

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.24 + phase: 0
(456 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
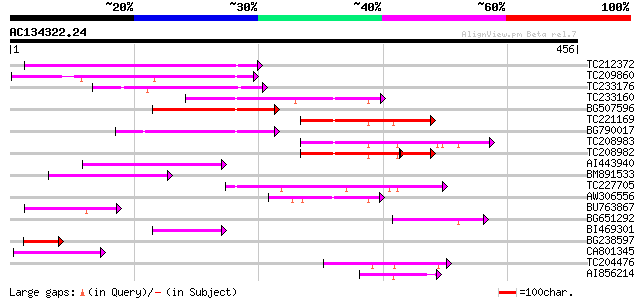
Score E
Sequences producing significant alignments: (bits) Value
TC212372 similar to GB|AAN46837.1|24111405|BT001072 At5g62000/mt... 126 2e-29
TC209860 similar to GB|AAB62404.1|2245390|ATU89296 auxin respons... 117 8e-27
TC233176 weakly similar to GB|AAL07033.1|15810291|AY056184 auxin... 103 2e-22
TC233160 similar to UP|Q8S976 (Q8S976) Auxin response factor 10 ... 99 5e-21
BG507596 homologue to GP|9757747|dbj auxin response factor 4 {Ar... 90 2e-18
TC221169 similar to UP|ARFJ_ARATH (Q9SKN5) Auxin response factor... 85 6e-17
BG790017 similar to PIR|T08917|T08 auxin response factor 9 - Ara... 82 7e-16
TC208983 similar to UP|Q8S976 (Q8S976) Auxin response factor 10 ... 81 9e-16
TC208982 similar to UP|Q8S976 (Q8S976) Auxin response factor 10 ... 57 2e-12
AI443940 homologue to GP|19352043|db auxin response factor 6b {O... 69 6e-12
BM891533 similar to GP|10176918|db auxin response factor-like pr... 68 1e-11
TC227705 similar to GB|AAM91657.1|22136676|AY133723 auxin respon... 66 3e-11
AW306556 weakly similar to GP|19352051|dbj auxin response factor... 64 2e-10
BU763867 similar to GP|10176918|dbj auxin response factor-like p... 58 8e-09
BG651292 55 7e-08
BI469301 homologue to GP|19352043|db auxin response factor 6b {O... 52 4e-07
BG238597 similar to GP|19352051|dbj auxin response factor 10 {Or... 49 4e-06
CA801345 similar to GP|19352045|db auxin response factor 7a {Ory... 49 5e-06
TC204476 similar to UP|Q84QI6 (Q84QI6) Auxin response factor-lik... 48 1e-05
AI856214 47 2e-05
>TC212372 similar to GB|AAN46837.1|24111405|BT001072 At5g62000/mtg10_20
{Arabidopsis thaliana;} , partial (31%)
Length = 878
Score = 126 bits (317), Expect = 2e-29
Identities = 71/191 (37%), Positives = 97/191 (50%)
Frame = +1
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVD 72
E+WH CA V +P+ RV+YFPQGH+E S++ + H + P LC V V
Sbjct: 280 ELWHACAGPLVTVPRERERVFYFPQGHIEQVEASTNQVAEQHMPVYDLPPKILCRVINVM 459
Query: 73 LLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFH 132
L A+P TDEVF ++ L P N KE R V SF KTLT SD + F
Sbjct: 460 LKAEPDTDEVFAQVTLLPEPNQDENAVEKEGPPAAPPRFHVHSFCKTLTASDTSTHGGFS 639
Query: 133 IPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKR 192
+ R AD P LD+ + +Q L D+HG +F H+ RG P++++L S WN FV
Sbjct: 640 VLRRHADECLPPLDMTKQPPTQELVAKDLHGNXWRFRHIFRGQPRKHLLQ-SGWNVFVTP 816
Query: 193 KKLVAGDSVIF 203
+ G + F
Sbjct: 817 XSXLLGMRLXF 849
