

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
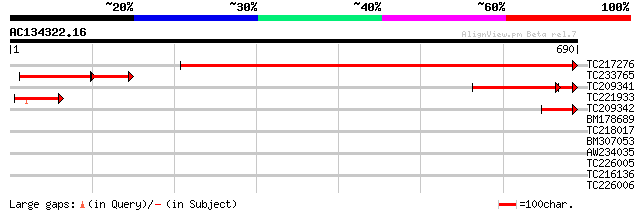
Query= AC134322.16 - phase: 0
(690 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC217276 weakly similar to GB|AAH11654.1|15079675|BC011654 cleav... 896 0.0
TC233765 similar to UP|Q8WZS6 (Q8WZS6) Related to BRR5 (Componen... 155 4e-60
TC209341 174 3e-47
TC221933 weakly similar to UP|Q9VE51 (Q9VE51) CG7698-PA (RE31408... 99 5e-21
TC209342 80 4e-15
BM178689 similar to SP|Q9LKF9|CPSB Cleavage and polyadenylation ... 35 0.085
TC218017 similar to GB|AAF82809.1|9082326|AF283277 polyadenylati... 32 0.72
BM307053 similar to GP|9082326|gb|A polyadenylation cleavage/spe... 32 0.94
AW234035 31 1.6
TC226005 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), parti... 30 3.6
TC216136 30 4.7
TC226006 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), parti... 29 6.1
>TC217276 weakly similar to GB|AAH11654.1|15079675|BC011654 cleavage and
polyadenylation specific factor 3, 73kDa {Homo sapiens;}
, partial (31%)
Length = 1853
Score = 896 bits (2315), Expect = 0.0
Identities = 450/482 (93%), Positives = 469/482 (96%)
Frame = +3
Query: 209 DVCIIESTYGVQHHQPRHTREKRFTDVIHSTISQGGRVLIPAYALGRAQELLLILDEYWA 268
DVCIIESTYGVQHHQPRHTREKRFTDVIHSTISQGGRVLIPA+ALGRAQELLLILDEYWA
Sbjct: 6 DVCIIESTYGVQHHQPRHTREKRFTDVIHSTISQGGRVLIPAFALGRAQELLLILDEYWA 185
Query: 269 NHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRIQNAKSNPFAFKHISALSSIDIFKDV 328
NHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRIQNAKSNPF+FKH+SALSSI++FKDV
Sbjct: 186 NHPELQNIPIYYASPLAKKCLTVYETYTLSMNDRIQNAKSNPFSFKHVSALSSIEVFKDV 365
Query: 329 GPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVIPGYVVEGTLAKTILNEPKEVTLMNGL 388
GPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCV+PGYVVEGTLAKTI+NEPKEVTLMNGL
Sbjct: 366 GPSVVMASPGGLQSGLSRQLFDMWCSDKKNSCVLPGYVVEGTLAKTIINEPKEVTLMNGL 545
Query: 389 SAPLHMQVHYISFSAHADSAQTSAFLEELNPPNIILVHGAANEMGRLKQKLMTQFADRNT 448
+APL+MQVHYISFSAHADSAQTSAFLEELNPPNIILVHG ANEMGRLKQKL++QFADRNT
Sbjct: 546 TAPLNMQVHYISFSAHADSAQTSAFLEELNPPNIILVHGEANEMGRLKQKLISQFADRNT 725
Query: 449 KILTPKNCQSVEMYFNSQKMAKTIGKLAEKTPEVGETVSGLLVKKGFTYQIMAPDDLHVF 508
KILTPKNCQSVEMYFNSQKMAKTIGKLAEKTPEVGETVSGLLVKKGFTYQIMA DDLHVF
Sbjct: 726 KILTPKNCQSVEMYFNSQKMAKTIGKLAEKTPEVGETVSGLLVKKGFTYQIMAADDLHVF 905
Query: 509 SQLSTANVTQRITIPYSGAFCVIQSRLKQIYESVEPSVDEESGVPMLLVHDRVTVKHESE 568
