

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
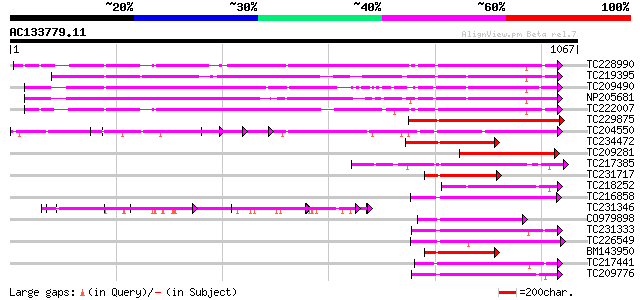
Query= AC133779.11 - phase: 0
(1067 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC228990 UP|Q9LKZ6 (Q9LKZ6) Receptor-like protein kinase 1, comp... 460 e-129
TC219395 UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kinase 3, part... 456 e-128
TC209490 UP|Q8GSN9 (Q8GSN9) LRR receptor-like kinase (Fragment),... 440 e-123
NP205681 receptor protein kinase-like protein 439 e-123
TC222007 UP|Q9LKZ5 (Q9LKZ5) Receptor-like protein kinase 2, comp... 433 e-121
TC229875 weakly similar to UP|Q75WU3 (Q75WU3) Leucine-rich repea... 369 e-102
TC204550 UP|Q8L3Y5 (Q8L3Y5) Receptor-like kinase RHG1, complete 275 6e-74
TC234472 weakly similar to UP|Q75WU3 (Q75WU3) Leucine-rich repea... 255 6e-68
TC209281 similar to UP|Q75WU3 (Q75WU3) Leucine-rich repeat recep... 235 9e-62
TC217385 similar to UP|Q6ZGC7 (Q6ZGC7) Receptor protein kinase P... 204 2e-52
TC231717 weakly similar to UP|Q8VZG8 (Q8VZG8) AT4g08850/T32A17_1... 198 1e-50
TC218252 weakly similar to UP|Q03407 (Q03407) Ydr490cp, partial ... 185 8e-47
TC216858 homologue to UP|Q76FZ8 (Q76FZ8) Brassinosteroid recepto... 183 3e-46
TC231346 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2,... 175 8e-44
CO979898 173 4e-43
TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, parti... 170 4e-42
TC226549 UP|Q84P43 (Q84P43) Protein kinase Pti1, complete 168 1e-41
BM143950 166 4e-41
TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic em... 164 2e-40
TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-l... 163 4e-40
>TC228990 UP|Q9LKZ6 (Q9LKZ6) Receptor-like protein kinase 1, complete
Length = 3346
Score = 460 bits (1183), Expect = e-129
Identities = 334/1045 (31%), Positives = 510/1045 (47%), Gaps = 12/1045 (1%)
Frame = +1
Query: 7 LIMILCVLPTLSVAEDSEAKLALLKWK-DSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDK 65
L++ L +L A SE + ALL +K S D LS+W ++T C W G+ CD
Sbjct: 109 LVLFFLFLHSLQAARISEYR-ALLSFKASSLTDDPTHALSSWNSSTPFCS--WFGLTCDS 279
Query: 66 SNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKN 125
++++ L +L L GTL + +L +S L+ +
Sbjct: 280 RRHVTSLNLTSLSLSGTLSD-------------------------DLSHLPFLSHLSLAD 384
Query: 126 NYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEI 185
N F G IP L+ L+FL++S N P + L NL L L NN +G +P +
Sbjct: 385 NKFSGPIPASFSALSALRFLNLSNNVFNATFPSQLNRLANLEVLDLYNNNMTG-ELPLSV 561
Query: 186 GKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLS 245
+ L HL + + G IP E G +L Y+ LS N L+G I +GNLS L L +
Sbjct: 562 AAMPLLRHLHLGGNFFSGQIPPEYGTWQHLQYLALSGNELAGTIAPELGNLSSLRELYIG 741
Query: 246 NNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPST 305
Y++ SG IP I NL NL L LSG IP+
Sbjct: 742 ----------------------YYNTY--SGGIPPEIGNLSNLVRLDAAYCGLSGEIPAE 849
Query: 306 IGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEV 365
+G L+NL L+L N LSG + +G+L +L+ + + N L+G +PAS LK LT+ +
Sbjct: 850 LGKLQNLDTLFLQVNALSGSLTPELGSLKSLKSMDLSNNMLSGEVPASFAELKNLTLLNL 1029
Query: 366 ATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTS 425
NKLHG IP +FVG LP+ L +L N FTG IP +
Sbjct: 1030 FRNKLHGAIP-----------------EFVGELPA-------LEVLQLWENNFTGSIPQN 1137
Query: 426 LKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFII 485
L + + L N+I G + + +LQ L N G I + GK +L +
Sbjct: 1138 LGNNGRLTLVDLSSNKITGTLPPNMCYGNRLQTLITLGNYLFGPIPDSLGKCKSLNRIRM 1317
Query: 486 SNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKS-LFDLKISNNHFSDNIP 544
N ++G IP GL KL + L N LTG+ P + G + + L + +SNN S ++P
Sbjct: 1318 GENFLNGSIPKGLFGLPKLTQVELQDNLLTGQFPED--GSIATDLGQISLSNNQLSGSLP 1491
Query: 545 SEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDSG--LESL 602
S IG +Q+L L GNE +G+IP ++ L L ++ S NK G I + L +
Sbjct: 1492 STIGNFTSMQKLLLNGNEFTGRIPPQIGMLQQLSKIDFSHNKFSGPIAPEISKCKLLTFI 1671
Query: 603 DLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFG--RNLVFVNISDNQLEGPLP 660
DLSGN L G IP + + L+ LNLS N L G+IP N ++L V+ S N G +P
Sbjct: 1672 DLSGNELSGEIPNKITSMRILNYLNLSRNHLDGSIPGNIASMQSLTSVDFSYNNFSGLVP 1851
Query: 661 KIPAFLSASFESLKNNNHLCGNIRGLDPCATSHSRKRKNVLRPVFIALGAVILVLCVVGA 720
F ++ S N LCG G ++ ++ +V P +L ++++ +V +
Sbjct: 1852 GTGQFGYFNYTSFLGNPELCGPYLGPCKDGVANGPRQPHVKGPFSSSLKLLLVIGLLVCS 2031
Query: 721 LMYIMCGRKKPNEESQTEEVQRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGN 780
+++ + K + E + L + D +++++ + ++G G G
Sbjct: 2032 ILFAVAAIFKARALKKASEARAWKLTAFQRLD--FTVDDVLDC---LKEDNIIGKGGAGI 2196
Query: 781 VYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKF 840
VYK + G VAVK+L ++ F +EI+TL I+HR+I++L GFCS+ +
Sbjct: 2197 VYKGAMPNGDNVAVKRLPAMS---RGSSHDHGFNAEIQTLGRIRHRHIVRLLGFCSNHET 2367
Query: 841 SFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISS 900
+ LVY+++ GSL ++L+ + W+ R + A L YLHHDCSP I+HRD+ S
Sbjct: 2368 NLLVYEYMPNGSLGEVLHGK-KGGHLHWDTRYKIAVEAAKGLCYLHHDCSPLIVHRDVKS 2544
Query: 901 KNVLLNLDYEAHVSDFGTAKFLKPGLHS--WTQFAGTFGYAAPELAQTMEVNEKCDVYSF 958
N+LL+ ++EAHV+DFG AKFL+ S + AG++GY APE A T++V+EK DVYSF
Sbjct: 2545 NNILLDSNFEAHVADFGLAKFLQDSGASECMSAIAGSYGYIAPEYAYTLKVDEKSDVYSF 2724
Query: 959 GVLALETIMGKHP----GDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILI 1014
GV+ LE + G+ P GD + + +N + VLD R V P+ EV+ +
Sbjct: 2725 GVVLLELVTGRKPVGEFGDGVDIVQWVRKMTDSNKEGVLKVLDSRLPSV--PL-HEVMHV 2895
Query: 1015 ARLAFACLSQNPRLRPSMGQVCKML 1039
+A C+ + RP+M +V ++L
Sbjct: 2896 FYVAMLCVEEQAVERPTMREVVQIL 2970
>TC219395 UP|Q9LKZ4 (Q9LKZ4) Receptor-like protein kinase 3, partial (92%)
Length = 3037
Score = 456 bits (1173), Expect = e-128
Identities = 322/971 (33%), Positives = 478/971 (49%), Gaps = 10/971 (1%)
Frame = +3
Query: 79 LKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCT 138
L G L S + P L + + +N F G IP + LS + L NN F+ + P E+
Sbjct: 3 LSGPL-SADVAHLPFLSNLSLASNKFSGPIPPSLSALSGLRFLNLSNNVFNETFPSELSR 179
Query: 139 LTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQK 198
L L+ LD+ + G +P ++ + NL +L LGGN +SG IPPE G+ L +LA+
Sbjct: 180 LQNLEVLDLYNNNMTGVLPLAVAQMQNLRHLHLGGNFFSG-QIPPEYGRWQRLQYLAVSG 356
Query: 199 SNLVGSIPQEIGFLTNLAYIDLSK-NSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHS 257
+ L G+IP EIG L++L + + N+ +GGIP IGNLS+L L + +SG IP +
Sbjct: 357 NELEGTIPPEIGNLSSLRELYIGYYNTYTGGIPPEIGNLSELVRLDAAY-CGLSGEIPAA 533
Query: 258 LWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYL 317
L + L L+ LSGS+ + NL +LK + L N LSG IP+ G+LKN+ L L
Sbjct: 534 LGKLQKLDTLFLQVNALSGSLTPELGNLKSLKSMDLSNNMLSGEIPARFGELKNITLLNL 713
Query: 318 GSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNG 377