>TC209860 similar to GB|AAB62404.1|2245390|ATU89296 auxin response
transcription factor 3 {Arabidopsis thaliana;} , partial
(32%)
Length = 687
Score = 117 bits (294), Expect = 8e-27
Identities = 73/209 (34%), Positives = 104/209 (48%), Gaps = 10/209 (4%)
Frame = +3
Query: 2 SLQKQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSF----L 57
S S V E+WH CA + +PK S V YFPQGHLE H H F
Sbjct: 90 SSSSSSSTVCLELWHACAGPMISLPKKGSVVVYFPQGHLEQ---------HLHDFPLPAS 242
Query: 58 QSFRPFTLCIVSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVS-- 115
+ C V V L A+ +DEV+ +++L P + E + D V+
Sbjct: 243 ANIPSHVFCRVLDVKLHAEEGSDEVYCQVVLVPESEQKLREGEFDADGEEEDAEAVMKST 422
Query: 116 ----FVKTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHV 171
F KTLT SD + F +PR A++ FP LD + SQ L D+HG+ +F H+
Sbjct: 423 TPHMFCKTLTASDTSTHGGFSVPRRAAEDCFPPLDYSQQRPSQELVAKDLHGQEWRFRHI 602
Query: 172 CRGFPKRNMLYISEWNSFVKRKKLVAGDS 200
RG P+R++L + W++FV +KKLV+GD+
Sbjct: 603 YRGQPRRHLL-TTGWSAFVNKKKLVSGDA 686
>TC233176 weakly similar to GB|AAL07033.1|15810291|AY056184 auxin response
factor ARF17 {Arabidopsis thaliana;} , partial (17%)
Length = 958
Score = 103 bits (256), Expect = 2e-22
Identities = 60/146 (41%), Positives = 81/146 (55%), Gaps = 5/146 (3%)
Frame = +2
Query: 67 IVSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLND-----RNEVVSFVKTLT 121
I SA+ P + +F+ LL+ + +N +EV N +D N V+SF K LT
Sbjct: 59 ITSALSSALSPSSPAMFLLTLLSQ-SQQQPFQNDTKEV*NDDDDKDRENNNVISFAKILT 235
Query: 122 RSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNML 181
SD NN F +PRFCAD+ FP LD + L V D+HG +F H+ G P+ +
Sbjct: 236 PSDANNGDDFSVPRFCADSCFPPLDFRTDLPV*LLSVADIHGVEWRFRHIYHGTPRHHFF 415
Query: 182 YISEWNSFVKRKKLVAGDSVIFMKDS 207
I W+ FV KKLVAGD+VIF+KDS
Sbjct: 416 TIG-WSKFVNHKKLVAGDTVIFVKDS 490
>TC233160 similar to UP|Q8S976 (Q8S976) Auxin response factor 10 (Fragment),
partial (26%)
Length = 658
Score = 98.6 bits (244), Expect = 5e-21
Identities = 67/210 (31%), Positives = 100/210 (46%), Gaps = 49/210 (23%)
Frame = +2
Query: 142 FPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKRKKLVAGDSV 201
FP+LD + Q++ DVHGE KF H+ RG P+R++L + W++FV KKL AGDS+
Sbjct: 2 FPRLDYSVDPPVQNILAKDVHGETWKFRHIYRGTPRRHLL-TTGWSTFVNHKKLFAGDSI 178
Query: 202 IFMKDSTGKIFVGIRRNTQFVAAAAEQ--------------------------------- 228
+F++ G + VGIRR + + E
Sbjct: 179 VFLRAENGDLCVGIRRAKKGICGGLETSSGWNPAGGNCHIPYGGFSPFFREDDNRISRNG 358
Query: 229 -------------KKDELEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDESM 275
K +AV EA LA K FE+VYYP+ +F V ++V+ ++
Sbjct: 359 NSNGLNPSVSMMGKGKVRPEAVSEASNLAANKKPFEVVYYPRAST-PEFCVKASLVEAAL 535
