

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133780.8 - phase: 0 /pseudo
(287 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
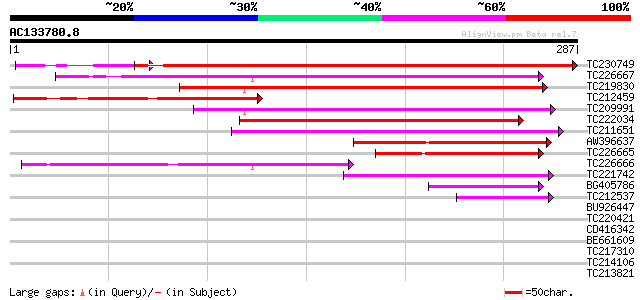
Score E
Sequences producing significant alignments: (bits) Value
TC230749 similar to UP|Q9LYV6 (Q9LYV6) ABA-responsive protein-li... 376 e-105
TC226667 similar to UP|Q9SE96 (Q9SE96) FH protein interacting pr... 202 2e-52
TC219830 weakly similar to UP|Q8S8F8 (Q8S8F8) Expressed protein,... 160 8e-40
TC212459 157 7e-39
TC209991 132 2e-31
TC222034 weakly similar to UP|Q6H5X3 (Q6H5X3) ABA-responsive pro... 119 2e-27
TC211651 similar to UP|Q9FMW4 (Q9FMW4) Similarity to ABA-respons... 119 2e-27
AW396637 similar to GP|20197888|gb Expressed protein {Arabidopsi... 110 7e-25
TC226665 106 1e-23
TC226666 similar to UP|Q9SE96 (Q9SE96) FH protein interacting pr... 85 3e-17
TC221742 weakly similar to UP|Q6H5X2 (Q6H5X2) ABA-responsive pro... 83 2e-16
BG405786 47 1e-05
TC212537 weakly similar to UP|Q9FMW6 (Q9FMW6) Similarity to ABA-... 47 1e-05
BU926447 similar to GP|8453100|gb|A short-root protein {Arabidop... 36 0.023
TC220421 weakly similar to UP|GP1_CHLRE (Q9FPQ6) Vegetative cell... 35 0.030
CD416342 similar to SP|P42698|D111_ DNA-damage-repair/toleration... 35 0.039
BE661609 similar to SP|Q9ZPY7|CSE1 Importin-alpha re-exporter (C... 35 0.051
TC217310 similar to UP|Q9SWF9 (Q9SWF9) Zinc finger protein, part... 35 0.051
TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 pr... 34 0.067
TC213821 weakly similar to UP|O81373 (O81373) Cotton fiber expre... 34 0.067
>TC230749 similar to UP|Q9LYV6 (Q9LYV6) ABA-responsive protein-like
(At5g13200), partial (58%)
Length = 972
Score = 376 bits (965), Expect = e-105
Identities = 182/225 (80%), Positives = 202/225 (88%), Gaps = 1/225 (0%)
Frame = +3
Query: 64 KKAALQTASADGQGQPQVHYYQQDH-PYVQHSPVDKPSSSPMESILHMFDSWSKKAEATA 122
KKAALQ QP +Y+ Q H PYVQHS +DKPS+SPMESIL+MFDSWS+KAEATA
Sbjct: 147 KKAALQP-------QPVQYYHDQHHHPYVQHSTLDKPSNSPMESILNMFDSWSRKAEATA 305
Query: 123 NNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTT 182
+N+WHNLKTGPSVSSAA+GKMNLTVKAISEGGFESLYKQ FTTYPNEKLKK+FACYLST+
Sbjct: 306 HNVWHNLKTGPSVSSAALGKMNLTVKAISEGGFESLYKQTFTTYPNEKLKKSFACYLSTS 485
Query: 183 TGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSE 242
TGPVAGTLYLS+IH+AFCSDRPL FTAPSGQ TW+YYKVMVPLGK+G VNPV MREN SE
Sbjct: 486 TGPVAGTLYLSNIHVAFCSDRPLCFTAPSGQETWTYYKVMVPLGKVGVVNPVTMRENPSE 665
Query: 243 RYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGISHFVVSGVAAPST 287
+YIQ+VTV+GHDFWFMGFVN+DKAVKN+SEGISHFV GVA PST
Sbjct: 666 KYIQVVTVEGHDFWFMGFVNFDKAVKNISEGISHFVAPGVAVPST 800