N L G IP IG L L+V+ + ENN TG+IP +G NG
Sbjct: 714 FRNKLHGAIPEFIGELPALEVVQLWENNFTGSIPEGLGK-------------------NG 836
Query: 378 LYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITL 437
N+ + +S N G LP+ +CSG +L+ L N GPIP SL +C S+ RI +
Sbjct: 837 RLNLVD-----LSSNKLTGTLPTYLCSGNTLQTLITLGNFLFGPIPESLGSCESLTRIRM 1001
Query: 438 EVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLD 497
N + G I + PKL ++L DN G+ ++NL +SNN +SGV+P
Sbjct: 1002 GENFLNGSIPRGLFGLPKLTQVELQDNYLSGEFPEVGSVAVNLGQITLSNNQLSGVLPPS 1181
Query: 498 FIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELD 557
+ + L L N TG++P ++ G ++ L + S N FS I EI + L LD
Sbjct: 1182 IGNFSSVQKLILDGNMFTGRIPPQI-GRLQQLSKIDFSGNKFSGPIVPEISQCKLLTFLD 1358
Query: 558 LGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFDSGLESLDLSGNFLKGNIPTGL 617
L NELSG IP E+ + L LNLSR N L G IP+ +
Sbjct: 1359 LSRNELSGDIPNEITGMRILNYLNLSR----------------------NHLVGGIPSSI 1472
Query: 618 ADLVRLSKLNLSHNMLSGTIPQNFGRNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNN 677
+ + L+ ++ S+N LSG +P F ++ S N
Sbjct: 1473 SSMQSLTSVDFSYNNLSGLVPGT----------------------GQFSYFNYTSFLGNP 1586
Query: 678 HLCGNIRGL--DPCATSHSRKRKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNEES 735
LCG G D A + L F L V L+LC + + + + + S
Sbjct: 1587 DLCGPYLGACKDGVANGAHQPHVKGLSSSFKLLLVVGLLLCSIAFAVAAIFKARSLKKAS 1766
Query: 736 QTEEVQRGVLFSIWSHDGKMMFENIIEATAN-FDDKYLVGVGSQGNVYKAELSEGLVVAV 794
W + ++ + + ++G G G VYK + G VAV
Sbjct: 1767 GAR---------AWKLTAFQRLDFTVDDVLHCLKEDNIIGKGGAGIVYKGAMPNGDHVAV 1919
Query: 795 KKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLD 854
K+L ++ F +EI+TL I+HR+I++L GFCS+ + + LVY+++ GSL
Sbjct: 1920 KRLPAMS---RGSSHDHGFNAEIQTLGRIRHRHIVRLLGFCSNHETNLLVYEYMPNGSLG 2090
Query: 855 QILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVS 914
++L+ + W+ R + A L YLHHDCSP I+HRD+ S N+LL+ ++EAHV+
Sbjct: 2091 EVLHG-KKGGHLHWDTRYKIAVEAAKGLCYLHHDCSPLIVHRDVKSNNILLDSNHEAHVA 2267
Query: 915 DFGTAKFLKPGLHS--WTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP- 971
DFG AKFL+ S + AG++GY APE A T++V+EK DVYSFGV+ LE I G+ P
Sbjct: 2268 DFGLAKFLQDSGTSECMSAIAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELITGRKPV 2447
Query: 972 ---GDLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRL 1028
GD + + +N + VLD R V P+ EV+ + +A C+ +
Sbjct: 2448 GEFGDGVDIVQWVRKMTDSNKEGVLKVLDPRLPSV--PL-HEVMHVFYVAMLCVEEQAVE 2618
Query: 1029 RPSMGQVCKML 1039
RP+M +V ++L
Sbjct: 2619 RPTMREVVQIL 2651
>TC209490 UP|Q8GSN9 (Q8GSN9) LRR receptor-like kinase (Fragment), complete
Length = 3298
Score = 440 bits (1132), Expect = e-123
Identities = 336/1031 (32%), Positives = 514/1031 (49%), Gaps = 18/1031 (1%)
Frame = +1
Query: 28 ALLKWKDSF--DDQSQTLLSTWKNNTN-PCKPKWRGIKCDKSNFISTIGLANLGLKGTLH 84
+LLK KDS D L WK + + G+KCD+ + I ++ + L
Sbjct: 181 SLLKLKDSMKGDKAKDDALHDWKFFPSLSAHCFFSGVKCDRELRVVAINVSFVPL----- 345
Query: 85 SLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQF 144
+G +P +IG L + LT N G +P+E+ LT L+
Sbjct: 346 --------------------FGHLPPEIGQLDKLENLTVSQNNLTGVLPKELAALTSLKH 465
Query: 145 LDISFCKLNGAIP-KSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVG 203
L+IS +G P + I +T L L + NN++G P+P E+ KL L +L + + G
Sbjct: 466 LNISHNVFSGHFPGQIILPMTKLEVLDVYDNNFTG-PLPVELVKLEKLKYLKLDGNYFSG 642
Query: 204 SIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSS 263
SIP+ +L ++ LS NSLSG IP+++ L L L L N G IP +M S
Sbjct: 643 SIPESYSEFKSLEFLSLSTNSLSGKIPKSLSKLKTLRYLKLGYNNAYEGGIPPEFGSMKS 822
Query: 264 LTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLS 323
L L + LSG IP S+ NL NL L L IN+L+G+IPS + + +L+ L L N+L+
Sbjct: 823 LRYLDLSSCNLSGEIPPSLANLTNLDTLFLQINNLTGTIPSELSAMVSLMSLDLSINDLT 1002
Query: 324 GPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITN 383
G IP S L NL +++ +NNL G++P+ +G L L ++ N +P L
Sbjct: 1003 GEIPMSFSQLRNLTLMNFFQNNLRGSVPSFVGELPNLETLQLWDNNFSFVLPPNLGQNGK 1182
Query: 384 WISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIE 443
F V +N F G +P +C G L+ + N F GPIP + C S+ +I N +
Sbjct: 1183 LKFFDVIKNHFTGLIPRDLCKSGRLQTIMITDNFFRGPIPNEIGNCKSLTKIRASNNYLN 1362
Query: 444 GDIAQDFGVYPKLQYLDLSDNKFHGQISPN-WGKSLNLQTFIISNNNISGVIPLDFIGLT 502
G + P + ++L++N+F+G++ P G+SL + T +SNN SG IP L
Sbjct: 1363 GVVPSGIFKLPSVTIIELANNRFNGELPPEISGESLGILT--LSNNLFSGKIPPALKNLR 1536
Query: 503 KLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNE 562
L L L +N+ G++P EV + L + IS N+ + IP+ + L +DL N
Sbjct: 1537 ALQTLSLDANEFVGEIPGEVF-DLPMLTVVNISGNNLTGPIPTTLTRCVSLTAVDLSRNM 1713
Query: 563 LSGKIPKELVELPNLRMLNLSRNKIEGIIP--IKFDSGLESLDLSGNFLKGNIPTGLADL 620
L GKIPK + L +L + N+S N+I G +P I+F L +LDLS N G +PTG
Sbjct: 1714 LEGKIPKGIKNLTDLSIFNVSINQISGPVPEEIRFMLSLTTLDLSNNNFIGKVPTG---- 1881
Query: 621 VRLSKLNLSHNMLSGTIPQNFGRNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLC 680
G+ VF S+ G P + S SL ++ L
Sbjct: 1882 ---------------------GQFAVF---SEKSFAGN-PNLCTSHSCPNSSLYPDDAL- 1983
Query: 681 GNIRGLDPCATSHSRKRKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNEESQTEEV 740
RG P + +R + + IALG L++ V +Y+M RK ++
Sbjct: 1984 KKRRG--PWSLKSTR-----VIVIVIALGTAALLVAVT---VYMMRRRKMNLAKTWKLTA 2133
Query: 741 QRGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLV 800
+ + F E+++E ++ ++G G G VY+ + G VA+K+L
Sbjct: 2134 FQRLNFKA---------EDVVEC---LKEENIIGKGGAGIVYRGSMPNGTDVAIKRL--- 2268
Query: 801 TDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNND 860
S + F +EIETL I+HRNI++L G+ S+ + + L+Y+++ GSL + L+
Sbjct: 2269 -VGAGSGRNDYGFKAEIETLGKIRHRNIMRLLGYVSNKETNLLLYEYMPNGSLGEWLHG- 2442
Query: 861 TQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAK 920
+ WE R + A L YLHHDCSP IIHRD+ S N+LL+ D EAHV+DFG AK
Sbjct: 2443 AKGGHLKWEMRYKIAVEAAKGLCYLHHDCSPLIIHRDVKSNNILLDGDLEAHVADFGLAK 2622
Query: 921 FL-KPGL-HSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP----GDL 974
FL PG S + AG++GY APE A T++V+EK DVYSFGV+ LE I+G+ P GD
Sbjct: 2623 FLYDPGASQSMSSIAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELIIGRKPVGEFGDG 2802
Query: 975 ISL--FLSPSTRPMA---NNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLR 1029
+ + +++ + +A + L+ V+D P+ P+ VI + +A C+ + R
Sbjct: 2803 VDIVGWVNKTRLELAQPSDAALVLAVVD--PRLSGYPL-TSVIYMFNIAMMCVKEMGPAR 2973
Query: 1030 PSMGQVCKMLA 1040
P+M +V ML+
Sbjct: 2974 PTMREVVHMLS 3006
>NP205681 receptor protein kinase-like protein
Length = 2946
Score = 439 bits (1128), Expect = e-123
Identities = 315/1031 (30%), Positives = 499/1031 (47%), Gaps = 18/1031 (1%)
Frame = +1
Query: 28 ALLKWKDSF--DDQSQTLLSTWKNNTN-PCKPKWRGIKCDKSNFISTIGLANLGLKGTLH 84
ALLK K+S D L WK +T+ + G+ CD+ + I ++ + L