Query: 276 KIQWNPRMRVKM--KTDKSSRIP-YQGTIS 302
+I+W +R KM +T+ SSRI + GTIS
Sbjct: 536 QIRWCSGIRFKMAFETEDSSRISWFMGTIS 625
>BG507596 homologue to GP|9757747|dbj auxin response factor 4 {Arabidopsis
thaliana}, partial (15%)
Length = 433
Score = 90.1 bits (222), Expect = 2e-18
Identities = 44/102 (43%), Positives = 66/102 (64%)
Frame = +3
Query: 116 FVKTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRGF 175
F KTLT SD + F +PR A++ FP LD + + SQ L D+HG KF H+ RG
Sbjct: 72 FCKTLTASDTSTHGGFSVPRRAAEDCFPPLDYKKQRPSQELVAKDLHGVEWKFRHIYRGQ 251
Query: 176 PKRNMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRR 217
P+R++L + W+ FV +K LV+GD+V+F++ G++ +GIRR
Sbjct: 252 PRRHLL-TTGWSIFVSQKNLVSGDAVLFLRGENGELRLGIRR 374
>TC221169 similar to UP|ARFJ_ARATH (Q9SKN5) Auxin response factor 10, partial
(18%)
Length = 562
Score = 85.1 bits (209), Expect = 6e-17
Identities = 49/116 (42%), Positives = 75/116 (64%), Gaps = 8/116 (6%)
Frame = +2
Query: 235 KAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKS 292
+AV+EA LA + FE+VYYP+ +F V N+V +++++W P MR KM +T+ S
Sbjct: 110 EAVIEAATLAANMQPFEVVYYPRASA-PEFCVKANLVRAALQVRWCPGMRFKMPFETEDS 286
Query: 293 SRIP-YQGTISIVSRT-----SNLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKP 342
SRI + GTIS V+ ++ WR+LQV WDE ++ Q +RV+PW VE++S+ P
Sbjct: 287 SRISWFMGTISSVNFADPRWPNSPWRLLQVTWDEPELLQNVKRVSPWLVEIVSNMP 454
>BG790017 similar to PIR|T08917|T08 auxin response factor 9 - Arabidopsis
thaliana, partial (21%)
Length = 440
Score = 81.6 bits (200), Expect = 7e-16
Identities = 47/132 (35%), Positives = 73/132 (54%)
Frame = +1
Query: 86 LLLTPITNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSFHIPRFCADNVFPQL 145
+ L P +N +P A L R V SF K LT SD + F + R A P L
Sbjct: 1 ITLVPESNQTEPTSPDPCPAEL-PRPRVHSFCKVLTASDTSTHGGFSVLRKHATECLPAL 177
Query: 146 DLEAESSSQHLFVTDVHGEVSKFYHVCRGFPKRNMLYISEWNSFVKRKKLVAGDSVIFMK 205
D+ + +Q L D+ G +F H+ RG P+R++L + W++FV K+LVAGD+ +F++
Sbjct: 178 DMSKSTPTQELVAKDLQGFEWRFKHIFRGQPRRHLL-TTGWSTFVTSKRLVAGDTFVFLR 354
Query: 206 DSTGKIFVGIRR 217
+ G++ VG+RR
Sbjct: 355 GNNGELRVGVRR 390
>TC208983 similar to UP|Q8S976 (Q8S976) Auxin response factor 10 (Fragment),
partial (26%)
Length = 958
Score = 81.3 bits (199), Expect = 9e-16
Identities = 64/180 (35%), Positives = 89/180 (48%), Gaps = 24/180 (13%)
Frame = +1
Query: 235 KAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKS 292
++V EA LA N+AFE+VYYP+ + +F + + V +M+IQW MR KM +T+ S
Sbjct: 100 ESVREAGTLAASNQAFEVVYYPRANT-PEFCIRTSAVRGAMRIQWCSGMRFKMPFETEDS 276
Query: 293 SRIP-YQGTISIVSRTSNL------WRMLQVNWDEFQVSQIPRRVNPWWVELISHKP--- 342
