

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC134049.2 - phase: 0
(170 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
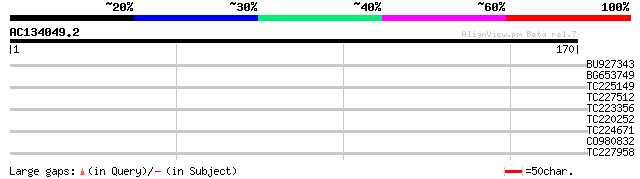
Score E
Sequences producing significant alignments: (bits) Value
BU927343 32 0.15
BG653749 28 1.6
TC225149 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S... 28 2.1
TC227512 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {A... 28 2.8
TC223356 similar to GB|AAO44032.1|28466847|BT004766 At5g47710 {A... 27 4.7
TC220252 weakly similar to GB|AAR24736.1|38603986|BT010958 At4g3... 27 4.7
TC224671 similar to UP|Q8S8Z0 (Q8S8Z0) Protein phosphatase 2C, p... 27 6.2
CO980832 26 8.1
TC227958 similar to UP|O48721 (O48721) Expressed protein, partia... 26 8.1
>BU927343
Length = 451
Score = 32.0 bits (71), Expect = 0.15
Identities = 16/47 (34%), Positives = 28/47 (59%)
Frame = -3
Query: 34 AEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEECSPSWN 80
+ ++++RG E+N Q G S AT +L CR R+ + ++E +P N
Sbjct: 179 SSQSSRRGLELNPQPGLSNAATVQLL---CRARDPITCLQEKTPPPN 48
>BG653749
Length = 417
Score = 28.5 bits (62), Expect = 1.6
Identities = 11/19 (57%), Positives = 12/19 (62%)
Frame = +2
Query: 38 AKRGFEINNQDGWSQHATC 56
+KR F I NQ W QH TC
Sbjct: 161 SKRSFPIPNQRTWEQHRTC 217
>TC225149 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S16
(Fragment), partial (97%)
Length = 768
Score = 28.1 bits (61), Expect = 2.1
Identities = 13/34 (38%), Positives = 17/34 (49%)
Frame = -3
Query: 67 EAVEFMEECSPSWNSFLSFMLTHNWWHVALCYLE 100
EA+ ++ C P L L H W+ V LC LE
Sbjct: 130 EALHLLQRCLPVLRLRLRLRLRHCWFVVLLCELE 29
>TC227512 similar to GB|AAQ89615.1|37202000|BT010593 At3g17205 {Arabidopsis
thaliana;} , partial (17%)
Length = 541
Score = 27.7 bits (60), Expect = 2.8
Identities = 12/23 (52%), Positives = 14/23 (60%)
Frame = -1
Query: 38 AKRGFEINNQDGWSQHATCHVLQ 60
AK+ F+I WS ATCH LQ
Sbjct: 484 AKQRFKILESKQWSPRATCHKLQ 416
>TC223356 similar to GB|AAO44032.1|28466847|BT004766 At5g47710 {Arabidopsis
thaliana;} , partial (96%)
Length = 705
Score = 26.9 bits (58), Expect = 4.7
Identities = 23/74 (31%), Positives = 33/74 (44%), Gaps = 1/74 (1%)
Frame = +3
Query: 76 SPSWNSFLSFMLTHNWWHVALCYLEGNAPMQRV-LEVYDNYIWKELDKTDATVPEVYLNA 134
+P WN L+F LT P+ + LEV+D +WK DK + YLN
Sbjct: 210 NPVWNEELNFTLTE--------------PLGVLNLEVFDKDLWKADDK----MGNSYLNL 335
Query: 135 VALLLRLCVRDELE 148
L+ +RD L+
Sbjct: 336 QPLISAARLRDILK 377
>TC220252 weakly similar to GB|AAR24736.1|38603986|BT010958 At4g32870
{Arabidopsis thaliana;} , partial (41%)
Length = 867
Score = 26.9 bits (58), Expect = 4.7
Identities = 10/24 (41%), Positives = 12/24 (49%), Gaps = 1/24 (4%)
Frame = +2
Query: 79 WNSFLSFMLTHNWWHVALCY-LEG 101
W + F H WW + CY LEG
Sbjct: 131 WTALEDFCNLHKWWPIETCYQLEG 202
>TC224671 similar to UP|Q8S8Z0 (Q8S8Z0) Protein phosphatase 2C, partial (87%)
Length = 1803
Score = 26.6 bits (57), Expect = 6.2
Identities = 13/31 (41%), Positives = 21/31 (66%)
Frame = +3
Query: 31 MKEAEEAAKRGFEINNQDGWSQHATCHVLQY 61
+++AEEAAKR + Q G S + TC V+++
Sbjct: 1239 IEDAEEAAKRLMQEAYQRGSSDNITCVVVRF 1331
>CO980832
Length = 838
Score = 26.2 bits (56), Expect = 8.1
Identities = 11/32 (34%), Positives = 17/32 (52%)
Frame = +3
Query: 59 LQYECRFREAVEFMEECSPSWNSFLSFMLTHN 90
LQ + + E +F C +WN+ SF+ HN
Sbjct: 288 LQLQTKI*EVTKFKNLCKFNWNNCSSFIDNHN 383
>TC227958 similar to UP|O48721 (O48721) Expressed protein, partial (19%)
Length = 704
Score = 26.2 bits (56), Expect = 8.1
Identities = 22/91 (24%), Positives = 32/91 (34%), Gaps = 4/91 (4%)
Frame = +3
Query: 23 FPLLELGQMKEAEEAAKRGFEINNQDGWSQHATCHVLQYECRFREAVEFMEECSPS---- 78
+P + E + RGFE+ + + HV + A + SPS
Sbjct: 60 YPXSAKASNEXEEXXSNRGFEVTRMNTYXTVPVQHVGHTVVKIALAAPVVTVASPSSIRA 239
Query: 79 WNSFLSFMLTHNWWHVALCYLEGNAPMQRVL 109
W + LS W + C E A M R L
Sbjct: 240 WKNLLS---DSEWSNSVACIGETTAAMARSL 323
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.137 0.436
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,472,539
Number of Sequences: 63676
Number of extensions: 124105
Number of successful extensions: 631
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 629
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 631
length of query: 170
length of database: 12,639,632
effective HSP length: 90
effective length of query: 80
effective length of database: 6,908,792
effective search space: 552703360
effective search space used: 552703360
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 55 (25.8 bits)
Medicago: description of AC134049.2