SQLSTAN+TQRITIPYSGAF VIQ RLKQIYESV SVDEESGVP L VH+ VTVKHESE
Sbjct: 906 SQLSTANITQRITIPYSGAFNVIQHRLKQIYESVAQSVDEESGVPTLQVHECVTVKHESE 1085
Query: 569 KHVSLHWASDPINDMVSDSVVALVLNINRDLPKIVAESDATKIEEENEKKTEKVMQALLN 628
KHVSLHWASDP++DMVSDS+VALVLNINRD+PKIV ESDA KIEEENEKK EKVMQALL
Sbjct: 1086KHVSLHWASDPMSDMVSDSIVALVLNINRDVPKIVNESDAIKIEEENEKKAEKVMQALLV 1265
Query: 629 SLFGNVKVGENGKLIINIDGNVAELNKESGEVESENEGLKERVRTAFRRIQSSVKPIPLS 688
SLFG+VKVGENGKLIINIDGNVAELNKESGEVESENEGLKERVRTAFRRIQSSVKPIP+S
Sbjct: 1266SLFGDVKVGENGKLIINIDGNVAELNKESGEVESENEGLKERVRTAFRRIQSSVKPIPVS 1445
Query: 689 AP 690
AP
Sbjct: 1446AP 1451
>TC233765 similar to UP|Q8WZS6 (Q8WZS6) Related to BRR5 (Component of
pre-mRNA polyadenylation factor PF I), partial (9%)
Length = 423
Score = 155 bits (392), Expect(2) = 4e-60
Identities = 70/92 (76%), Positives = 82/92 (89%)
Frame = +2
Query: 13 TINRETEDQLIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPS 72
++ E ++ +IVTPLGAGNEVGRSCVYM+YKGK++LFDCGIH G+SGM+ALPYFDEIDPS
Sbjct: 8 SVREEDDEVMIVTPLGAGNEVGRSCVYMSYKGKSILFDCGIHLGFSGMSALPYFDEIDPS 187
Query: 73 TVDVLLITHFHLDHAASLPYFLEKTTFKGRVF 104
T+DVLLITHFHLDHAASLPYFLEKT + F
Sbjct: 188 TLDVLLITHFHLDHAASLPYFLEKTNLRRPAF 283
Score = 95.5 bits (236), Expect(2) = 4e-60
Identities = 47/51 (92%), Positives = 50/51 (97%)
Frame = +1
Query: 100 KGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQDINRSMDKIEVIDF 150
K RVFMTYATKAIYKLLLSD+VKVSKVSV+DML+DEQDINRSMDKIEVIDF
Sbjct: 271 KARVFMTYATKAIYKLLLSDFVKVSKVSVEDMLFDEQDINRSMDKIEVIDF 423
>TC209341
Length = 412
Score = 174 bits (440), Expect(2) = 3e-47
Identities = 84/107 (78%), Positives = 97/107 (90%)
Frame = +3
Query: 564 KHESEKHVSLHWASDPINDMVSDSVVALVLNINRDLPKIVAESDATKIEEENEKKTEKVM 623
KHE+EKH+SLHW SDPI+DMVSDS+VAL+LNINRD+PKI+AESD KIEEEN+KK EKVM
Sbjct: 3 KHEAEKHISLHWTSDPISDMVSDSIVALILNINRDVPKIMAESDVIKIEEENKKKAEKVM 182
Query: 624 QALLNSLFGNVKVGENGKLIINIDGNVAELNKESGEVESENEGLKER 670
ALL SLFG+VK GENGKLIINIDGNVA LNKESGEVE++NE L+ +
Sbjct: 183 HALLVSLFGDVKAGENGKLIINIDGNVAVLNKESGEVENKNESLERK 323
Score = 33.9 bits (76), Expect(2) = 3e-47
Identities = 12/23 (52%), Positives = 20/23 (86%)
Frame = +2
Query: 668 KERVRTAFRRIQSSVKPIPLSAP 690
+++++ AF+ IQ+S+KPIPLS P
Sbjct: 314 RKKIKXAFQHIQNSIKPIPLSTP 382
>TC221933 weakly similar to UP|Q9VE51 (Q9VE51) CG7698-PA (RE31408p), partial
(7%)
Length = 434
Score = 99.4 bits (246), Expect = 5e-21