SRI + GTI+ V + WR+LQV WDE + +RV+PW VEL+S+ P
Sbjct: 277 SRISWFMGTIASVQVLDPIRWPNSPWRLLQVTWDEPDLLHNVKRVSPWLVELVSNVPIIH 456
Query: 343 --APTP------FPQTKKFRTTQSS----AQLSDKKETLLNGDGFPVDIQRVRHDLVSIS 390
A +P FP +F S S D P IQ RH + IS
Sbjct: 457 LAAFSPPRKKLRFPLDVQFPIPSFSGNPFGSSSSSSPFCCLSDNAPAGIQGARHAQIGIS 636
>TC208982 similar to UP|Q8S976 (Q8S976) Auxin response factor 10 (Fragment),
partial (19%)
Length = 675
Score = 56.6 bits (135), Expect(2) = 2e-12
Identities = 36/92 (39%), Positives = 56/92 (60%), Gaps = 9/92 (9%)
Frame = +2
Query: 235 KAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKS 292
++V EA+ LA N+ FE+VYYP+ + +F + + V +M+IQW+ MR KM +T+ S
Sbjct: 122 ESVREAVTLAASNQPFEVVYYPRANT-PEFCIRTSAVRGAMRIQWSSGMRFKMPFETEDS 298
Query: 293 SRIP-YQGTISIVSRTSNL------WRMLQVN 317
SRI + GTI+ V + WR+LQV+
Sbjct: 299 SRISWFMGTIASVQLLDPIRWPNSPWRLLQVH 394
Score = 33.9 bits (76), Expect(2) = 2e-12
Identities = 14/28 (50%), Positives = 20/28 (71%)
Frame = +3
Query: 315 QVNWDEFQVSQIPRRVNPWWVELISHKP 342
QV+WDE + +RV+PW VEL+S+ P
Sbjct: 453 QVSWDEPDLLHNVKRVSPWLVELVSNVP 536
>AI443940 homologue to GP|19352043|db auxin response factor 6b {Oryza
sativa}, partial (14%)
Length = 391
Score = 68.6 bits (166), Expect = 6e-12
Identities = 38/116 (32%), Positives = 54/116 (45%)
Frame = +3
Query: 59 SFRPFTLCIVSAVDLLADPHTDEVFVKLLLTPITNDVHLENPKEEVANLNDRNEVVSFVK 118
S P +C + + + AD TDEV+ ++ L P+ E + F K
Sbjct: 39 SLPPQLICQLHNMTMHADVETDEVYAQMTLQPLNPQEQNEAYLPAELGTASKQPTNYFCK 218
Query: 119 TLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRG 174
TLT SD + F +PR A+ VFP LD + +Q L D+HG KF H+ RG
Sbjct: 219 TLTASDTSTHGGFSVPRRAAEKVFPPLDFSQQPPAQELIARDLHGNEWKFRHIFRG 386
>BM891533 similar to GP|10176918|db auxin response factor-like protein
{Arabidopsis thaliana}, partial (11%)
Length = 306
Score = 67.8 bits (164), Expect = 1e-11
Identities = 39/100 (39%), Positives = 49/100 (49%)
Frame = +3
Query: 32 VYYFPQGHLENASPSSSSITHTHSFLQSFRPFTLCIVSAVDLLADPHTDEVFVKLLLTPI 91
V+YFPQGH+E S++ + H + P LC V V L A+P TDEVF ++ L P
Sbjct: 3 VFYFPQGHIEQVEASTNQVADQHMPVYDLPPKILCRVINVQLKAEPDTDEVFAQVTLLPE 182
Query: 92 TNDVHLENPKEEVANLNDRNEVVSFVKTLTRSDVNNARSF 131
N KE R V SF KTLT SD + F
Sbjct: 183 PNQYFFAVEKEPPPPPPPRFHVHSFCKTLTASDTSTHGGF 302
>TC227705 similar to GB|AAM91657.1|22136676|AY133723 auxin response factor 1
{Arabidopsis thaliana;} , partial (54%)
Length = 1772
Score = 66.2 bits (160), Expect = 3e-11
Identities = 55/191 (28%), Positives = 91/191 (46%), Gaps = 12/191 (6%)
Frame = +3
Query: 174 GFPKRNMLYISEWNSFVKRKKLVAGDSVIFMKDSTGKIFVGIRR--NTQFVAAAAEQKKD 231
G PKR++L + W+ FV KKL AGD+ IF++ G++ VG+RR Q ++