Sbjct: 91 ALLKLKESMKGDRAKDDALHDWKFSTSLSAHCFFSGVSCDQELRVVAINVSFVPL----- 255
Query: 85 SLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQF 144
+G +P +IG L + LT N G +P+E+ LT L+
Sbjct: 256 --------------------FGHVPPEIGELDKLENLTISQNNLTGELPKELAALTSLKH 375
Query: 145 LDISFCKLNGAIP-KSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVG 203
L+IS +G P K I +T L L + NN++G +P E KL L +L + + G
Sbjct: 376 LNISHNVFSGYFPGKIILPMTELEVLDVYDNNFTGS-LPEEFVKLEKLKYLKLDGNYFSG 552
Query: 204 SIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSS 263
SIP+ +L ++ LS NSLSG IP+++ L L L L N G IP M S
Sbjct: 553 SIPESYSEFKSLEFLSLSTNSLSGNIPKSLSKLKTLRILKLGYNNAYEGGIPPEFGTMES 732
Query: 264 LTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLS 323
L L + LSG IP S+ N+ NL L L +N+L+G+IPS + D+ +L+ L L N L+
Sbjct: 733 LKYLDLSSCNLSGEIPPSLANMRNLDTLFLQMNNLTGTIPSELSDMVSLMSLDLSFNGLT 912
Query: 324 GPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITN 383
G IP L NL +++ NNL G++P+ +G L L ++ N +P L
Sbjct: 913 GEIPTRFSQLKNLTLMNFFHNNLRGSVPSFVGELPNLETLQLWENNFSSELPQNLGQNGK 1092
Query: 384 WISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIE 443
+ F V++N F G +P +C G L+ N F GPIP + C S+ +I N +
Sbjct: 1093 FKFFDVTKNHFSGLIPRDLCKSGRLQTFLITDNFFHGPIPNEIANCKSLTKIRASNNYLN 1272
Query: 444 GDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTK 503
G + P + ++L++N+F+G++ P ISG
Sbjct: 1273 GAVPSGIFKLPSVTIIELANNRFNGELPP----------------EISG---------DS 1377
Query: 504 LGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNEL 563
LG+L LS+N TGK+P L +++L L + N F IP E+ L L +++ GN L
Sbjct: 1378 LGILTLSNNLFTGKIP-PALKNLRALQTLSLDTNEFLGEIPGEVFDLPMLTVVNISGNNL 1554
Query: 564 SGKIPKELVELPNLRMLNLSRNKIEGIIP--IKFDSGLESLDLSGNFLKGNIPTGLADLV 621
+G IP +L ++LSRN ++G IP +K + L ++S N + G++P + ++
Sbjct: 1555 TGPIPTTFTRCVSLAAVDLSRNMLDGEIPKGMKNLTDLSIFNVSINQISGSVPDEIRFML 1734
Query: 622 RLSKLNLSHNMLSGTIPQNFGRNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCG 681
L+ L+LS+N G +P G+ LVF S +S N +LC
Sbjct: 1735 SLTTLDLSYNNFIGKVPTG-GQFLVF---------------------SDKSFAGNPNLCS 1848
Query: 682 NIRGLDPCATSHSRKRKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNEESQTEEVQ 741
+ C S +KR+ P + VI+++ + ++ G + + +
Sbjct: 1849 S----HSCPNSSLKKRRG---PWSLKSTRVIVMVIALATAAILVAGTEYMRRRRKLK--- 1998
Query: 742 RGVLFSIWSHDG----KMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKL 797
L W G + E ++E ++ ++G G G VY+ + G VA+K+L
Sbjct: 1999 ---LAMTWKLTGFQRLNLKAEEVVEC---LKEENIIGKGGAGIVYRGSMRNGSDVAIKRL 2160
Query: 798 HLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQIL 857
S + F +EIET+ I+HRNI++L G+ S+ + + L+Y+++ GSL + L
Sbjct: 2161 ----VGAGSGRNDYGFKAEIETVGKIRHRNIMRLLGYVSNKETNLLLYEYMPNGSLGEWL 2328
Query: 858 NNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFG 917
+ + WE R + A L YLHHDCSP IIHRD+ S N+LL+ +EAHV+DFG
Sbjct: 2329 HG-AKGGHLKWEMRYKIAVEAAKGLCYLHHDCSPLIIHRDVKSNNILLDAHFEAHVADFG 2505
Query: 918 TAKFLKP--GLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP---- 971
AKFL S + AG++GY APE A T++V+EK DVYSFGV+ LE I+G+ P
Sbjct: 2506 LAKFLYDLGSSQSMSSIAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELIIGRKPVGEF 2685
Query: 972 GDLISL--FLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLR 1029
GD + + +++ + ++ VL ++ VI + +A C+ + R
Sbjct: 2686 GDGVDIVGWVNKTRLELSQPSDAAVVLAVVDPRLSGYPLISVIYMFNIAMMCVKEVGPTR 2865
Query: 1030 PSMGQVCKMLA 1040
P+M +V ML+
Sbjct: 2866 PTMREVVHMLS 2898
>TC222007 UP|Q9LKZ5 (Q9LKZ5) Receptor-like protein kinase 2, complete
Length = 3192
Score = 433 bits (1113), Expect = e-121
Identities = 315/1024 (30%), Positives = 491/1024 (47%), Gaps = 12/1024 (1%)
Frame = +3
Query: 28 ALLKWKDSFDDQSQTLLSTWKNNTNPCKPKWRGIKCDKSNFISTIGLANLGLKGTLHSLT 87
ALL + D + +LS+W + C W G+ CD ++ + L L L GTL
Sbjct: 105 ALLSLRSVITDATPPVLSSWNASIPYCS--WLGVTCDNRRHVTALNLTGLDLSGTLS--- 269
Query: 88 FSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDI 147
A + +L +S L+ N F G IP + L+GL++L++
Sbjct: 270 ----------------------ADVAHLPFLSNLSLAANKFSGPIPPSLSALSGLRYLNL 383
Query: 148 SFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQ 207
S N P + L +L L L NN +G +P + ++ NL HL + + G IP
Sbjct: 384 SNNVFNETFPSELWRLQSLEVLDLYNNNMTG-VLPLAVAQMQNLRHLHLGGNFFSGQIPP 560
Query: 208 EIGFLTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVL 267
E G L Y+ +S N L G IP IGNL+ L L +
Sbjct: 561 EYGRWQRLQYLAVSGNELDGTIPPEIGNLTSLRELYIG---------------------- 674
Query: 268 YFDNIGLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIP 327
Y++ +G IP I NL L L + LSG IP+ +G L+ L L+L N LSG +
Sbjct: 675 YYNTY--TGGIPPEIGNLSELVRLDVAYCALSGEIPAALGKLQKLDTLFLQVNALSGSLT 848
Query: 328 ASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISF 387
+GNL +L+ + + N L+G IPAS G LK +T+ + NKLHG IP + +
Sbjct: 849 PELGNLKSLKSMDLSNNMLSGEIPASFGELKNITLLNLFRNKLHGAIPEFIGELPALEVV 1028
Query: 388 VVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIA 447
+ EN+ G +P + G L L++ N+ TG +P L + ++++ + N + G I
Sbjct: 1029 QLWENNLTGSIPEGLGKNGRLNLVDLSSNKLTGTLPPYLCSGNTLQTLITLGNFLFGPIP 1208
Query: 448 QDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVL 507
+ G L + + +N +G I L + +N +SG P LG +
Sbjct: 1209 ESLGTCESLTRIRMGENFLNGSIPKGLFGLPKLTQVELQDNYLSGEFPEVGSVAVNLGQI 1388
Query: 508 HLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKI 567
LS+NQL+G L + G S+ L + N F+ IP++IG LQ+L ++D GN+ SG I
Sbjct: 1389 TLSNNQLSGALSPSI-GNFSSVQKLLLDGNMFTGRIPTQIGRLQQLSKIDFSGNKFSGPI 1565
Query: 568 PKELVELPNLRMLNLSRNKIEGIIPIKFDSGLESLDLSGNFLKGNIPTGLADLVRLSKLN 627
E+ + L L+LSRN+ L G+IP + + L+ LN
Sbjct: 1566 APEISQCKLLTFLDLSRNE----------------------LSGDIPNEITGMRILNYLN 1679
Query: 628 LSHNMLSGTIPQNFG--RNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRG 685
LS N L G+IP + ++L V+ S N L G +P F ++ S N LCG G
Sbjct: 1680 LSKNHLVGSIPSSISSMQSLTSVDFSYNNLSGLVPGTGQFSYFNYTSFLGNPDLCGPYLG 1859
Query: 686 LDPCATSHSRKRKNVLRPVFIALGAVILVLCVVGALM----YIMCGRKKPNEESQTEEVQ 741
++ + +V L + + +L VVG L+ + + K + E +
Sbjct: 1860 ACKGGVANGAHQPHVK-----GLSSSLKLLLVVGLLLCSIAFAVAAIFKARSLKKASEAR 2024
Query: 742 RGVLFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVT 801
L + D ++++ + ++G G G VYK + G VAVK+L ++
Sbjct: 2025 AWKLTAFQRLD--FTVDDVLHC---LKEDNIIGKGGAGIVYKGAMPNGDHVAVKRLPAMS 2189
Query: 802 DEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDT 861
F +EI+TL I+HR+I++L GFCS+ + + LVY+++ GSL ++L+
Sbjct: 2190 ---RGSSHDHGFNAEIQTLGRIRHRHIVRLLGFCSNHETNLLVYEYMPNGSLGEVLHGK- 2357
Query: 862 QAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKF 921
+ W+ R + A L YLHHDCSP I+HRD+ S N+LL+ ++EAHV+DFG AKF