Score = 61.2 bits (147), Expect = 5e-10
Identities = 30/70 (42%), Positives = 41/70 (57%)
Frame = +2
Query: 4 NNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDN 63
N N+N +NQS P+ + +PP TENWGTH+MG PA+PSSHPDN
Sbjct: 20 NTNSNKKNQSFPEAS-----SPPKVG-------------TENWGTHIMGTPAIPSSHPDN 145
Query: 64 KKAALQTASA 73
KK + T+++
Sbjct: 146KKGSFTTSTS 175
>TC226667 similar to UP|Q9SE96 (Q9SE96) FH protein interacting protein FIP1
(At1g28200/F3H9_12), partial (83%)
Length = 1252
Score = 202 bits (513), Expect = 2e-52
Identities = 111/256 (43%), Positives = 151/256 (58%), Gaps = 9/256 (3%)
Frame = +2
Query: 24 TPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADGQGQPQVHY 83
T P S S PQ H T ++ + P P + L T S +
Sbjct: 191 TKPEESHSQSQSQPQPH--TADYAPY-------PKLDPTDVAPPLNTESRAPISEDAATT 343
Query: 84 YQQD-HPYVQHSPVDKPSS-SPMESILHMFDSWSKKA-EAT------ANNIWHNLKTGPS 134
+D +PYV +PV S+ + ++S+ + W KKA EAT A N+W +LKTGPS
Sbjct: 344 MPKDSNPYVTPAPVPASSTKTTLDSVKDVLGKWGKKAAEATKKAEDLAGNMWQHLKTGPS 523
Query: 135 VSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSD 194
+ AA+G++ K ++EGG+E +++Q F T P E+L KT+ACYLST+ GPV G LYLS
Sbjct: 524 FADAAVGRIAQGTKVLAEGGYEKIFRQTFETVPEEQLLKTYACYLSTSAGPVMGVLYLST 703
Query: 195 IHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHD 254
LAFCSD PLS+ Q WSYYKV++PL ++ VN R N SE+YIQI++VD H+
Sbjct: 704 AKLAFCSDNPLSYQVGGDQTQWSYYKVVIPLHQLRAVNASTSRTNQSEKYIQIISVDNHE 883
Query: 255 FWFMGFVNYDKAVKNL 270
FWFMGFV+YD AVKN+
Sbjct: 884 FWFMGFVHYDSAVKNI 931
>TC219830 weakly similar to UP|Q8S8F8 (Q8S8F8) Expressed protein, partial
(28%)
Length = 909
Score = 160 bits (404), Expect = 8e-40
Identities = 83/194 (42%), Positives = 128/194 (65%), Gaps = 8/194 (4%)
Frame = +3
Query: 87 DHPYVQHSPVDKPSSSPMESILHMFDSWSKK-------AEATANNIWHNLKTGPSVSSAA 139
++PYVQ SP+ PM+++ + S+K AE A+N W++++ G S++ AA
Sbjct: 111 NNPYVQISPLHTNRPKPMDTVCDALNRCSRKVGKATRRAETMADNFWNHIRIGSSLADAA 290
Query: 140 MGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAF 199
+ ++ K ++ GG + L++Q F +P EKL K+FACYLST+TGPV GT+Y+S +AF
Sbjct: 291 VARIVQGTKVLTLGGPDILFQQSFGNFPGEKLIKSFACYLSTSTGPVIGTIYVSTKRVAF 470
Query: 200 CSDRPLSFTAPSGQVTWS-YYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFM 258
CSD PL S Q S +YKV++ L ++ TV+P R N +E+Y+Q+VTVDG++F+FM
Sbjct: 471 CSDYPLCNYPLSLQQNQSVHYKVVLQLDQLSTVSPFSNRFNPAEKYMQLVTVDGYEFYFM 650
Query: 259 GFVNYDKAVKNLSE 272
GF+ YDKA+K + E
Sbjct: 651 GFIAYDKALKTVRE 692
>TC212459
Length = 395
Score = 157 bits (396), Expect = 7e-39
Identities = 82/127 (64%), Positives = 94/127 (73%), Gaps = 1/127 (0%)
Frame = +2
Query: 3 NNNNNNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPD 62