Sbjct: 165 GQPKRHLL-TTGWSVFVSSKKLAAGDAFIFLRGENGELRVGVRRVMRQQSNVPSSVISSH 341
Query: 232 ELEKAVMEALKLAEENKAFEIVYYPQGDDWCDFVVDGN--VVDESMKIQWNPRMRVKMKT 289
+ V+ A V+Y +F+V N + +S K+ R +++ +
Sbjct: 342 SMHLGVLATASHAIATGTLFSVFYKPRTSRSEFIVSVNKYLEVQSHKLSVGMRFKMRFEG 521
Query: 290 DKSSRIPYQGTISIV--SRTSNL-----WRMLQVNWDEFQVSQIPRRVNPWWVE-LISHK 341
D+ + GTI V +++S++ WR L+V WDE P RV+ W +E L+S
Sbjct: 522 DEIPERRFSGTIVGVGDNKSSSVWPDSEWRSLKVQWDEPSSIMRPDRVSSWELEPLVSTT 701
Query: 342 PAPTPFPQTKK 352
A + Q K
Sbjct: 702 LANSQTTQRNK 734
>AW306556 weakly similar to GP|19352051|dbj auxin response factor 10 {Oryza
sativa}, partial (9%)
Length = 396
Score = 63.5 bits (153), Expect = 2e-10
Identities = 43/121 (35%), Positives = 64/121 (52%), Gaps = 28/121 (23%)
Frame = +1
Query: 209 GKIFVGIRRNTQFVAAAA----------EQKKDELE---------------KAVMEALKL 243
G + VGIRR T F E++++E E K V EA +L
Sbjct: 10 GGLLVGIRRTTLFSPGKGGDVGTRIKVDEEEEEEEEVREVFSRDGRGKLSAKVVAEAAEL 189
Query: 244 AEENKAFEIVYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKM--KTDKSSRIPY-QGT 300
A + FE+VYYP+G W +FVV V+E+M ++W+ M+VK+ +TD SSR+ + QGT
Sbjct: 190 AARSMPFEVVYYPKGG-WSEFVVKTEAVNEAMSVEWSHGMKVKIATETDDSSRVSWCQGT 366
Query: 301 I 301
+
Sbjct: 367 V 369
>BU763867 similar to GP|10176918|dbj auxin response factor-like protein
{Arabidopsis thaliana}, partial (8%)
Length = 452
Score = 58.2 bits (139), Expect = 8e-09
Identities = 27/80 (33%), Positives = 47/80 (58%), Gaps = 2/80 (2%)
Frame = +1
Query: 13 EIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTHSFLQSF--RPFTLCIVSA 70
E+W CA + V +P++ RV+YFPQGHLE + + + +H + + LC +
Sbjct: 139 ELWRACAGSFVYVPRVDDRVFYFPQGHLEQVAAYTQNQPDSHLEIPVYDLPSKILCKIMN 318
Query: 71 VDLLADPHTDEVFVKLLLTP 90
V+L A+ ++DEV+ ++ L P
Sbjct: 319 VELKAEAYSDEVYAQVTLVP 378
>BG651292
Length = 348
Score = 55.1 bits (131), Expect = 7e-08
Identities = 33/96 (34%), Positives = 46/96 (47%), Gaps = 19/96 (19%)
Frame = +3
Query: 309 NLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA-PTPFPQTKKFRTTQSS-------- 359
+LWRMLQV WDE + QI + V+PW VEL+S PA + FP K+ + S
Sbjct: 57 SLWRMLQVTWDEPEGLQIAKWVSPWQVELVSTTPALHSAFPPIKRIKAAHDSGVFTNGER 236
Query: 360 ----------AQLSDKKETLLNGDGFPVDIQRVRHD 385
+ + + LL+ FP +Q RHD
Sbjct: 237 DPFPMTGFTNSTMGQLNQALLSYGTFPAGMQGARHD 344
>BI469301 homologue to GP|19352043|db auxin response factor 6b {Oryza
sativa}, partial (15%)
Length = 422
Score = 52.4 bits (124), Expect = 4e-07
Identities = 25/59 (42%), Positives = 32/59 (53%)
Frame = +3
Query: 116 FVKTLTRSDVNNARSFHIPRFCADNVFPQLDLEAESSSQHLFVTDVHGEVSKFYHVCRG 174
