

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134521.1 - phase: 0 /pseudo
(324 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
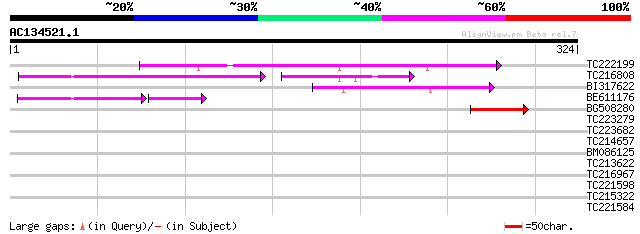
Score E
Sequences producing significant alignments: (bits) Value
TC222199 118 3e-27
TC216808 59 9e-20
BI317622 56 3e-08
BE611176 39 3e-05
BG508280 45 4e-05
TC223279 32 0.51
TC223682 homologue to UP|SYE_ARATH (Q9FEA2) Glutamyl-tRNA synthe... 31 0.67
TC214657 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxid... 29 2.5
BM086125 29 2.5
TC213622 29 3.3
TC216967 28 7.4
TC221598 homologue to GB|AAM19966.1|20466089|AY098956 At2g32600/... 28 7.4
TC215322 similar to UP|Q9LSL3 (Q9LSL3) Oligopeptidase A, partial... 27 9.6
TC221584 weakly similar to UP|Q9FH22 (Q9FH22) Gb|AAF17687.1, par... 27 9.6
>TC222199
Length = 691
Score = 118 bits (296), Expect = 3e-27
Identities = 79/229 (34%), Positives = 117/229 (50%), Gaps = 22/229 (9%)
Frame = +2
Query: 75 DFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIP---KIPTIAKALHLEVADIKDKFTTKDG 131
DFQL PT+EE+ +G P+ + P+ + G +P +IPT+ K + +K T++G
Sbjct: 5 DFQLVPTIEEFEEILGCPLGERKPYLSSGCLPSLSRIPTMVKDSARGLDRVKQ---TRNG 175
Query: 132 LPCLPYNFLHQKATTCFEKSKTDDFESILALLIYGIVLFPKVDKFVDMNAIRIFLT---- 187
+ LP +L KA + F +LALLI+G+VLFP VD VD+ AI FL
Sbjct: 176 IAGLPRKYLEDKARGIANQGDWVPFMDVLALLIFGVVLFPNVDGLVDLAAIDAFLAYHHS 355
Query: 188 -QNPVPVLLADTYVAIHDRTSHGKGTIVCCTPLLHRWITSHL-PRPSIRPEP-------- 237
++PV +LAD + + R I+CC P L W+ SHL + + RP P
Sbjct: 356 KESPVVAVLADLFDTFNRRYEKSSARIICCLPALCVWLVSHLFQQDTRRPCPLLSHRSCT 535
Query: 238 ----IPWSQKLMTWTPKDIIWFNPSCD-PEFIIDSCGDFNNVPLLGTQG 281
I W Q L + I WF + E ++ SCG + N+PL+GT+G
Sbjct: 536 EKRRIDWDQLLAGIGGRTISWFPRWKEGKEGVLFSCGRYPNIPLVGTRG 682
>TC216808
Length = 1144
Score = 58.9 bits (141), Expect(2) = 9e-20
Identities = 38/141 (26%), Positives = 68/141 (47%)
Frame = +2
Query: 6 RRTHGYKFKAPDTESLEGLRGLTKMLKNPGHFQERYGNLLDILKTKVDVMLLNTLVQFYD 65
+R + K K+ + SL L L L+ F++ YG + D+ + +V + + +L Q+YD
Sbjct: 146 KRFYQVKIKSLEVTSLGELGQLMGQLQR*A-FRKAYGKIWDLARVEVSMEPIASLSQYYD 322
Query: 66 PIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIAKALHLEVADIKDK 125
FTF DFQL PT++E G + P+ G P +AK + + ++
Sbjct: 323 QPLRCFTFEDFQLVPTVKELEEIRGCQLGGRKPYLFSGFYPSTTRVAKVVKISAQELNRV 502
Query: 126 FTTKDGLPCLPYNFLHQKATT 146
++G+ +P L ++A T
Sbjct: 503 KQNRNGVVGVPRKCLEERAKT 565
Score = 55.5 bits (132), Expect(2) = 9e-20
Identities = 35/83 (42%), Positives = 49/83 (58%), Gaps = 7/83 (8%)
Frame = +3
Query: 156 FESILALLIYGIVLFPKVDKFVDMNAIRIFLT-----QNPVPVLLA--DTYVAIHDRTSH 208
F ILALLI+G+VLFP ++ VD+ AI FL ++PV +LA DT+ +++S
Sbjct: 594 FIDILALLIFGVVLFPNMEGLVDLAAIDAFLAFHHGKESPVVAILAMYDTFDRGCEKSS- 770
Query: 209 GKGTIVCCTPLLHRWITSHLPRP 231
IVCC L+ W+ SHL P
