWWADLRAINQHQSMIAA---SNQFCWKVGRGDQILFWED 92
>TC212055
Length = 776
Score = 68.6 bits (166), Expect = 8e-12
Identities = 41/111 (36%), Positives = 62/111 (54%), Gaps = 1/111 (0%)
Frame = +1
Query: 42 LFLLAAEGLNIVMKSLVEAQLFSGYSIGAANPIVVSHLQFADDTLLLGSKSWANVRALRA 101
LF + AEGL +M+ ++ L+S +G N I V+ LQ+ADDT+ LG + NV +++
Sbjct: 1 LFNIVAEGLTGLMREALDKSLYSSLMVGK-NKIPVNILQYADDTIFLGEATMQNVMTIKS 177
Query: 102 ALVLFEAMSGLKVNFHKSMLVGVNIGSSWLYEAA-SVLSCKVGIVPFLYLD 151
L +FE SGLK++F KS + W EA + +C+ IV LD
Sbjct: 178 MLRVFELASGLKIHFAKSSFGAIGKPDHWQKEAGLNTFNCRNLIVHIRQLD 330
>BU762692
Length = 423
Score = 58.5 bits (140), Expect = 8e-09
Identities = 27/55 (49%), Positives = 35/55 (63%)
Frame = -3
Query: 106 FEAMSGLKVNFHKSMLVGVNIGSSWLYEAASVLSCKVGIVPFLYLDMPIGGNPRR 160
FE +SGLK+NF KS + W EAAS L+C + +PF+Y +PIG NPRR
Sbjct: 169 FELVSGLKINFAKSCFGAFGMTDRWKTEAASCLNCSLLAIPFVYQGIPIGANPRR 5
>BI470652 similar to PIR|T17102|T171 RNA-directed DNA polymerase (EC
2.7.7.49) (clone AmLi2) - garden snapdragon (fragment),
partial (22%)
Length = 111
Score = 54.7 bits (130), Expect = 1e-07
Identities = 24/36 (66%), Positives = 29/36 (79%)
Frame = +1
Query: 6 KECVSTATASVLVNGSPTDEFPLKRGLRQGDPLSPF 41
KEC+S+A+ S+LVNGSPT+EF RGLRQG P PF
Sbjct: 1 KECLSSASISILVNGSPTEEFKPXRGLRQGGPFGPF 108
>BI698930
Length = 384
Score = 52.8 bits (125), Expect = 5e-07
Identities = 28/78 (35%), Positives = 43/78 (54%)
Frame = +3
Query: 175 LSGWNSRFLSFGGRLVLLKSVLTSLPVYALSFFKAPTGIISSIDSLFINFFLGGGGGSED 234
L+ W LSF GR L+ S+L+SLP+++ SFFK P + S+ NF G +
Sbjct: 81 LALWKHEVLSFAGRTCLINSMLSSLPLFSFSFFKVPLNVGMRFISI*RNFV----GEWRE 248
Query: 235 NRKISWLSWQTVCLKKEC 252
+ I W++W V +K+ C
Sbjct: 249 S*MIPWVNWDKV*VKRVC 302
>TC234877
Length = 534
Score = 50.1 bits (118), Expect = 3e-06
Identities = 25/89 (28%), Positives = 45/89 (50%)
Frame = +1
Query: 136 SVLSCKVGIVPFLYLDMPIGGNPRRLCFWEPIVNRIKSRLSGWNSRFLSFGGRLVLLKSV 195
++L VG +PF+YL +P+ R PI +RIK+ L W LSF G + L+ S+
Sbjct: 10 NILGFSVGSLPFIYLSVPLFQEKPRPVHLRPIADRIKAELQSWKGNLLSFMGCVQLIISI 189
Query: 196 LTSLPVYALSFFKAPTGIISSIDSLFINF 224
+ + + + + P ++ + + NF
Sbjct: 190 IDGMLL*SFHLYSWPKALLIEMQTWTRNF 276
>BI427218
Length = 421
Score = 48.5 bits (114), Expect = 9e-06
Identities = 25/79 (31%), Positives = 36/79 (44%), Gaps = 1/79 (1%)
Frame = +1
Query: 269 GKWCWRMLVDRGGLWYRVLVARYGELGGRLE-VGGRSVSCWWSEVRRLRDGVGDGGSGWF 327
GKW W ++G LW R+L ++YG+ E G S WW ++ + DG F
Sbjct: 1 GKWKWSFFYNKGELWARILDSKYGDWRHLEEPTRGNRASAWWKDINIIDPSGADG--SLF 174
Query: 328 GDQVLRRVGDGVVTSFWSD 346
+ +V G T FW D
Sbjct: 175 DKGIKWKVACGEKTKFWED 231
>BE554941
Length = 383
Score = 47.8 bits (112), Expect = 1e-05
Identities = 29/92 (31%), Positives = 40/92 (42%)
Frame = +1
Query: 256 GVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARYGELGGRLEVGGRSVSCWWSEVRRL 315
G++ L FN++LL K LV+ +W +L YG L S WW ++ L
Sbjct: 7 GIKNLSIFNISLLVKRR*HFLVETKAIWAPILKFHYGSLDLSNTGSASRTSQWWRDI--L 180
Query: 316 RDGVGDGGSGWFGDQVLRRVGDGVVTSFWSDR 347
R G GW R++GD T FW R
Sbjct: 181 RIGNCQYQPGWLTSTFCRKIGDRKDTLFWRHR 276
>BU090318 weakly similar to GP|9743345|gb| F21J9.30 {Arabidopsis thaliana},
partial (1%)
Length = 422
Score = 44.3 bits (103), Expect = 2e-04
Identities = 30/113 (26%), Positives = 49/113 (42%), Gaps = 3/113 (2%)
Frame = +3
Query: 237 KISWLSWQTVCLKKECGGLGVRKLKEFNLALLGKWCWRMLVDRGGLWYRVLVARY---GE 293
K+ W SW+ +C K+ GGLG+R L N +LL K M L RV+ ++Y +
Sbjct: 3 KVHWTSWKALCSPKKSGGLGLRLLSHMNKSLLMKST*NMCTKPNDL*VRVVRSKYK*GKD 182
Query: 294 LGGRLEVGGRSVSCWWSEVRRLRDGVGDGGSGWFGDQVLRRVGDGVVTSFWSD 346
+ ++ + W G+ D + R++G+G FW D
Sbjct: 183 ILP*IDPKKTGTNFW--------QGIVHCWEKITHDNIARKIGNGRSVHFWKD 317
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.348 0.155 0.570
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,324,234
Number of Sequences: 63676
Number of extensions: 570831
Number of successful extensions: 6363
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 4711
Number of HSP's successfully gapped in prelim test: 180
Number of HSP's that attempted gapping in prelim test: 1467
Number of HSP's gapped (non-prelim): 5122
length of query: 624
length of database: 12,639,632
effective HSP length: 103
effective length of query: 521
effective length of database: 6,081,004
effective search space: 3168203084
effective search space used: 3168203084
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC134967.10