N NN+ +NQS P+ SSSSSS P + TENWGTH+MG PAVPSSHPD
Sbjct: 29 NTNNDTEKNQSFPEAQG------ASSSSSSPP-----NGGTENWGTHIMGTPAVPSSHPD 175
Query: 63 NKKAALQTASADGQGQPQVHYYQQ-DHPYVQHSPVDKPSSSPMESILHMFDSWSKKAEAT 121
NKKAALQ+ GQ QP +Y+ Q HPYVQHSPVDKPS+SPMESIL+MFDSWSKKAEAT
Sbjct: 176 NKKAALQS----GQPQPVQYYHDQHQHPYVQHSPVDKPSNSPMESILNMFDSWSKKAEAT 343
Query: 122 ANNIWHN 128
A+N+WHN
Sbjct: 344 AHNVWHN 364
>TC209991
Length = 793
Score = 132 bits (331), Expect = 2e-31
Identities = 73/191 (38%), Positives = 111/191 (57%), Gaps = 8/191 (4%)
Frame = +3
Query: 94 SPVDKPSSSPMESILHMFDSWSKK-------AEATANNIWHNLKTGPSVSSAAMGKMNLT 146
SP + +PM+ I + + KK AE NI ++L+ + AA+ ++
Sbjct: 138 SPAEAKRPNPMDRIYGAINHYGKKVEEATKQAETMVGNIRNHLRVSSRPADAAIARLIQG 317
Query: 147 VKAISEGGFESLYKQIFTTYPNEKLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLS 206
K ++ GG + L++Q F +P EKL + ACY+ST +GP+ GTLY+S LAFCSD PL
Sbjct: 318 TKVLTSGGPDKLFQQTFGVFPGEKLLQPCACYISTNSGPLIGTLYISTKRLAFCSDYPLC 497
Query: 207 FTAPS-GQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDK 265
S Q YYKV+V L ++ V+ V N SE+ +Q++T DG++F FMGF +YDK
Sbjct: 498 HHPFSLQQHECVYYKVIVLLDQLSNVSSVTNGLNPSEKRMQVITTDGYEFNFMGFFSYDK 677
Query: 266 AVKNLSEGISH 276
A+K ++E + H
Sbjct: 678 ALKAVNEALKH 710
>TC222034 weakly similar to UP|Q6H5X3 (Q6H5X3) ABA-responsive protein-like,
partial (47%)
Length = 557
Score = 119 bits (298), Expect = 2e-27
Identities = 56/144 (38%), Positives = 92/144 (63%)
Frame = +1
Query: 117 KAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKTFA 176
K+ + + I ++K GP++S GK++L + I EGG S++K +F E+L K
Sbjct: 124 KSRSFTHRIHDHVKMGPNLSEILKGKLSLGARIIQEGGRGSIFKSVFGMQEKEQLLKASQ 303
Query: 177 CYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIM 236
CYL TT GP+AG L++S +AF S+RP++F++ +G++ + YKV++P+G+I VN
Sbjct: 304 CYLYTTAGPIAGILFVSTEKVAFYSERPITFSSATGELVRAPYKVLIPIGRIKEVNESQN 483
Query: 237 RENHSERYIQIVTVDGHDFWFMGF 260
++YI+IVT D +FWF+GF
Sbjct: 484 VNKAEQKYIEIVTEDDSEFWFVGF 555
>TC211651 similar to UP|Q9FMW4 (Q9FMW4) Similarity to ABA-responsive protein,
partial (53%)
Length = 799
Score = 119 bits (297), Expect = 2e-27
Identities = 61/168 (36%), Positives = 96/168 (56%)
Frame = +1
Query: 113 SWSKKAEATANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLK 172
+WS + + N+ ++ ++S K++L + + GG E ++KQ F+ E+L
Sbjct: 148 TWSCHSMSIFNHNIKVMRLRTNISETVKRKISLGARILRVGGVEKVFKQFFSMEEGERLL 327
Query: 173 KTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVN 232
K CYLSTT+GP+AG L++S +AFCS+R + G + YKV++PL KI VN
Sbjct: 328 KVSQCYLSTTSGPLAGFLFISTDKVAFCSERSMKVFTRKGHMLRIRYKVVIPLKKIKCVN 507
Query: 233 PVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGISHFVVS 280