Identities = 49/66 (74%), Positives = 51/66 (77%), Gaps = 7/66 (10%)
Frame = +1
Query: 7 RESNGGTINRET-------EDQLIVTPLGAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSG 59
RE N T R EDQLIVTPLGAGNEVGRSCVYM+YKGKTVLFDCGIHP YSG
Sbjct: 235 REMNNCTAKRRENARSSREEDQLIVTPLGAGNEVGRSCVYMSYKGKTVLFDCGIHPAYSG 414
Query: 60 MAALPY 65
MAA+PY
Sbjct: 415 MAAMPY 432
>TC209342
Length = 629
Score = 79.7 bits (195), Expect = 4e-15
Identities = 40/43 (93%), Positives = 41/43 (95%)
Frame = +3
Query: 648 GNVAELNKESGEVESENEGLKERVRTAFRRIQSSVKPIPLSAP 690
GNVA LNKESGEVESENEGLKERVR AF+RIQSSVKPIPLSAP
Sbjct: 3 GNVAVLNKESGEVESENEGLKERVRAAFQRIQSSVKPIPLSAP 131
>BM178689 similar to SP|Q9LKF9|CPSB Cleavage and polyadenylation specificity
factor 100 kDa subunit (CPSF 100 kDa subunit)., partial
(16%)
Length = 419
Score = 35.4 bits (80), Expect = 0.085
Identities = 32/113 (28%), Positives = 53/113 (46%), Gaps = 5/113 (4%)
Frame = +1
Query: 24 VTPL-GAGNEVGRSCVYMTYKGKTVLFDCGIHPGYSGMAALPYFDEIDPSTVDVLLITHF 82
VTPL G NE S + ++ G L DCG + + P ST+D +L++H
Sbjct: 64 VTPLCGVYNENPLSYL-VSIDGFNFLVDCGWNDHFDPSHLQPLARVA--STIDAVLLSHA 234
Query: 83 HLDHAASLPYFLEKTTFKGRVFMTYATKAIYKL----LLSDYVKVSKVSVDDM 131
H +LPY +++ V Y+T+ +Y+L + Y+ +VS D+
Sbjct: 235 DTLHLGALPYAMKRLGLSAPV---YSTEPVYRLGLLTMYDQYLSRKQVSEFDL 384
>TC218017 similar to GB|AAF82809.1|9082326|AF283277 polyadenylation
cleavage/specificity factor 100 kDa subunit {Arabidopsis
thaliana;} , partial (35%)
Length = 1292
Score = 32.3 bits (72), Expect = 0.72
Identities = 19/76 (25%), Positives = 35/76 (46%)
Frame = +2
Query: 362 IPGYVVEGTLAKTILNEPKEVTLMNGLSAPLHMQVHYISFSAHADSAQTSAFLEELNPPN 421
I G + EG + + +P +V + + + + + Y+ F +D L + P
Sbjct: 71 INGKLDEGAASLILDTKPSKV-VSDERTVQVRCSLVYMDFEGRSDGRSIKNILSHVAPLK 247
Query: 422 IILVHGAANEMGRLKQ 437
++LVHG+A LKQ
Sbjct: 248 LVLVHGSAEATEHLKQ 295
>BM307053 similar to GP|9082326|gb|A polyadenylation cleavage/specificity
factor 100 kDa subunit {Arabidopsis thaliana}, partial
(4%)
Length = 379
Score = 32.0 bits (71), Expect = 0.94
Identities = 14/46 (30%), Positives = 23/46 (49%)
Frame = +1
Query: 161 FWCYTAGHVLGAAMFMVDIAGVRVLYTGDYSREEDRHLRAAETPQF 206
F AGH+LG ++ + G V+Y D++ ++RHL F
Sbjct: 241 FGTRVAGHLLGGTIWKITKDGEDVIYAVDFNHRKERHLNGTVLGSF 378
>AW234035
Length = 417
Score = 31.2 bits (69), Expect = 1.6
Identities = 20/71 (28%), Positives = 35/71 (49%)
Frame = +2