Sbjct: 771 --ARIVCCALALYVWLVSHLFSP 833
>BI317622
Length = 370
Score = 55.8 bits (133), Expect = 3e-08
Identities = 40/123 (32%), Positives = 54/123 (43%), Gaps = 19/123 (15%)
Frame = -2
Query: 174 DKFVDMNAIRIFLTQN-----PVPVLLADTYVAIHDRTSHGKGTIVCCTPLLHRWITSHL 228
D VD+ AI FL + P+ +LAD Y + R IVCCTP L+ W+ SHL
Sbjct: 369 DGLVDLAAIGAFLAYHDYKEIPIVAMLADLYDTFNRRCEKSSTRIVCCTPALYVWLVSHL 190
Query: 229 PRPSIRPE-PIP------------WSQKLMTWTPKDIIWFNPSCDPEF-IIDSCGDFNNV 274
R +R P+ W Q L + WF + ++ SCG F NV
Sbjct: 189 FRQEVRHACPLESHCSCTEKGGENWGQLLANKEGASVNWFPRWKEGRTRVLISCGVFLNV 10
Query: 275 PLL 277
PL+
Sbjct: 9 PLM 1
>BE611176
Length = 424
Score = 38.9 bits (89), Expect(2) = 3e-05
Identities = 24/74 (32%), Positives = 38/74 (50%)
Frame = -2
Query: 5 GRRTHGYKFKAPDTESLEGLRGLTKMLKNPGHFQERYGNLLDILKTKVDVMLLNTLVQFY 64
G+R + K K D S++ L L L+ F++ Y +LD+ ++ + +L Q+Y
Sbjct: 363 GKRFYQVKIKGLDITSIKELGRLMGPLQMQA-FRKAYEKILDLTIAEISTEAIASLTQYY 187
Query: 65 DPIYHGFTFPDFQL 78
D FTF DFQL
Sbjct: 186 D*PLRCFTFGDFQL 145
Score = 26.2 bits (56), Expect(2) = 3e-05
Identities = 11/33 (33%), Positives = 18/33 (54%)
Frame = -1
Query: 80 PTLEEYSHWVGLPVLDEVPFHAFGPIPKIPTIA 112
PT+EE+ +G P+ P+ G +P + IA
Sbjct: 142 PTVEEFEEILGCPLRGRKPYFFSGFLPSLSKIA 44
>BG508280
Length = 418
Score = 45.1 bits (105), Expect = 4e-05
Identities = 19/33 (57%), Positives = 26/33 (78%)
Frame = +1
Query: 264 IIDSCGDFNNVPLLGTQGGISYSPILARRQFEY 296
++ SCG F NVPL+GT+G ISY+P+LA +Q Y
Sbjct: 70 VLISCGGFPNVPLMGTRGCISYNPVLAIKQLGY 168
>TC223279
Length = 539
Score = 31.6 bits (70), Expect = 0.51
Identities = 17/44 (38%), Positives = 23/44 (51%)
Frame = -3
Query: 173 VDKFVDMNAIRIFLTQNPVPVLLADTYVAIHDRTSHGKGTIVCC 216
++KF AI+IFLT P +A Y+ HGK I+CC
Sbjct: 447 LNKFDRSEAIQIFLTSTFDPFHIATLYLHRGP*HLHGKNCIICC 316
>TC223682 homologue to UP|SYE_ARATH (Q9FEA2) Glutamyl-tRNA synthetase
(Glutamate--tRNA ligase) (GluRS) , partial (19%)
Length = 461
Score = 31.2 bits (69), Expect = 0.67
Identities = 18/63 (28%), Positives = 25/63 (39%), Gaps = 2/63 (3%)
Frame = -2
Query: 203 HDRTSHGKGTIVCCTPLLHRWITSHLP--RPSIRPEPIPWSQKLMTWTPKDIIWFNPSCD 260
HDR G IVC H P ++ P + + L W P+ W +PS
Sbjct: 436 HDRIGVGSPHIVCIKNSSQTAYKDHNPLQHQALHPNQVQAKKDL*AWPPQTPYWTSPSQC 257
Query: 261 PEF 263
P+F
Sbjct: 256 PQF 248
>TC214657 similar to UP|Q9XGV6 (Q9XGV6) Bacterial-induced peroxidase
precursor , partial (72%)
Length = 1750
Score = 29.3 bits (64), Expect = 2.5
Identities = 16/43 (37%), Positives = 19/43 (43%)
Frame = +3
Query: 203 HDRTSHGKGTIVCCTPLLHRWITSHLPRPSIRPEPIPWSQKLM 245
H H K + C HRW H P PS P P P Q+L+
Sbjct: 369 HQLRHHQKSQLPC-----HRW--EHQPGPSRHPHPQPLRQQLL 476
>BM086125
Length = 428
Score = 29.3 bits (64), Expect = 2.5
Identities = 18/55 (32%), Positives = 22/55 (39%), Gaps = 7/55 (12%)