Sbjct: 2358 KGGHLHWDTRYKIAVEAAKGLCYLHHDCSPLIVHRDVKSNNILLDSNHEAHVADFGLAKF 2537
Query: 922 LKPGLHS--WTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHP----GDLI 975
L+ S + AG++GY APE A T++V+EK DVYSFGV+ LE I G+ P GD +
Sbjct: 2538 LQDSGTSECMSAIAGSYGYIAPEYAYTLKVDEKSDVYSFGVVLLELITGRKPVGEFGDGV 2717
Query: 976 SLFLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQV 1035
+ +N + VLD R V P+ EV+ + +A C+ + RP+M +V
Sbjct: 2718 DIVQWVRKMTDSNKEGVLKVLDPRLPSV--PL-HEVMHVFYVAMLCVEEQAVERPTMREV 2888
Query: 1036 CKML 1039
++L
Sbjct: 2889 VQIL 2900
>TC229875 weakly similar to UP|Q75WU3 (Q75WU3) Leucine-rich repeat
receptor-like protein kinase 1, partial (23%)
Length = 1141
Score = 369 bits (946), Expect = e-102
Identities = 191/298 (64%), Positives = 228/298 (76%), Gaps = 4/298 (1%)
Frame = +2
Query: 751 HDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSS 810
H GKM+FENIIEAT +FDDK+L+GVG QG VYKA L G VVAVKKLH V + EM +
Sbjct: 2 HQGKMVFENIIEATEDFDDKHLIGVGGQGCVYKAVLPTGQVVAVKKLHSVPNGEM--LNL 175
Query: 811 KSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEK 870
K+F EI+ LT I+HRNI+KL+GFCSHS+FSFLV +FLE GS+ + L +D QA+AFDW K
Sbjct: 176 KAFTCEIQALTEIRHRNIVKLYGFCSHSQFSFLVCEFLENGSVGKTLKDDGQAMAFDWYK 355
Query: 871 RVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPGLHSWT 930
RVNVVK VANAL Y+HH+CSP I+HRDISSKNVLL+ +Y AHVSDFGTAKFL P +WT
Sbjct: 356 RVNVVKDVANALCYMHHECSPRIVHRDISSKNVLLDSEYVAHVSDFGTAKFLNPDSSNWT 535
Query: 931 QFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFL--SPST--RPM 986
F GTFGYAAPELA TMEVNEKCDVYSFGVLA E ++GKHPGD+IS L SPST
Sbjct: 536 SFVGTFGYAAPELAYTMEVNEKCDVYSFGVLAWEILIGKHPGDVISCLLGSSPSTLVAST 715
Query: 987 ANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKMLAIGKS 1044
++M L D LDQR +PI +EV IA++A ACL+++PR RP+M QV L + S
Sbjct: 716 LDHMALMDKLDQRLPHPTKPIGKEVASIAKIAMACLTESPRSRPTMEQVANELVMSSS 889
>TC204550 UP|Q8L3Y5 (Q8L3Y5) Receptor-like kinase RHG1, complete
Length = 2659
Score = 275 bits (704), Expect = 6e-74
Identities = 216/734 (29%), Positives = 342/734 (46%), Gaps = 56/734 (7%)
Frame = +2
Query: 362 VFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGP 421
V ++ L GRI + + + + +N G +PS + +LR + +NR TG
Sbjct: 449 VIQLPWKGLRGRITDKIGQLQGLRKLSLHDNQIGGSIPSTLGLLPNLRGVQLFNNRLTGS 628
Query: 422 IPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQ 481
IP SL C ++ + L N + G I KL +L+LS N F G + + S +L
Sbjct: 629 IPLSLGFCPLLQSLDLSNNLLTGAIPYSLANSTKLYWLNLSFNSFSGPLPASLTHSFSLT 808
Query: 482 TFIISNNNISGVIPLDFIGLTKLGVLHLSS-----NQLTGKLPMEVLGGMKSLFDLKISN 536
+ NNN+SG +P + G +K G L + N TG +P LG ++ L ++ +S+
Sbjct: 809 FLSLQNNNLSGSLPNSWGGNSKNGFFRLQNLILDHNFFTGDVPAS-LGSLRELNEISLSH 985
Query: 537 NHFSDNIPSEIGLLQRLQELDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFD 596
N FS IP+EIG L RL+ LD+ N L+G +P L L +L +LN N ++ IP
Sbjct: 986 NKFSGAIPNEIGTLSRLKTLDISNNALNGNLPATLSNLSSLTLLNAENNLLDNQIPQSLG 1165
Query: 597 S--GLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQNFG--RNLVFVNISD 652
L L LS N G+IP+ +A++ L +L+LS N SG IP +F R+L N+S
Sbjct: 1166 RLRNLSVLILSRNQFSGHIPSSIANISSLRQLDLSLNNFSGEIPVSFDSQRSLNLFNVSY 1345
Query: 653 NQLEGPLPKIPA--FLSASFESLKNNNHLCG--------------NIRGLDPCATSHSRK 696
N L G +P + A F S+SF N LCG + P + H
Sbjct: 1346 NSLSGSVPPLLAKKFNSSSFVG---NIQLCGYSPSTPCFSQAPSQGVIAPPPEVSKHHHH 1516
Query: 697 RKNVLRPVFIALGAVILVLCVVGALMYIMCGRKKPNEESQ-----------TEEVQRGVL 745
RK + + + + V+LV+ ++ + + C +K + T E ++
Sbjct: 1517 RKLSTKDIILVVAGVLLVVLIILCCVLLFCLIRKRSTSKAGNGQATEGRAVTYEDRKRSP 1696
Query: 746 FSIW---------------SHDGKMMF--ENIIEATANFDDKYLVGVGSQGNVYKAELSE 788
S W DG M F ++++ ATA ++G + G VYKA L +
Sbjct: 1697 SSCWWLMLKQVGRLEGNLVHFDGPMAFTADDLLCATAE-----IMGKSTNGTVYKAILED 1861
Query: 789 GLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFS-FLVYKF 847
G VAVK+L E + F SE+ L I+H N++ L + K LV+ +
Sbjct: 1862 GSQVAVKRLR-----EKIAKGHREFESEVSVLGKIRHPNVLALRAYYLGPKGEKLLVFDY 2026
Query: 848 LEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNL 907
+ GSL L+ DW R+ + + +A L LH + IIH +++S NVLL+
Sbjct: 2027 MSKGSLASFLHGGGTETFIDWPTRMKIAQDLARGLFCLHSQEN--IIHGNLTSSNVLLDE 2200
Query: 908 DYEAHVSDFGTAKFLKPGLHSWT-QFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETI 966
+ A ++DFG ++ + +S AG GY APEL++ + N K D+YS GV+ LE +
Sbjct: 2201 NTNAKIADFGLSRLMSTTANSNVIATAGALGYRAPELSKLKKANTKTDIYSLGVILLELL 2380
Query: 967 MGKHPG-DLISLFLSPSTRPMANNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQN 1025
K PG + L L + +V D + + +E++ +LA C+ +
Sbjct: 2381 TRKSPGVSMNGLDLPQWVASVVKEEWTNEVFDADLMRDASTVGDELLNTLKLALHCVDPS 2560
Query: 1026 PRLRPSMGQVCKML 1039
P RP + QV + L
Sbjct: 2561 PSARPEVHQVLQQL 2602
Score = 188 bits (478), Expect = 1e-47
Identities = 140/415 (33%), Positives = 205/415 (48%), Gaps = 12/415 (2%)
Frame = +2
Query: 1 MMVLPTLIMILCVLPTL-------SVAEDSEAKLALLKWKDSFDDQSQTLLSTWKNNTNP 53
++VLP+ CV P L V + LAL +K D L S +
Sbjct: 236 LVVLPS-----CVRPVLCEDEGWDGVVVTASNLLALEAFKQELADPEGFLRSWNDSGYGA 400
Query: 54 CKPKWRGIKCDKSNFISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIG 113
C W GIKC + I I L GL+G + L + + +N G+IP+ +G
Sbjct: 401 CSGGWVGIKCAQGQVI-VIQLPWKGLRGRITD-KIGQLQGLRKLSLHDNQIGGSIPSTLG 574
Query: 114 NLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGG 173
L N+ + NN GSIP + LQ LD+S L GAIP S+ N T L +L L
Sbjct: 575 LLPNLRGVQLFNNRLTGSIPLSLGFCPLLQSLDLSNNLLTGAIPYSLANSTKLYWLNLSF 754
Query: 174 NNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIG-----FLTNLAYIDLSKNSLSGG 228
N++S GP+P + +L L++Q +NL GS+P G L + L N +G
Sbjct: 755 NSFS-GPLPASLTHSFSLTFLSLQNNNLSGSLPNSWGGNSKNGFFRLQNLILDHNFFTGD 931
Query: 229 IPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNL 288
+P ++G+L +L+ + LS+N K SG IP+ + +S L L N L+G++P ++ NL +L
Sbjct: 932 VPASLGSLRELNEISLSHN-KFSGAIPNEIGTLSRLKTLDISNNALNGNLPATLSNLSSL 1108
Query: 289 KELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTG 348
L + N L IP ++G L+NL L L N SG IP+SI N+ +L+ L + NN +G
Sbjct: 1109TLLNAENNLLDNQIPQSLGRLRNLSVLILSRNQFSGHIPSSIANISSLRQLDLSLNNFSG 1288
Query: 349 TIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQIC 403
IP S + + L +F V+ N L G +P L N SF V G+ PS C
Sbjct: 1289EIPVSFDSQRSLNLFNVSYNSLSGSVPPLLAKKFNSSSF-VGNIQLCGYSPSTPC 1450
Score = 188 bits (478), Expect = 1e-47
Identities = 115/300 (38%), Positives = 158/300 (52%), Gaps = 5/300 (1%)
Frame = +2
Query: 152 LNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGF 211
L G I IG L L L L N GG IP +G L NL + + + L GSIP +GF
Sbjct: 473 LRGRITDKIGQLQGLRKLSLHDNQ-IGGSIPSTLGLLPNLRGVQLFNNRLTGSIPLSLGF 649
Query: 212 LTNLAYIDLSKNSLSGGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDN 271