+N +++YI+IVT D DFWFMG + Y K K L + +S +S
Sbjct: 508 QSQNIQNPTQKYIEIVTEDNFDFWFMGVLKYQKTFKYLEQAVSQA*IS 651
>AW396637 similar to GP|20197888|gb Expressed protein {Arabidopsis thaliana},
partial (33%)
Length = 469
Score = 110 bits (275), Expect = 7e-25
Identities = 49/100 (49%), Positives = 72/100 (72%)
Frame = +1
Query: 175 FACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPV 234
+ACYLS + GPV G LY+S +A+ SD P+S+ + Q WSYYKV++PL ++ +VNP
Sbjct: 1 YACYLSMSAGPVMGVLYVSTAKIAYSSDNPISYKNDN-QTEWSYYKVVIPLLELKSVNPS 177
Query: 235 IMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGI 274
N +E+YIQ+++VD H+FWFMGF+NY+ AV++L I
Sbjct: 178 SNTSNPAEKYIQVISVDNHEFWFMGFLNYEGAVESLQGAI 297
>TC226665
Length = 562
Score = 106 bits (264), Expect = 1e-23
Identities = 49/85 (57%), Positives = 61/85 (71%)
Frame = +3
Query: 186 VAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIGTVNPVIMRENHSERYI 245
V G LYLS LAFCSD PLS+ Q WSYYKV++PL ++ VN + N SE+YI
Sbjct: 3 VMGVLYLSTAKLAFCSDNPLSYQV-GDQTQWSYYKVVIPLHQLRAVNASTSKTNQSEKYI 179
Query: 246 QIVTVDGHDFWFMGFVNYDKAVKNL 270
QI++VD H+FWFMGFV+YD AVKN+
Sbjct: 180 QIISVDNHEFWFMGFVHYDSAVKNI 254
>TC226666 similar to UP|Q9SE96 (Q9SE96) FH protein interacting protein FIP1
(At1g28200/F3H9_12), partial (44%)
Length = 655
Score = 85.1 bits (209), Expect = 3e-17
Identities = 55/176 (31%), Positives = 86/176 (48%), Gaps = 8/176 (4%)
Frame = +1
Query: 7 NNNQNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKA 66
NNN + +P + A+ T P S S PQ H+ + P P N ++
Sbjct: 145 NNNMSADAPPQTQ-ATNTKPEESQSQSQSQPQRHSGDYAPYPKLDPTDVAPPQQPLNTES 321
Query: 67 ALQTASADGQGQPQVHYYQQDHPYVQHSPVDKPSS-SPMESILHMFDSWSKKA-EAT--- 121
+ P+ +PYV +PV S+ + ++S+ + W KKA EAT
Sbjct: 322 RAPISEDAATTMPK-----DSNPYVTPAPVTASSTKTTLDSVKDVLGKWGKKAAEATKKA 486
Query: 122 ---ANNIWHNLKTGPSVSSAAMGKMNLTVKAISEGGFESLYKQIFTTYPNEKLKKT 174
A N+W +LKTGPS + AA+G++ K ++EGG+E +++Q F T P E+L KT
Sbjct: 487 EDLAGNMWQHLKTGPSFADAAVGRIAQGTKVLAEGGYEKIFRQTFETVPEEQLLKT 654
>TC221742 weakly similar to UP|Q6H5X2 (Q6H5X2) ABA-responsive protein-like,
partial (35%)
Length = 714
Score = 82.8 bits (203), Expect = 2e-16
Identities = 42/106 (39%), Positives = 60/106 (55%)
Frame = +2
Query: 170 KLKKTFACYLSTTTGPVAGTLYLSDIHLAFCSDRPLSFTAPSGQVTWSYYKVMVPLGKIG 229
+L K CYLSTT+ P+A L++S +A S+R + G + YKV +PL KI
Sbjct: 2 RLLKVSQCYLSTTSXPLAXXLFISTDKVAXXSERSMKVFTQKGHMLRIRYKVTIPLKKIK 181
Query: 230 TVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
VN + +++YI+IVT D DFWFMG + Y K K L + +S
Sbjct: 182 CVNQSANVQKPTQKYIEIVTEDNFDFWFMGVLKYQKTFKYLEQAVS 319
>BG405786
Length = 201
Score = 47.0 bits (110), Expect = 1e-05
Identities = 24/59 (40%), Positives = 34/59 (56%), Gaps = 1/59 (1%)