Query: 77 LLITHFHLDHAASLPYFLEKTTFKGRVFMTYATKAIYKLLLSDYVKVSKVSVDDMLYDEQ 136
LLITH H+DH LP ++ AT+ +Y+ + + +SV + +
Sbjct: 104 LLITHAHMDHIGGLPMYV-------------ATRGLYR--MKPPTIIVPISVKEDVEKLF 238
Query: 137 DINRSMDKIEV 147
+I+R MD+ E+
Sbjct: 239 EIHRKMDQSEL 271
>TC226005 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), partial (64%)
Length = 792
Score = 30.0 bits (66), Expect = 3.6
Identities = 21/70 (30%), Positives = 31/70 (44%)
Frame = +1
Query: 601 KIVAESDATKIEEENEKKTEKVMQALLNSLFGNVKVGENGKLIINIDGNVAELNKESGEV 660
K E+D KIEEE EKK + + + N + K E + II E K +
Sbjct: 292 KAAVEADLKKIEEELEKKKAEAAEKIKNKIAAIHKEAEERRAII-------EAKKGEDLL 450
Query: 661 ESENEGLKER 670
++E + K R
Sbjct: 451 KAEEQAAKYR 480
>TC216136
Length = 1291
Score = 29.6 bits (65), Expect = 4.7
Identities = 26/122 (21%), Positives = 46/122 (37%), Gaps = 7/122 (5%)
Frame = +3
Query: 500 MAPDDLHVFSQLSTANVTQRITIPYSGAFCVIQSRLKQIYESVEPSVDEESGVPMLLVHD 559
M P L + N+ QR+ +P+ G+ Q LKQ+++ P D
Sbjct: 501 MFPPSLSPLQEERLRNLRQRLEVPFDGSKAEHQDALKQLWKLAYP--------------D 638
Query: 560 RVTVKHESEKHVSLHW-ASDPINDMVSDSVVAL------VLNINRDLPKIVAESDATKIE 612
R +S+ + W SDP D ++L + +++ + D T+ E
Sbjct: 639 RELPSLKSDLWKEMGWQGSDPSTDFRGGGFISLENLIFFAMKYPDSFQRLLHKQDGTRAE 818
Query: 613 EE 614
E
Sbjct: 819 WE 824
>TC226006 similar to UP|REMO_SOLTU (P93788) Remorin (pp34), partial (64%)
Length = 991
Score = 29.3 bits (64), Expect = 6.1
Identities = 21/70 (30%), Positives = 31/70 (44%)
Frame = +2
Query: 601 KIVAESDATKIEEENEKKTEKVMQALLNSLFGNVKVGENGKLIINIDGNVAELNKESGEV 660
K E+D KIEEE EKK + + + N + K E + II E K +
Sbjct: 398 KAAVEADLKKIEEELEKKKAEAAEKIKNKIATIHKEAEERRAII-------EAKKGEDLL 556
Query: 661 ESENEGLKER 670
++E + K R
Sbjct: 557 KAEEQAAKYR 586
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.134 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,986,353
Number of Sequences: 63676
Number of extensions: 330737
Number of successful extensions: 1566
Number of sequences better than 10.0: 24
Number of HSP's better than 10.0 without gapping: 1560
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1566
length of query: 690
length of database: 12,639,632
effective HSP length: 104
effective length of query: 586
effective length of database: 6,017,328
effective search space: 3526154208
effective search space used: 3526154208
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 62 (28.5 bits)
Medicago: description of AC134322.16