L +DLS N L+G IP ++ N +KL L LS N+ SGP+P SL + SLT L N
Sbjct: 650 CPLLQSLDLSNNLLTGAIPYSLANSTKLYWLNLSFNS-FSGPLPASLTHSFSLTFLSLQN 826
Query: 272 IGLSGSIPDSI-----QNLVNLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPI 326
LSGS+P+S L+ L LD N +G +P+++G L+ L ++ L N SG I
Sbjct: 827 NNLSGSLPNSWGGNSKNGFFRLQNLILDHNFFTGDVPASLGSLRELNEISLSHNKFSGAI 1006
Query: 327 PASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRIPNGLYNITNWIS 386
P IG L L+ L + N L G +PA++ NL LT+ N L +IP L + N
Sbjct: 1007PNEIGTLSRLKTLDISNNALNGNLPATLSNLSSLTLLNAENNLLDNQIPQSLGRLRNLSV 1186
Query: 387 FVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIERITLEVNQIEGDI 446
++S N F GH+PS I + SLR L+ N F+G IP S + S+ + N + G +
Sbjct: 1187LILSRNQFSGHIPSSIANISSLRQLDLSLNNFSGEIPVSFDSQRSLNLFNVSYNSLSGSV 1366
Score = 164 bits (416), Expect = 1e-40
Identities = 109/322 (33%), Positives = 166/322 (50%), Gaps = 2/322 (0%)
Frame = +2
Query: 176 WSG--GPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETI 233
W G G I +IG+L L L++ + + GSIP +G L NL + L N L+G IP ++
Sbjct: 464 WKGLRGRITDKIGQLQGLRKLSLHDNQIGGSIPSTLGLLPNLRGVQLFNNRLTGSIPLSL 643
Query: 234 GNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELAL 293
G L +L LSNN ++G IP+SL N + L L SG +P S+ + +L L+L
Sbjct: 644 GFCPLLQSLDLSNNL-LTGAIPYSLANSTKLYWLNLSFNSFSGPLPASLTHSFSLTFLSL 820
Query: 294 DINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPAS 353
N+LSGS+P++ G N+ +G LQ L + N TG +PAS
Sbjct: 821 QNNNLSGSLPNS-----------WGGNSKNG--------FFRLQNLILDHNFFTGDVPAS 943
Query: 354 IGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNA 413
+G+L+ L ++ NK G IPN + ++ + +S N G+LP+ + + SL LLNA
Sbjct: 944 LGSLRELNEISLSHNKFSGAIPNEIGTLSRLKTLDISNNALNGNLPATLSNLSSLTLLNA 1123
Query: 414 DHNRFTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPN 473
++N IP SL ++ + L NQ G I L+ LDLS N F G+I +
Sbjct: 1124ENNLLDNQIPQSLGRLRNLSVLILSRNQFSGHIPSSIANISSLRQLDLSLNNFSGEIPVS 1303
Query: 474 WGKSLNLQTFIISNNNISGVIP 495
+ +L F +S N++SG +P
Sbjct: 1304FDSQRSLNLFNVSYNSLSGSVP 1369
>TC234472 weakly similar to UP|Q75WU3 (Q75WU3) Leucine-rich repeat
receptor-like protein kinase 1, partial (16%)
Length = 553
Score = 255 bits (652), Expect = 6e-68
Identities = 117/178 (65%), Positives = 154/178 (85%)
Frame = +1
Query: 745 LFSIWSHDGKMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEE 804
LF+IWS DGK+++ENI+EAT +FD+K+L+GVG QG+VYKA+L G ++AVKKLHLV + E
Sbjct: 22 LFAIWSFDGKLVYENIVEATEDFDNKHLIGVGGQGSVYKAKLHTGQILAVKKLHLVQNGE 201
Query: 805 MSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAV 864
+S + K+F SEI+ L I+HRNI+KL+GFCSHS+ SFLVY+FLE GS+D+IL +D QA+
Sbjct: 202 LS--NIKAFTSEIQALINIRHRNIVKLYGFCSHSQSSFLVYEFLEKGSIDKILKDDEQAI 375
Query: 865 AFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFL 922
AFDW+ R+N +KGVANALSY+HHDCSPPI+HRDISSKN++L+L+Y AHVSDFG A+ L
Sbjct: 376 AFDWDPRINAIKGVANALSYMHHDCSPPIVHRDISSKNIVLDLEYVAHVSDFGAARLL 549
>TC209281 similar to UP|Q75WU3 (Q75WU3) Leucine-rich repeat receptor-like
protein kinase 1, partial (14%)
Length = 586
Score = 235 bits (599), Expect = 9e-62
Identities = 118/193 (61%), Positives = 144/193 (74%), Gaps = 3/193 (1%)
Frame = +2
Query: 846 KFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLL 905
+FLE G + +IL +D QA+A DW KRV++VKGVANAL Y+HHDCSPPI+HRDISSKNVLL
Sbjct: 8 EFLEKGDVKKILKDDEQAIALDWNKRVDIVKGVANALCYMHHDCSPPIVHRDISSKNVLL 187
Query: 906 NLDYEAHVSDFGTAKFLKPGLHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALET 965
+ D AHV+DFGTAKFL P +WT FAGT+GYAAPELA TME NEKCDVYSFGV ALE
Sbjct: 188 DSDDVAHVADFGTAKFLNPDSSNWTSFAGTYGYAAPELAYTMEANEKCDVYSFGVFALEI 367
Query: 966 IMGKHPGDLISLFLSPSTRPMA---NNMLLTDVLDQRPQQVMEPIDEEVILIARLAFACL 1022
+ G+HPGD+ S L S+ M ++M L LD+R PID+EVI I ++A A L
Sbjct: 368 LFGEHPGDVTSSLLLSSSSTMTSTLDHMSLMVKLDERLPHPTSPIDKEVISIVKIAIARL 547
Query: 1023 SQNPRLRPSMGQV 1035
+++ R RPSM QV
Sbjct: 548 TESTRSRPSMEQV 586
>TC217385 similar to UP|Q6ZGC7 (Q6ZGC7) Receptor protein kinase PERK1-like,
partial (80%)
Length = 1604
Score = 204 bits (518), Expect = 2e-52
Identities = 137/430 (31%), Positives = 217/430 (49%), Gaps = 22/430 (5%)
Frame = +1
Query: 644 NLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNIRGLDPCATSHSRKRKNVLRP 703
+L +N+S N+L G +P F +S N LCGN L PC + +R + +
Sbjct: 4 SLSLLNVSYNKLFGVIPTSNNFTRFPPDSFIGNPGLCGNWLNL-PCHGARPSERVTLSKA 180
Query: 704 VF--IALGAVILVLCVVGALMYIMCGRKKPNEESQTEE--VQRGVLFS------IWSHDG 753
I LGA++++L V+ A +P+ S + + + FS + +
Sbjct: 181 AILGITLGALVILLMVLVAAC-------RPHSPSPFPDGSFDKPINFSPPKLVILHMNMA 339
Query: 754 KMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSF 813
++E+I+ T N +KY++G G+ VYK L VA+K+++ C K F
Sbjct: 340 LHVYEDIMRMTENLSEKYIIGYGASSTVYKCVLKNCKPVAIKRIY---SHYPQCI--KEF 504
Query: 814 MSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVN 873
+E+ET+ IKHRN++ L G+ L Y ++E GSL +L+ T+ DWE R+
Sbjct: 505 ETELETVGSIKHRNLVSLQGYSLSPYGHLLFYDYMENGSLWDLLHGPTKKKKLDWELRLK 684
Query: 874 VVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPG-LHSWTQF 932
+ G A L+YLHHDC P IIHRD+ S N+LL+ D+E H++DFG AK L P H+ T
Sbjct: 685 IALGAAQGLAYLHHDCCPRIIHRDVKSSNILLDADFEPHLTDFGIAKSLCPSKSHTSTYI 864
Query: 933 AGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNMLL 992
GT GY PE A+T + EK DVYS+G++ LE + G+ D ++++L
Sbjct: 865 MGTIGYIDPEYARTSHLTEKSDVYSYGIVLLELLTGRKAVD---------NESNLHHLIL 1017
Query: 993 TDVLDQRPQQVMEPIDEEVIL----------IARLAFACLSQNPRLRPSMGQVCKML-AI 1041
+ VME +D ++ + +LA C + P RP+M +V ++L ++
Sbjct: 1018 SKA---ATNAVMETVDPDITATCKDLGAVKKVYQLALLCTKRQPADRPTMHEVTRVLGSL 1188
Query: 1042 GKSPLVGKQL 1051
S + KQL
Sbjct: 1189 VPSSIPPKQL 1218
>TC231717 weakly similar to UP|Q8VZG8 (Q8VZG8) AT4g08850/T32A17_160, partial
(11%)
Length = 453
Score = 198 bits (503), Expect = 1e-50
Identities = 93/144 (64%), Positives = 117/144 (80%)
Frame = +3
Query: 781 VYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKF 840
VYKA+L G +VAVKKLH +EE SK+F +E++ L IKHRNI+K G+C H +F
Sbjct: 6 VYKAKLPAGQIVAVKKLHAAPNEETP--DSKAFSTEVKALAEIKHRNIVKSLGYCLHPRF 179
Query: 841 SFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISS 900
SFL+Y+FLEGGSLD++L +DT+A FDWE+RV VVKGVA+AL ++HH C PPI+HRDISS
Sbjct: 180 SFLIYEFLEGGSLDKVLTDDTRATMFDWERRVKVVKGVASALYHMHHGCFPPIVHRDISS 359
Query: 901 KNVLLNLDYEAHVSDFGTAKFLKP 924
KNVL++LDYEAH+SDFGTAK LKP
Sbjct: 360 KNVLIDLDYEAHISDFGTAKILKP 431
>TC218252 weakly similar to UP|Q03407 (Q03407) Ydr490cp, partial (5%)
Length = 1279
Score = 185 bits (470), Expect = 8e-47
Identities = 100/233 (42%), Positives = 141/233 (59%), Gaps = 6/233 (2%)
Frame = +1
Query: 813 FMSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRV 872