Frame = +3
Query: 213 QVTWSYYKVMVPLGKIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGF-VNYDKAVKNL 270
Q WSYYK +PL + VN + S ++I I++VD FW +G +YD AVKN+
Sbjct: 12 QTQWSYYKGGIPLHPLRAVNRSNKGTHPSGKFIPIISVDTMKFWVIGLGAHYDSAVKNI 188
>TC212537 weakly similar to UP|Q9FMW6 (Q9FMW6) Similarity to ABA-repsonsive
protein, partial (15%)
Length = 657
Score = 46.6 bits (109), Expect = 1e-05
Identities = 20/49 (40%), Positives = 29/49 (58%)
Frame = +3
Query: 227 KIGTVNPVIMRENHSERYIQIVTVDGHDFWFMGFVNYDKAVKNLSEGIS 275
K+ VN + +++YI+IVT D DFWFMG + Y K K L + +S
Sbjct: 3 KVKCVNQSANAQKPTQKYIEIVTEDNFDFWFMGVLKYQKTFKYLEQAVS 149
>BU926447 similar to GP|8453100|gb|A short-root protein {Arabidopsis
thaliana}, partial (7%)
Length = 450
Score = 35.8 bits (81), Expect = 0.023
Identities = 24/70 (34%), Positives = 32/70 (45%), Gaps = 13/70 (18%)
Frame = +3
Query: 1 MNNNNNNNN-------QNQSSPDENPFA-SPTPP-----SSSSSSVPLPPQNHATTENWG 47
+ NNNNNNN Q S+P P +P+PP +S+S+S P P TT +W
Sbjct: 219 ITNNNNNNNTTYPTFLQTPSAPPPTPITPTPSPPPHKFKTSTSNSPPTGP----TTSSWN 386
Query: 48 THMMGAPAVP 57
H P
Sbjct: 387 PHAPSPTTTP 416
Score = 28.1 bits (61), Expect = 4.8
Identities = 15/44 (34%), Positives = 25/44 (56%), Gaps = 2/44 (4%)
Frame = +3
Query: 5 NNNNNQNQSSP-DENPFASPTPPSSSSSSV-PLPPQNHATTENW 46
NNNN S+P P A+P PP +++ + P+ + +A+T W
Sbjct: 33 NNNNPIPPSTPRPAEPPAAPDPPDKTTTLILPIKKKKNASTFTW 164
>TC220421 weakly similar to UP|GP1_CHLRE (Q9FPQ6) Vegetative cell wall
protein gp1 precursor (Hydroxyproline-rich glycoprotein
1), partial (13%)
Length = 1409
Score = 35.4 bits (80), Expect = 0.030
Identities = 25/90 (27%), Positives = 34/90 (37%)
Frame = +3
Query: 13 SSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTAS 72
SSP +P A PT PS S S P PP + + + +P S A+ +A+
Sbjct: 294 SSPSASPTAPPTSPSPSPPSSPSPPSAPSPSSTAPSPPSASPPTSPSPSPPSPASTASAN 473
Query: 73 ADGQGQPQVHYYQQDHPYVQHSPVDKPSSS 102
+ G P S PSSS
Sbjct: 474 SPASGSPTSRTSPTSPSPTSTSKPPAPSSS 563
>CD416342 similar to SP|P42698|D111_ DNA-damage-repair/toleration protein
DRT111 chloroplast precursor. [Mouse-ear cress],
partial (24%)
Length = 624
Score = 35.0 bits (79), Expect = 0.039
Identities = 19/61 (31%), Positives = 26/61 (42%)
Frame = -1
Query: 13 SSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTAS 72
S P PF TPPSS S PP + +TT T A S+ P + +++
Sbjct: 216 SPPPPPPFPRKTPPSSPPSLASSPPSSRSTTRPGPTTTRNTAATSSAXPARPRCCASSSA 37
Query: 73 A 73
A
Sbjct: 36 A 34
>BE661609 similar to SP|Q9ZPY7|CSE1 Importin-alpha re-exporter (Cellular
apoptosis susceptibility protein homolog). [Mouse-ear
cress], partial (14%)
Length = 742
Score = 34.7 bits (78), Expect = 0.051
