

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134322.2 + phase: 0 /pseudo
(840 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
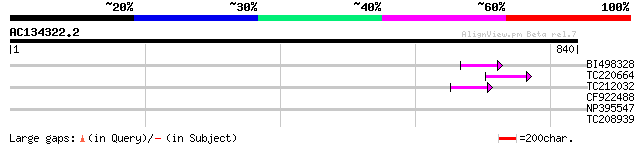
Sequences producing significant alignments: (bits) Value
BI498328 49 7e-06
TC220664 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial... 45 1e-04
TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 43 7e-04
CF922488 33 0.68
NP395547 reverse transcriptase [Glycine max] 30 4.4
TC208939 UP|Q39830 (Q39830) BiP isoform A, complete 29 7.6
>BI498328
Length = 335
Score = 49.3 bits (116), Expect = 7e-06
Identities = 25/62 (40%), Positives = 33/62 (52%)
Frame = +1
Query: 668 KWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSSNNQAEYEALFAGMRLALKVV 727
KW++ DG+SN G G G L P I + R F +NN AEYEA G++ A+
Sbjct: 148 KWIVCFDGASNALGHGVGAVLVSPDDQCIPFTARLGFDCTNNMAEYEACALGVQAAIDFD 327
Query: 728 VK 729
VK
Sbjct: 328 VK 333
>TC220664 weakly similar to UP|Q84TE0 (Q84TE0) At5g51080, partial (31%)
Length = 724
Score = 45.4 bits (106), Expect = 1e-04
Identities = 24/68 (35%), Positives = 37/68 (54%)
Frame = +1
Query: 706 SSNNQAEYEALFAGMRLALKVVVKELEAKSDSQLVFRQVTGNFQTKDPQLLMYLESVMYL 765
++NN AEY A+ GM+ ALK + + DS+LV Q+ G+++ K+ L L
Sbjct: 61 ATNNAAEYRAMILGMKYALKKGFTGIRIQGDSKLVCMQIDGSWKVKNENLSTLYNVAKEL 240
Query: 766 TQKFVSFK 773
KF SF+
Sbjct: 241 KDKFSSFQ 264
>TC212032 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (3%)
Length = 803
Score = 42.7 bits (99), Expect = 7e-04
Identities = 22/62 (35%), Positives = 32/62 (51%)
Frame = +1
Query: 654 LVEMTAEKGSMEAAKWVLSVDGSSNLKGSGAGVTLEGPYGVLIEQSLRFDFKSSNNQAEY 713
++ + EK + KW++ D +SN+ G G G L P I + R F +NN AEY
Sbjct: 7 IMALFEEKLDEDRDKWIVWFDRASNVLGHGVGAILVSPDNQCIPFTTRLGFDCTNNMAEY 186
Query: 714 EA 715
EA
Sbjct: 187 EA 192
>CF922488
Length = 741
Score = 32.7 bits (73), Expect = 0.68
Identities = 61/221 (27%), Positives = 86/221 (38%)
Frame = +1
Query: 308 ECARTTLI*IRCVPRMLIPCRA*INWYITLQGMTFLASWMLTRATTKFECT*RMKKRQLL 367
ECA T I* + V R+ R +IT SWM + T+ R+ KRQL
Sbjct: 28 ECAWTIEI*TKPVQRISFLYRISTFSWITRPVFPNSPSWMDFQVITR*R*HQRIWKRQLS 207
Query: 368 *QTKLIIVTKVCHLG*GILELLFKDWWIKSSRVR*ARRSRCM*MTWW*NRQNKMIIKASS 427
++C LG* +L W SR RR R MT *N++ + +
Sbjct: 208 LLYGEPSAIRLCRLG*RMLGQHTSGPWWHYSRT*CTRR*RSTWMT*S*NQERRRNTLSIC 387
Query: 428 TRYLTCLRGTISSLT*RNVHLEYKQESF*ASCLLGGE*RSIQKSARPSWI*GVQLMRRRY 487
L T ++V L ES + E*R I+ + S + R +
Sbjct: 388 ESCLGDYVNTG*D*IPQSVCLR*NPESCSTLLIAREE*RWIRTR*K*SLRWPSHIQRSKS 567
Query: 488 KN*LGGLLHSRGFYHARLIKTYLFSNCCQKMKIFVGRKSVR 528
K GG S YH+ L LFS CC ++ + G V+
Sbjct: 568 KVSWGG*TTS*DSYHS*LPLASLFSYCCARISLSNGTMIVK 690
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 30.0 bits (66), Expect = 4.4
Identities = 18/56 (32%), Positives = 28/56 (49%)
Frame = +2
Query: 337 LQGMTFLASWMLTRATTKFECT*RMKKRQLL*QTKLIIVTKVCHLG*GILELLFKD 392
LQG SW T+ T + + R+KK+QLL + ++ C + LLF+D
Sbjct: 251 LQGNPSTVSWTDTQVTIRLQWILRIKKKQLLHVLSVFLLIAACRSVYVMPLLLFRD 418
>TC208939 UP|Q39830 (Q39830) BiP isoform A, complete
Length = 2452
Score = 29.3 bits (64), Expect = 7.6
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = -3
Query: 251 IQWQGLYNKESVRWVWRNKRQYK 273
I ++G +N W WRN RQ+K
Sbjct: 1589 INFEGHFNLRGTPWSWRNSRQFK 1521
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.358 0.156 0.572
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,235,798
Number of Sequences: 63676
Number of extensions: 530732
Number of successful extensions: 5048
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 2708
Number of HSP's successfully gapped in prelim test: 138
Number of HSP's that attempted gapping in prelim test: 2285
Number of HSP's gapped (non-prelim): 2951
length of query: 840
length of database: 12,639,632
effective HSP length: 105
effective length of query: 735
effective length of database: 5,953,652
effective search space: 4375934220
effective search space used: 4375934220
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC134322.2