F E+E L IKHR ++ L G+C+ L+Y +L GGSLD+ L+ +A DW+ R+
Sbjct: 46 FERELEILGSIKHRYLVNLRGYCNSPTSKLLIYDYLPGGSLDEALHE--RADQLDWDSRL 219
Query: 873 NVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLK-PGLHSWTQ 931
N++ G A L+YLHHDCSP IIHRDI S N+LL+ + EA VSDFG AK L+ H T
Sbjct: 220 NIIMGAAKGLAYLHHDCSPRIIHRDIKSSNILLDGNLEARVSDFGLAKLLEDEESHITTI 399
Query: 932 FAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNML 991
AGTFGY APE Q+ EK DVYSFGVL LE + GK P D + P+ N +
Sbjct: 400 VAGTFGYLAPEYMQSGRATEKSDVYSFGVLTLEVLSGKRPTDAAFIEKGPNIVGWLNFL- 576
Query: 992 LTDVLDQRPQQVMEPIDEEVIL-----IARLAFACLSQNPRLRPSMGQVCKML 1039
+ + RP+++++P+ E V + + +A C+S +P RP+M +V ++L
Sbjct: 577 ---ITENRPREIVDPLCEGVQMESLDALLSVAIQCVSSSPEDRPTMHRVVQLL 726
>TC216858 homologue to UP|Q76FZ8 (Q76FZ8) Brassinosteroid receptor, partial
(29%)
Length = 1320
Score = 183 bits (465), Expect = 3e-46
Identities = 106/290 (36%), Positives = 169/290 (57%), Gaps = 5/290 (1%)
Frame = +1
Query: 754 KMMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSF 813
K+ F ++++AT F + L+G G G+VYKA+L +G VVA+KKL V+ + + F
Sbjct: 43 KLTFADLLDATNGFHNDSLIGSGGFGDVYKAQLKDGSVVAIKKLIHVSGQ-----GDREF 207
Query: 814 MSEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQA-VAFDWEKRV 872
+E+ET+ IKHRN++ L G+C + LVY++++ GSL+ +L++ +A + +W R
Sbjct: 208 TAEMETIGKIKHRNLVPLLGYCKVGEERLLVYEYMKYGSLEDVLHDQKKAGIKLNWAIRR 387
Query: 873 NVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKP-GLH-SWT 930
+ G A L++LHH+C P IIHRD+ S NVLL+ + EA VSDFG A+ + H S +
Sbjct: 388 KIAIGAARGLAFLHHNCIPHIIHRDMKSSNVLLDENLEARVSDFGMARLMSAMDTHLSVS 567
Query: 931 QFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANN- 989
AGT GY PE Q+ + K DVYS+GV+ LE + GK P D + + +
Sbjct: 568 TLAGTPGYVPPEYYQSFRCSTKGDVYSYGVVLLELLTGKRPTDSADFGDNNLVGWVKQHA 747
Query: 990 -MLLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKM 1038
+ ++D+ D + ++ E++ ++A +CL P RP+M QV M
Sbjct: 748 KLKISDIFDPELMKEDPNLEMELLQHLKIAVSCLDDRPWRRPTMIQVMAM 897
>TC231346 weakly similar to UP|Q6QM04 (Q6QM04) LRR protein WM1.2, partial
(14%)
Length = 1790
Score = 175 bits (444), Expect = 8e-44
Identities = 153/555 (27%), Positives = 253/555 (45%), Gaps = 75/555 (13%)
Frame = +1
Query: 89 SSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYFDGSIPQEMCTLTGLQFLDIS 148
SS N+ +D++NN G +P +G L ++ +L NN F IP L+ L+ L+++
Sbjct: 46 SSLQNIKNLDLQNNQLSGPLPDSLGQLKHLEVLDLSNNTFTCPIPSPFANLSSLRTLNLA 225
Query: 149 FCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPEIGKLNNLLHLAIQKSNLVGSIPQE 208
+LNG IPKS L NL L LG N+ + G +P +G L+NL+ L + + L GSI +E
Sbjct: 226 HNRLNGTIPKSFEFLKNLQVLNLGANSLT-GDVPVTLGTLSNLVTLDLSSNLLEGSI-KE 399
Query: 209 IGFL------------TN--------------LAYIDLSKNSLSGGIPETIGNLSKLDTL 242
F+ TN L Y+ LS + PE + S + L
Sbjct: 400 SNFVKLFTLKELRLSWTNLFLSVNSGWAPPFQLEYVLLSSFGIGPKFPEWLKRQSSVKVL 579
Query: 243 VLSNNTKMSGPIPHSLWNMSSLTVLYFD-----------NIGLSGSIPDSIQNL------ 285
+S ++ +P W + +L + + D NI L+ S+ + NL
Sbjct: 580 TMS-KAGIADLVPSWFW-IWTLQIEFLDLSNNLLRGDLSNIFLNSSVINLSSNLFKGRLP 753
Query: 286 ---VNLKELALDINHLSGSIPSTI---------------------GDL-------KNLIK 314
N++ L + N +SG+I + GDL + L+
Sbjct: 754 SVSANVEVLNVANNSISGTISPFLCGKPNATNKLSVLDFSNNVLSGDLGHCWVHWQALVH 933
Query: 315 LYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLTVFEVATNKLHGRI 374
+ LGSNN+SG IP S+G L L+ L + +N +G IP+++ N + ++ N+L I
Sbjct: 934 VNLGSNNMSGEIPNSMGYLSQLESLLLDDNRFSGYIPSTLQNCSTMKFIDMGNNQLSDTI 1113
Query: 375 PNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSLKTCSSIE- 433
P+ ++ + + + N+F G + ++C SL +L+ +N +G IP L ++
Sbjct: 1114PDWMWEMQYLMVLRLRSNNFNGSITQKMCQLSSLIVLDHGNNSLSGSIPNCLDDMKTMAG 1293
Query: 434 RITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGV 493
N DF + L L K + N + ++ +S+N +SG
Sbjct: 1294EDDFFANPSSYSYGSDFSYNHYKETLVLVPKKDELEYRDN---LILVRMIDLSSNKLSGA 1464
Query: 494 IPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRL 553
IP + L+ L L+LS N L+G++P + +G MK L L +S N+ S IP + L L
Sbjct: 1465IPSEISKLSALRFLNLSRNHLSGEIPND-MGKMKLLESLDLSLNNISGQIPQSLSDLSFL 1641
Query: 554 QELDLGGNELSGKIP 568
L+L + LSG+IP
Sbjct: 1642SFLNLSYHNLSGRIP 1686
Score = 174 bits (442), Expect = 1e-43
Identities = 158/584 (27%), Positives = 259/584 (44%), Gaps = 81/584 (13%)
Frame = +1
Query: 179 GPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIGFLTNLAYIDLSKNSLSGGIPETIGNLSK 238
G IP I L N+ +L +Q + L G +P +G L +L +DLS N+ + IP NLS
Sbjct: 25 GKIPQIISSLQNIKNLDLQNNQLSGPLPDSLGQLKHLEVLDLSNNTFTCPIPSPFANLSS 204
Query: 239 LDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVNLKELALDINHL 298
L TL L++N +++G IP S + +L VL L+G +P ++ L NL L L N L
Sbjct: 205 LRTLNLAHN-RLNGTIPKSFEFLKNLQVLNLGANSLTGDVPVTLGTLSNLVTLDLSSNLL 381
Query: 299 SGSIPS-------TIGDLK------------------NLIKLYLGSNNLSGPIPASIGNL 333
GSI T+ +L+ L + L S + P +
Sbjct: 382 EGSIKESNFVKLFTLKELRLSWTNLFLSVNSGWAPPFQLEYVLLSSFGIGPKFPEWLKRQ 561
Query: 334 INLQVLSVQENNLTGTIPA--SIGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSE 391
+++VL++ + + +P+ I L+ + +++ N L G + N N + +S
Sbjct: 562 SSVKVLTMSKAGIADLVPSWFWIWTLQ-IEFLDLSNNLLRGDLSNIFLNSS---VINLSS 729
Query: 392 NDFVGHLPSQICSGGSLRLLNADHNRFTGPIPTSL----KTCSSIERITLEVNQIEGDIA 447
N F G LPS ++ +LN +N +G I L + + + N + GD+
Sbjct: 730 NLFKGRLPS---VSANVEVLNVANNSISGTISPFLCGKPNATNKLSVLDFSNNVLSGDLG 900
Query: 448 QDFGVYPKLQYLDLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVL 507
+ + L +++L N G+I + G L++ ++ +N SG IP + + +
Sbjct: 901 HCWVHWQALVHVNLGSNNMSGEIPNSMGYLSQLESLLLDDNRFSGYIPSTLQNCSTMKFI 1080
Query: 508 HLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGNELSGKI 567
+ +NQL+ +P + + M+ L L++ +N+F+ +I ++ L L LD G N LSG I
Sbjct: 1081DMGNNQLSDTIP-DWMWEMQYLMVLRLRSNNFNGSITQKMCQLSSLIVLDHGNNSLSGSI 1257
Query: 568 P----------------------------------KELVELPN------------LRMLN 581
P + LV +P +RM++
Sbjct: 1258PNCLDDMKTMAGEDDFFANPSSYSYGSDFSYNHYKETLVLVPKKDELEYRDNLILVRMID 1437
Query: 582 LSRNKIEGIIPIKFD--SGLESLDLSGNFLKGNIPTGLADLVRLSKLNLSHNMLSGTIPQ 639
LS NK+ G IP + S L L+LS N L G IP + + L L+LS N +SG IPQ
Sbjct: 1438LSSNKLSGAIPSEISKLSALRFLNLSRNHLSGEIPNDMGKMKLLESLDLSLNNISGQIPQ 1617
Query: 640 NFG--RNLVFVNISDNQLEGPLPKIPAFLSASFESLKNNNHLCG 681
+ L F+N+S + L G +P S S N LCG
Sbjct: 1618SLSDLSFLSFLNLSYHNLSGRIPTSTQLQSFEELSYTGNPELCG 1749
Score = 154 bits (389), Expect = 2e-37
Identities = 147/527 (27%), Positives = 245/527 (45%), Gaps = 30/527 (5%)
Frame = +1
Query: 69 ISTIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTIPAQIGNLSNISILTFKNNYF 128
+ T+ LA+ L GT+ +F NL ++++ NS G +P +G LSN+ L +N
Sbjct: 205 LRTLNLAHNRLNGTIPK-SFEFLKNLQVLNLGANSLTGDVPVTLGTLSNLVTLDLSSNLL 381