Frame = -2
Query: 203 HDRTSHGKGTIVCCTPLLHR---WITSHLPR----PSIRPEPIPWSQKLMTWTPK 250
H G+ T+ CC R WI P PS+ P+P W KL PK
Sbjct: 196 HRNEERGRNTLRCCCCYRWR*LHWIPIQAPLALLLPSLLPQPRCWCSKLSL*APK 32
>TC213622
Length = 607
Score = 28.9 bits (63), Expect = 3.3
Identities = 12/31 (38%), Positives = 18/31 (57%)
Frame = +3
Query: 224 ITSHLPRPSIRPEPIPWSQKLMTWTPKDIIW 254
+ S +P PS P P+ + L T+TPK +W
Sbjct: 174 VHSRIPMPSFPPSPL--TSALSTFTPKPAVW 260
>TC216967
Length = 1117
Score = 27.7 bits (60), Expect = 7.4
Identities = 18/68 (26%), Positives = 32/68 (46%), Gaps = 7/68 (10%)
Frame = +1
Query: 66 PIYHGFTFPDFQLFPTLEEYSHWVGLPVLDEVP----FHAFGPIPKIPTIAKALHLEVAD 121
P+ + FP++ FPTL ++ + G+P P F+ P TIA + +D
Sbjct: 124 PLANPAGFPEYYYFPTL-QFEPFGGMPFFTHAPPPAMFYPVAETPLTNTIANQIDYYFSD 300
Query: 122 ---IKDKF 126
+KD++
Sbjct: 301 ANLVKDEY 324
>TC221598 homologue to GB|AAM19966.1|20466089|AY098956 At2g32600/T26B15.16
{Arabidopsis thaliana;} , partial (64%)
Length = 542
Score = 27.7 bits (60), Expect = 7.4
Identities = 14/47 (29%), Positives = 21/47 (43%), Gaps = 3/47 (6%)
Frame = -1
Query: 239 PWSQKLMTWTPKDIIWFN---PSCDPEFIIDSCGDFNNVPLLGTQGG 282
PW ++L W WF P C+P CG+ + V + +GG
Sbjct: 275 PWPRELHVWPS----WFGGAFPECEPSSCPRCCGE*DRVCIRSFRGG 147
>TC215322 similar to UP|Q9LSL3 (Q9LSL3) Oligopeptidase A, partial (56%)
Length = 1338
Score = 27.3 bits (59), Expect = 9.6
Identities = 20/72 (27%), Positives = 31/72 (42%), Gaps = 3/72 (4%)
Frame = -1
Query: 142 QKATTCFEKSKTDDFESILAL---LIYGIVLFPKVDKFVDMNAIRIFLTQNPVPVLLADT 198
Q+A+ + + F++ L +YG FP +K + Q P P L
Sbjct: 741 QQASNILLQGDSPQFQNALQCPLKFLYGNTSFP*TEKVTQQHPTLSL*YQQPDPHLA**A 562
Query: 199 YVAIHDRTSHGK 210
YV +HD+ HGK
Sbjct: 561 YVEVHDQI-HGK 529
>TC221584 weakly similar to UP|Q9FH22 (Q9FH22) Gb|AAF17687.1, partial (35%)
Length = 694
Score = 27.3 bits (59), Expect = 9.6
Identities = 14/40 (35%), Positives = 21/40 (52%)
Frame = -3
Query: 51 KVDVMLLNTLVQFYDPIYHGFTFPDFQLFPTLEEYSHWVG 90
K+ + L +TL P+Y +FP LFP + +S VG
Sbjct: 311 KICLHLQSTLQNALSPLYM*MSFPGKSLFPQRKHWSRKVG 192
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.144 0.458
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,655,657
Number of Sequences: 63676
Number of extensions: 289422
Number of successful extensions: 1378
Number of sequences better than 10.0: 29
Number of HSP's better than 10.0 without gapping: 1363
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1373
length of query: 324
length of database: 12,639,632
effective HSP length: 97
effective length of query: 227
effective length of database: 6,463,060
effective search space: 1467114620
effective search space used: 1467114620
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC134521.1