F KTLT SD + F +PR A+ VFP LD + +Q L D+H KF H+ RG
Sbjct: 243 FCKTLTASDTSTHGGFSVPRRAAEKVFPPLDYSQQPPAQELIARDLHDNEWKFRHIFRG 419
>BG238597 similar to GP|19352051|dbj auxin response factor 10 {Oryza sativa},
partial (5%)
Length = 487
Score = 49.3 bits (116), Expect = 4e-06
Identities = 17/32 (53%), Positives = 26/32 (81%)
Frame = +1
Query: 12 PEIWHTCATAAVKIPKLHSRVYYFPQGHLENA 43
P++WH CA V++P ++++VYYFPQGH E+A
Sbjct: 376 PQLWHACAGGMVQMPTVNTKVYYFPQGHAEHA 471
>CA801345 similar to GP|19352045|db auxin response factor 7a {Oryza sativa},
partial (6%)
Length = 424
Score = 48.9 bits (115), Expect = 5e-06
Identities = 24/75 (32%), Positives = 38/75 (50%), Gaps = 1/75 (1%)
Frame = +2
Query: 4 QKQPSHVRPEIWHTCATAAVKIPKLHSRVYYFPQGHLENASPSSSSITHTH-SFLQSFRP 62
+++ + PE+W CA V +P + V YFPQGH E + S + H+ +
Sbjct: 197 EEKKKSINPELWQACAGPLVNLPPSGTHVIYFPQGHSEQVAASLNKDPHSQIPNYPNLPS 376
Query: 63 FTLCIVSAVDLLADP 77
LC++ + LLADP
Sbjct: 377 KLLCLLHNLTLLADP 421
>TC204476 similar to UP|Q84QI6 (Q84QI6) Auxin response factor-like protein,
partial (43%)
Length = 2243
Score = 47.8 bits (112), Expect = 1e-05
Identities = 33/113 (29%), Positives = 55/113 (48%), Gaps = 10/113 (8%)
Frame = +3
Query: 253 VYYPQGDDWCDFVVDGNVVDESMKIQWNPRMRVKMKTD--KSSRIPYQGTISIVSRTS-- 308
VYY +F+V + ES+K ++ MR KM+ + ++ + GT+ + +
Sbjct: 54 VYYKPRTSPAEFIVPYDQYMESLKNNYSIGMRFKMRFEGEEAPEQRFTGTVVGIEDSDPK 233
Query: 309 ----NLWRMLQVNWDEFQVSQIPRRVNPWWVELISHKPA--PTPFPQTKKFRT 355
+ WR L+V WDE + P RV+PW +E PA P P+ K+ R+
Sbjct: 234 RWRDSKWRCLKVRWDETSNTPRPERVSPWKIEPALAPPALNPLSMPRPKRPRS 392
>AI856214
Length = 464
Score = 46.6 bits (109), Expect = 2e-05
Identities = 27/72 (37%), Positives = 38/72 (52%), Gaps = 6/72 (8%)
Frame = -1
Query: 282 RMRVKMKTDKSSRIPYQGTISIVSRT------SNLWRMLQVNWDEFQVSQIPRRVNPWWV 335
R R+ +T++S Y GTI+ +S S+ WR +QV WDE + PRRV+ W +
Sbjct: 440 RFRMMFETEESGVRGYMGTITGISDLDPVRWKSSQWRNIQVGWDESTAGERPRRVSIWEI 261
Query: 336 ELISHKPAPTPF 347
E P TPF
Sbjct: 260 E-----PVVTPF 240
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.133 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,544,012
Number of Sequences: 63676
Number of extensions: 323886
Number of successful extensions: 1844
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 1816
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1829
length of query: 456
length of database: 12,639,632
effective HSP length: 100
effective length of query: 356
effective length of database: 6,272,032
effective search space: 2232843392
effective search space used: 2232843392
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC134322.24