Query: 129 DGSIPQ-EMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGGNNWSGGPIPPE-IG 186
+GSI + L L+ L +S+ L ++ L Y++L +++ GP PE +
Sbjct: 382 EGSIKESNFVKLFTLKELRLSWTNLFLSVNSGWAPPFQLEYVLL--SSFGIGPKFPEWLK 555
Query: 187 KLNNLLHLAIQKSNLVGSIPQEIGFLT-NLAYIDLSKNSLSGGIPETIGNLSKLDTLVLS 245
+ +++ L + K+ + +P T + ++DLS N L G + N S ++ LS
Sbjct: 556 RQSSVKVLTMSKAGIADLVPSWFWIWTLQIEFLDLSNNLLRGDLSNIFLNSSVIN---LS 726
Query: 246 NNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLVN----LKELALDINHLSGS 301
+N G +P N+ L V N +SG+I + N L L N LSG
Sbjct: 727 SNL-FKGRLPSVSANVEVLNVA---NNSISGTISPFLCGKPNATNKLSVLDFSNNVLSGD 894
Query: 302 IPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPASIGNLKWLT 361
+ + L+ + LGSNN+SG IP S+G L L+ L + +N +G IP+++ N +
Sbjct: 895 LGHCWVHWQALVHVNLGSNNMSGEIPNSMGYLSQLESLLLDDNRFSGYIPSTLQNCSTMK 1074
Query: 362 VFEVATNKLHGRIPNGLYNITNWISFVVSENDFVGHLPSQICSGGSLRLLNADHNRFTGP 421
++ N+L IP+ ++ + + + N+F G + ++C SL +L+ +N +G
Sbjct: 1075FIDMGNNQLSDTIPDWMWEMQYLMVLRLRSNNFNGSITQKMCQLSSLIVLDHGNNSLSGS 1254
Query: 422 IPTSLKTCSSIE-RITLEVNQIEGDIAQDFG---------VYPK---LQY---------L 459
IP L ++ N DF + PK L+Y +
Sbjct: 1255IPNCLDDMKTMAGEDDFFANPSSYSYGSDFSYNHYKETLVLVPKKDELEYRDNLILVRMI 1434
Query: 460 DLSDNKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLP 519
DLS NK G I K L+ +S N++SG IP D + L L LS N ++G++P
Sbjct: 1435DLSSNKLSGAIPSEISKLSALRFLNLSRNHLSGEIPNDMGKMKLLESLDLSLNNISGQIP 1614
Query: 520 MEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQELDLGGN-ELSG 565
+ L + L L +S ++ S IP+ LQ +EL GN EL G
Sbjct: 1615-QSLSDLSFLSFLNLSYHNLSGRIPTST-QLQSFEELSYTGNPELCG 1749
Score = 150 bits (380), Expect = 2e-36
Identities = 146/514 (28%), Positives = 233/514 (44%), Gaps = 57/514 (11%)
Frame = +1
Query: 227 GGIPETIGNLSKLDTLVLSNNTKMSGPIPHSLWNMSSLTVLYFDNIGLSGSIPDSIQNLV 286
G IP+ I +L + L L NN ++SGP+P SL + L VL N + IP NL
Sbjct: 25 GKIPQIISSLQNIKNLDLQNN-QLSGPLPDSLGQLKHLEVLDLSNNTFTCPIPSPFANLS 201
Query: 287 NLKELALDINHLSGSIPSTIGDLKNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNL 346
+L+ L L N L+G+IP + LKNL L LG+N+L+G +P ++G L NL L + N L
Sbjct: 202 SLRTLNLAHNRLNGTIPKSFEFLKNLQVLNLGANSLTGDVPVTLGTLSNLVTLDLSSNLL 381
Query: 347 TGTIPAS-IGNLKWLTVFEVATNKLHGRIPNGLYNITNWISFVVSENDFVG-HLPSQICS 404
G+I S L L ++ L + +G + + +V+ + +G P +
Sbjct: 382 EGSIKESNFVKLFTLKELRLSWTNLFLSVNSG-WAPPFQLEYVLLSSFGIGPKFPEWLKR 558
Query: 405 GGSLRLLNADHNRFTGPIPTSLKTCS-SIERITLEVNQIEGDIAQDFGVYPKLQYLDLSD 463
S+++L +P+ + IE + L N + GD++ ++ ++LS
Sbjct: 559 QSSVKVLTMSKAGIADLVPSWFWIWTLQIEFLDLSNNLLRGDLSN---IFLNSSVINLSS 729
Query: 464 NKFHGQISPNWGKSLNLQTFIISNNNISGVIPLDFIG----LTKLGVL------------ 507
N F G++ S N++ ++NN+ISG I G KL VL
Sbjct: 730 NLFKGRLP---SVSANVEVLNVANNSISGTISPFLCGKPNATNKLSVLDFSNNVLSGDLG 900
Query: 508 ------------HLSSNQLTGKLPMEVLGGMKSLFDLKISNNHFSDNIPSEIGLLQRLQE 555
+L SN ++G++P +G + L L + +N FS IPS + ++
Sbjct: 901 HCWVHWQALVHVNLGSNNMSGEIPNS-MGYLSQLESLLLDDNRFSGYIPSTLQNCSTMKF 1077
Query: 556 LDLGGNELSGKIPKELVELPNLRMLNLSRNKIEGIIPIKFD--SGLESLDLSGNFLKGNI 613
+D+G N+LS IP + E+ L +L L N G I K S L LD N L G+I
Sbjct: 1078IDMGNNQLSDTIPDWMWEMQYLMVLRLRSNNFNGSITQKMCQLSSLIVLDHGNNSLSGSI 1257
Query: 614 PTGLADLVRLSK--------------LNLSHNMLSGT---IPQN----FGRNLVFV---N 649
P L D+ ++ + S+N T +P+ + NL+ V +
Sbjct: 1258PNCLDDMKTMAGEDDFFANPSSYSYGSDFSYNHYKETLVLVPKKDELEYRDNLILVRMID 1437
Query: 650 ISDNQLEGPLPKIPAFLSASFESLKNNNHLCGNI 683
+S N+L G +P + LSA + NHL G I
Sbjct: 1438LSSNKLSGAIPSEISKLSALRFLNLSRNHLSGEI 1539
Score = 118 bits (296), Expect = 1e-26
Identities = 105/344 (30%), Positives = 162/344 (46%), Gaps = 51/344 (14%)
Frame = +1
Query: 61 IKCDKSNFI---STIGLANLGLKGTLHSLTFSSFPNLLMIDIRNNSFYGTI-PAQIG--N 114
++ D SN S I L++ KG L S++ N+ ++++ NNS GTI P G N
Sbjct: 673 LRGDLSNIFLNSSVINLSSNLFKGRLPSVS----ANVEVLNVANNSISGTISPFLCGKPN 840
Query: 115 LSN-ISILTFKNNYFDGSIPQEMCTLTGLQFLDISFCKLNGAIPKSIGNLTNLSYLILGG 173
+N +S+L F NN G + L +++ ++G IP S+G L+ L L+L
Sbjct: 841 ATNKLSVLDFSNNVLSGDLGHCWVHWQALVHVNLGSNNMSGEIPNSMGYLSQLESLLLDD 1020
Query: 174 NNWSG-----------------------GPIPPEIGKLNNLLHLAIQKSNLVGSIPQEIG 210
N +SG IP + ++ L+ L ++ +N GSI Q++
Sbjct: 1021 NRFSGYIPSTLQNCSTMKFIDMGNNQLSDTIPDWMWEMQYLMVLRLRSNNFNGSITQKMC 1200
Query: 211 FLTNLAYIDLSKNSLSGGIPETIGNLSKL--DTLVLSNNTKMSGPIPHSLWNMSSLTVL- 267
L++L +D NSLSG IP + ++ + + +N + S S + VL
Sbjct: 1201 QLSSLIVLDHGNNSLSGSIPNCLDDMKTMAGEDDFFANPSSYSYGSDFSYNHYKETLVLV 1380
Query: 268 -------YFDNI-----------GLSGSIPDSIQNLVNLKELALDINHLSGSIPSTIGDL 309
Y DN+ LSG+IP I L L+ L L NHLSG IP+ +G +
Sbjct: 1381 PKKDELEYRDNLILVRMIDLSSNKLSGAIPSEISKLSALRFLNLSRNHLSGEIPNDMGKM 1560
Query: 310 KNLIKLYLGSNNLSGPIPASIGNLINLQVLSVQENNLTGTIPAS 353
K L L L NN+SG IP S+ +L L L++ +NL+G IP S
Sbjct: 1561 KLLESLDLSLNNISGQIPQSLSDLSFLSFLNLSYHNLSGRIPTS 1692
Score = 98.2 bits (243), Expect = 2e-20
Identities = 77/270 (28%), Positives = 131/270 (48%), Gaps = 28/270 (10%)
Frame = +1
Query: 418 FTGPIPTSLKTCSSIERITLEVNQIEGDIAQDFGVYPKLQYLDLSDNKFHGQISPNWGKS 477
+ G IP + + +I+ + L+ NQ+ G + G L+ LDLS+N F I +
Sbjct: 19 YRGKIPQIISSLQNIKNLDLQNNQLSGPLPDSLGQLKHLEVLDLSNNTFTCPIPSPFANL 198
Query: 478 LNLQTFIISNNNISGVIPLDFIGLTKLGVLHLSSNQLTGKLPMEVLGGMKSLFDLKISNN 537
+L+T +++N ++G IP F L L VL+L +N LTG +P+ LG + +L L +S+N
Sbjct: 199 SSLRTLNLAHNRLNGTIPKSFEFLKNLQVLNLGANSLTGDVPV-TLGTLSNLVTLDLSSN 375
Query: 538 HFSDNI-PSEIGLLQRLQELDLGGNEL-----SG-------------------KIPKELV 572
+I S L L+EL L L SG K P+ L
Sbjct: 376 LLEGSIKESNFVKLFTLKELRLSWTNLFLSVNSGWAPPFQLEYVLLSSFGIGPKFPEWLK 555
Query: 573 ELPNLRMLNLSRNKIEGIIPIKF---DSGLESLDLSGNFLKGNIPTGLADLVRLSKLNLS 629
++++L +S+ I ++P F +E LDLS N L+G++ + S +NLS
Sbjct: 556 RQSSVKVLTMSKAGIADLVPSWFWIWTLQIEFLDLSNNLLRGDLSN---IFLNSSVINLS 726
Query: 630 HNMLSGTIPQNFGRNLVFVNISDNQLEGPL 659
N+ G +P + N+ +N+++N + G +
Sbjct: 727 SNLFKGRLP-SVSANVEVLNVANNSISGTI 813
>CO979898
Length = 869
Score = 173 bits (438), Expect = 4e-43
Identities = 89/207 (42%), Positives = 122/207 (57%), Gaps = 1/207 (0%)
Frame = -1
Query: 768 DDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRN 827
+ K ++G G G VY+ +L + +A+K+L+ T E K F E+E + IKHRN
Sbjct: 860 NSKDIIGSGGYGVVYELKLDDSTALAIKRLNRGTAER-----DKGFERELEAMADIKHRN 696
Query: 828 IIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHH 887
I+ LHG+ + ++ L+Y+ + GSLD L+ ++ DW R + G A +SYLHH
Sbjct: 695 IVTLHGYYTAPLYNLLIYELMPHGSLDSFLHGRSREKVLDWPTRYRIAAGAARGISYLHH 516
Query: 888 DCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPG-LHSWTQFAGTFGYAAPELAQT 946
DC P IIHRDI S N+LL+ + +A VSDFG A ++P H T AGTFGY APE T