Identities = 29/103 (28%), Positives = 41/103 (39%)
Frame = +1
Query: 13 SSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTAS 72
++P + SP+PPS++ S+ PLP + T G P P S + + +S
Sbjct: 154 TTPSPSSAWSPSPPSTTRSARPLPSTSRTTXA-----XAGPPRTPPSPTPRRTRSKP*SS 318
Query: 73 ADGQGQPQVHYYQQDHPYVQHSPVDKPSSSPMESILHMFDSWS 115
PQ Q P SP P S P+ S H S S
Sbjct: 319 PHALRLPQNPISAQRSPCPNRSP-RFPKSGPLSSRTHRKSSKS 444
>TC217310 similar to UP|Q9SWF9 (Q9SWF9) Zinc finger protein, partial (93%)
Length = 1606
Score = 34.7 bits (78), Expect = 0.051
Identities = 28/92 (30%), Positives = 41/92 (44%), Gaps = 8/92 (8%)
Frame = +1
Query: 1 MNNNNNNNNQNQSSPDENPFAS--PTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAV-P 57
M NNNNN+N +P ++ + T + S S P P + T + A
Sbjct: 64 MENNNNNHNLIVPNPQDSLWMMNLRTGETMDSGSYPERPGEPDCSYYMRTGLCRFGATCR 243
Query: 58 SSHPDNKKAALQTASADGQ-----GQPQVHYY 84
+HP N+K A+ TA G+ GQP+ YY
Sbjct: 244 FNHPPNRKLAIATARMIGEFPERIGQPECQYY 339
>TC214106 weakly similar to UP|Q8L685 (Q8L685) Pherophorin-dz1 protein
precursor, partial (22%)
Length = 1296
Score = 34.3 bits (77), Expect = 0.067
Identities = 25/85 (29%), Positives = 31/85 (36%)
Frame = +1
Query: 19 PFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSSHPDNKKAALQTASADGQGQ 78
P SP PPS PP + T+ + +P PS P + +
Sbjct: 7 PPPSPPPPSPPPPVYSPPPPPPSPPPPSPTYCVRSPPPPSPPPPSPPPPPPPVFSP---P 177
Query: 79 PQVHYYQQDHPYVQHSPVDKPSSSP 103
P V YY P QHSP P SP
Sbjct: 178 PPVQYYYSSPPPPQHSPPPPPPHSP 252
>TC213821 weakly similar to UP|O81373 (O81373) Cotton fiber expressed protein
1, partial (16%)
Length = 444
Score = 34.3 bits (77), Expect = 0.067
Identities = 23/66 (34%), Positives = 31/66 (46%), Gaps = 3/66 (4%)
Frame = +1
Query: 10 QNQSSPDENPFASPTPPSSSSSSVPLPPQNHATTENWGTHMMGAPAVPSS---HPDNKKA 66
Q SSP P S + +SSSS LPP + TT +P PSS HP ++
Sbjct: 202 QPSSSPASLPLTSTSFSTSSSSPSSLPPNSTTTTTLRRIPPSSSPPTPSSTPYHPPQSRS 381
Query: 67 ALQTAS 72
+L+ S
Sbjct: 382 SLRKPS 399
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.312 0.127 0.383
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,156,452
Number of Sequences: 63676
Number of extensions: 257899
Number of successful extensions: 4780
Number of sequences better than 10.0: 289
Number of HSP's better than 10.0 without gapping: 3156
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4028
length of query: 287
length of database: 12,639,632
effective HSP length: 96
effective length of query: 191
effective length of database: 6,526,736
effective search space: 1246606576
effective search space used: 1246606576
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 58 (26.9 bits)
Medicago: description of AC133780.8