Sbjct: 515 DCIPHIIHRDIKSSNILLDQNMDARVSDFGLATLMQPNKTHVSTIVAGTFGYLAPEYFDT 336
Query: 947 MEVNEKCDVYSFGVLALETIMGKHPGD 973
K DVYSFGV+ LE + GK P D
Sbjct: 335 GRATLKGDVYSFGVVLLELLTGKKPSD 255
>TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, partial (38%)
Length = 1009
Score = 170 bits (430), Expect = 4e-42
Identities = 102/289 (35%), Positives = 153/289 (52%), Gaps = 6/289 (2%)
Frame = +3
Query: 757 FENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSE 816
F + AT F + G G+V++ L +G V+AVK+ L + + K F SE
Sbjct: 12 FSELQLATGGFSQANFLAEGGFGSVHRGVLPDGQVIAVKQYKLASTQ-----GDKEFCSE 176
Query: 817 IETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVK 876
+E L+ +HRN++ L GFC LVY+++ GSLD + Q V +W R +
Sbjct: 177 VEVLSCAQHRNVVMLIGFCVDDGRRLLVYEYICNGSLDSHIYRRKQNV-LEWSARQKIAV 353
Query: 877 GVANALSYLHHDCSPP-IIHRDISSKNVLLNLDYEAHVSDFGTAKFLKPG-LHSWTQFAG 934
G A L YLH +C I+HRD+ N+LL D+EA V DFG A++ G + T+ G
Sbjct: 354 GAARGLRYLHEECRVGCIVHRDMRPNNILLTHDFEALVGDFGLARWQPDGDMGVETRVIG 533
Query: 935 TFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLI----SLFLSPSTRPMANNM 990
TFGY APE AQ+ ++ EK DVYSFG++ LE + G+ D+ LS RP+
Sbjct: 534 TFGYLAPEYAQSGQITEKADVYSFGIVLLELVTGRKAVDINRPKGQQCLSEWARPLLEKQ 713
Query: 991 LLTDVLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
++D + +D+EV + + + C+ ++P LRP M QV +ML
Sbjct: 714 ATYKLIDPSLRNCY--VDQEVYRMLKCSSLCIGRDPHLRPRMSQVLRML 854
>TC226549 UP|Q84P43 (Q84P43) Protein kinase Pti1, complete
Length = 1627
Score = 168 bits (425), Expect = 1e-41
Identities = 111/303 (36%), Positives = 155/303 (50%), Gaps = 12/303 (3%)
Frame = +3
Query: 755 MMFENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFM 814
+ + + E T NF K L+G GS G VY A L+ G VAVKKL + ++ E S+ F+
Sbjct: 372 LSLDELKEKTDNFGSKALIGEGSYGRVYYATLNNGKAVAVKKLDVSSEPE----SNNEFL 539
Query: 815 SEIETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDT------QAVAFDW 868
+++ ++ +K+ N ++LHG+C L Y+F GSL IL+ DW
Sbjct: 540 TQVSMVSRLKNGNFVELHGYCVEGNLRVLAYEFATMGSLHDILHGRKGVQGAQPGPTLDW 719
Query: 869 EKRVNVVKGVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDF---GTAKFLKPG 925
+RV + A L YLH PPIIHRDI S NVL+ DY+A ++DF A +
Sbjct: 720 IQRVRIAVDAARGLEYLHEKVQPPIIHRDIRSSNVLIFEDYKAKIADFNLSNQAPDMAAR 899
Query: 926 LHSWTQFAGTFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRP 985
LHS T+ GTFGY APE A T ++ +K DVYSFGV+ LE + G+ P D S
Sbjct: 900 LHS-TRVLGTFGYHAPEYAMTGQLTQKSDVYSFGVVLLELLTGRKPVDHTMPRGQQSLVT 1076
Query: 986 MANNMLLTDVLDQ--RPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML-AIG 1042
A L D + Q P+ E + V + +A C+ RP+M V K L +
Sbjct: 1077 WATPRLSEDKVKQCVDPKLKGEYPPKGVAKLGAVAALCVQYEAEFRPNMSIVVKALQPLL 1256
Query: 1043 KSP 1045
KSP
Sbjct: 1257 KSP 1265
>BM143950
Length = 426
Score = 166 bits (421), Expect = 4e-41
Identities = 77/142 (54%), Positives = 109/142 (76%)
Frame = +2
Query: 781 VYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTGIKHRNIIKLHGFCSHSKF 840
VYKAE+S G V AVKKL ++ ++ S KSF +EIE +T +HRNIIKL+GFC
Sbjct: 2 VYKAEMSGGQVFAVKKLKCDSNN-LNIESIKSFENEIEAMTKTRHRNIIKLYGFCCEGMH 178
Query: 841 SFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVKGVANALSYLHHDCSPPIIHRDISS 900
+FL+Y+++ G+L +L +D A+ DW KR++++KGV +ALSY+HHDC+PP+IHRD+SS
Sbjct: 179 TFLIYEYMNRGNLADMLRDDKDALELDWHKRIHIIKGVTSALSYMHHDCAPPLIHRDVSS 358
Query: 901 KNVLLNLDYEAHVSDFGTAKFL 922
KN+LL+ + +AHVSDFGTA+FL
Sbjct: 359 KNILLSSNLQAHVSDFGTARFL 424
>TC217441 homologue to GB|AAK68074.1|14573459|AF384970 somatic embryogenesis
receptor-like kinase 3 {Arabidopsis thaliana;} , partial
(54%)
Length = 1008
Score = 164 bits (415), Expect = 2e-40
Identities = 101/285 (35%), Positives = 150/285 (52%), Gaps = 8/285 (2%)
Frame = +2
Query: 763 ATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSEIETLTG 822
AT NF +K+++G G G VYK L++G +VAVK+L EE + F +E+E ++
Sbjct: 143 ATDNFSNKHILGRGGFGKVYKGRLADGSLVAVKRLK----EERTQGG*LQFQTEVEMISM 310
Query: 823 IKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVA-FDWEKRVNVVKGVANA 881
HRN+++L GFC LVY ++ GS+ L ++ W +R + G A
Sbjct: 311 AVHRNLLRLRGFCMTPTERLLVYPYMANGSVASCLRERQESQPPLGWPERKRIALGSARG 490
Query: 882 LSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKFLK-PGLHSWTQFAGTFGYAA 940
L+YLH C P IIHRD+ + N+LL+ ++EA V DFG AK + H T GT G+ A
Sbjct: 491 LAYLHDHCDPKIIHRDVKAANILLDEEFEAVVGDFGLAKLMDYKDTHVTTAVRGTIGHIA 670
Query: 941 PELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISL------FLSPSTRPMANNMLLTD 994
PE T + +EK DV+ +GV+ LE I G+ DL L L + + + L
Sbjct: 671 PEYLSTGKSSEKTDVFGYGVMLLELITGQRAFDLARLANDDDVMLLDWVKGLLKDRKLET 850
Query: 995 VLDQRPQQVMEPIDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
++D Q DEEV + ++A C +P RP M +V +ML
Sbjct: 851 LVDADLQGSYN--DEEVEQLIQVALLCTQGSPMERPKMSEVVRML 979
>TC209776 similar to UP|O65470 (O65470) Serine/threonine kinase-like protein ,
partial (47%)
Length = 1305
Score = 163 bits (412), Expect = 4e-40
Identities = 99/291 (34%), Positives = 148/291 (50%), Gaps = 8/291 (2%)
Frame = +2
Query: 757 FENIIEATANFDDKYLVGVGSQGNVYKAELSEGLVVAVKKLHLVTDEEMSCFSSKSFMSE 816
F I AT F + +G G G VYK L G VAVK+L ++ + + F +E
Sbjct: 50 FSTIEAATQKFSEANKLGEGGFGEVYKGLLPSGQEVAVKRLSKISGQ-----GGEEFKNE 214
Query: 817 IETLTGIKHRNIIKLHGFCSHSKFSFLVYKFLEGGSLDQILNNDTQAVAFDWEKRVNVVK 876
+E + ++HRN+++L GFC + LVY+F+ SLD IL + + + DW +R +V+
Sbjct: 215 VEIVAKLQHRNLVRLLGFCLEGEEKILVYEFVANKSLDYILFDPEKQKSLDWTRRYKIVE 394
Query: 877 GVANALSYLHHDCSPPIIHRDISSKNVLLNLDYEAHVSDFGTAKF--LKPGLHSWTQFAG 934
G+A + YLH D IIHRD+ + NVLL+ D +SDFG A+ + + + G
Sbjct: 395 GIARGIQYLHEDSRLKIIHRDLKASNVLLDGDMNPKISDFGMARIFGVDQTQANTNRIVG 574
Query: 935 TFGYAAPELAQTMEVNEKCDVYSFGVLALETIMGKHPGDLISLFLSPSTRPMANNMLLTD 994
T+GY +PE A E + K DVYSFGVL LE I GK ++ A +
Sbjct: 575 TYGYMSPEYAMHGEYSAKSDVYSFGVLILEIISGKRNSSFYETDVAEDLLSYAWKLW--- 745
Query: 995 VLDQRPQQVMEP------IDEEVILIARLAFACLSQNPRLRPSMGQVCKML 1039
D+ P ++M+ EVI + C+ ++P RP+M V ML
Sbjct: 746 -KDEAPLELMDQSLRESYTRNEVIRCIHIGLLCVQEDPIDRPTMASVVLML 895
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.138 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 47,740,934
Number of Sequences: 63676
Number of extensions: 682613
Number of successful extensions: 7760
Number of sequences better than 10.0: 1096
Number of HSP's better than 10.0 without gapping: 4433
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5848
length of query: 1067
length of database: 12,639,632
effective HSP length: 107
effective length of query: 960
effective length of database: 5,826,300
effective search space: 5593248000
effective search space used: 5593248000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC133779.11


