

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133779.1 + phase: 0 /pseudo
(611 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
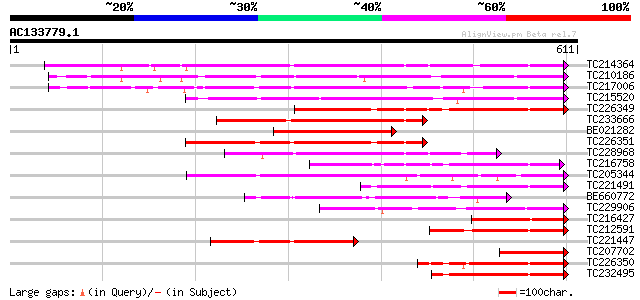
Score E
Sequences producing significant alignments: (bits) Value
TC214364 UP|Q6WNU4 (Q6WNU4) Subtilisin-like protease, complete 426 e-119
TC210186 UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, complete 281 5e-76
TC217006 UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1, complete 276 1e-74
TC215520 GB|AAG38994.1|11611651|AF160513 subtilisin-type proteas... 253 2e-67
TC226349 similar to PIR|JC7519|JC7519 subtilisin-like serine pro... 219 3e-57
TC233666 weakly similar to UP|Q9FJF3 (Q9FJF3) Serine protease-li... 216 2e-56
BE021282 similar to GP|21593457|gb| subtilisin-like serine prote... 197 9e-51
TC226351 similar to PIR|JC7519|JC7519 subtilisin-like serine pro... 196 3e-50
TC228968 similar to GB|BAB09208.1|9758669|AB012245 subtilisin-li... 187 9e-48
TC216758 similar to PIR|T05768|T05768 subtilisin-like proteinase... 185 4e-47
TC205344 similar to UP|Q9SZV5 (Q9SZV5) Proteinase-like protein (... 185 4e-47
TC221491 similar to UP|P93204 (P93204) SBT1 protein (Subtilisin-... 157 2e-38
BE660772 152 5e-37
TC229906 weakly similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like pr... 145 4e-35
TC216427 similar to UP|O82440 (O82440) Subtilisin-like protease ... 145 5e-35
TC212591 similar to UP|Q9FJF3 (Q9FJF3) Serine protease-like prot... 142 6e-34
TC221447 similar to UP|Q75I27 (Q75I27) Putaive subtilisin-like p... 136 2e-32
TC207702 weakly similar to UP|Q8S896 (Q8S896) Subtilisin-like se... 134 2e-31
TC226350 similar to PIR|JC7519|JC7519 subtilisin-like serine pro... 133 3e-31
TC232495 similar to UP|Q8W554 (Q8W554) AT3g14240/MLN21_2, partia... 125 5e-29
>TC214364 UP|Q6WNU4 (Q6WNU4) Subtilisin-like protease, complete
Length = 2667
Score = 426 bits (1094), Expect = e-119
Identities = 255/584 (43%), Positives = 342/584 (57%), Gaps = 19/584 (3%)
Frame = +1
Query: 38 SYLFLQCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQIN 97
S+ + Y+VYLGAH HGP SSVD T SH+D LGS LGS A+++I YSY + IN
Sbjct: 106 SFAVKKSYVVYLGAHSHGPELSSVDFNQVTQSHHDFLGSFLGSSNTAKDSIFYSYTRHIN 285
Query: 98 GFAAILEEEEAAQLANCTQHVL-----GNSLD*VQMMSTQLGKREGLVKIPL*LTLIQVF 152
GFAA L+EE A ++A + + G L + + G+++ + F
Sbjct: 286 GFAATLDEEVAVEIAKHPKVLSVFENRGRKLHTTRSWDFMELEHNGVIQSSSIWKKAR-F 462
Query: 153 GR-----NRRVLTTEE*VQFH*DGVGILLRKFHVTVYLNGLP------RKLIGARFFNKA 201
G N E F G+G + K+ + NG+ RKLIGAR+FNK
Sbjct: 463 GEGVIIGNLDTGVWPESKSFSEQGLGPIPSKWRGICH-NGIDHTFHCNRKLIGARYFNKG 639
Query: 202 YEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYK 261
Y + G L SS + RD G GTHTLSTAGGN V ++FG G+GT KGGSP +RVA YK
Sbjct: 640 YASVAGPLNSSFDSPRDNEGHGTHTLSTAGGNMVARVSVFGQGHGTAKGGSPMARVAAYK 819
Query: 262 ACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALA 321
CW +CF AD+LAA D AI+DG D++SVS GG +T F D ++IG+FHA
Sbjct: 820 VCWPPVAGDECFDADILAAFDLAIHDGVDVLSVSLGGSSST----FFKDSVAIGSFHAAK 987
Query: 322 RNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTINNK-TLTGASLFVNLP 380
R +++V SAGN GP + N+APW TVAAST+DR F + + + N T G SL
Sbjct: 988 RGVVVVCSAGNSGPAEATAENLAPWHVTVAASTMDRQFPTYVVLGNDITFKGESLSATKL 1167
Query: 381 PNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALS 440
++ + II +TDAK A+ DA C+ GTLDP+K GK+V C R G + +G++A
Sbjct: 1168AHKFYPIIKATDAKLASARAEDAVLCQNGTLDPNKAKGKIVVCLR-GINARVDKGEQAFL 1344
Query: 441 AGAVGVIMRNQPEVDGKTLLAEPHVV--STINYYDARSITTPKGSEITPEDIKTNATIRM 498
AGAVG+++ N + G ++A+PHV+ S IN+ D ++ S P T+ ++
Sbjct: 1345AGAVGMVLAND-KTTGNEIIADPHVLPASHINFTDGSAVFKYINSTKFPVAYITHPKTQL 1521
Query: 499 SPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNR 558
KPAP MA+FSS+GPN + P ILKPD+TAPGV+++AAY+ +N V D R
Sbjct: 1522D-------TKPAPFMAAFSSKGPNTIVPEILKPDITAPGVSVIAAYTEAQGPTNQVFDKR 1680
Query: 559 RGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
R PFN GTSMSCPHV G GL++ L+P WS AAIKSAIMTT
Sbjct: 1681R-IPFNSVSGTSMSCPHVSGIVGLLRALYPTWSTAAIKSAIMTT 1809
>TC210186 UP|Q93WQ0 (Q93WQ0) Subtilisin-type protease, complete
Length = 2401
Score = 281 bits (720), Expect = 5e-76
Identities = 202/581 (34%), Positives = 305/581 (51%), Gaps = 21/581 (3%)
Frame = +1
Query: 43 QCYIVYLGAHVHGPTPSSVDLETATY--SHYDLLGSILGSHEEAEEAIIYSYNKQINGFA 100
+ YIVY+GA D A++ H +L S+L +E A ++ +Y +GFA
Sbjct: 175 EVYIVYMGA---------ADSTDASFRNDHAQVLNSVLRRNENA---LVRNYKHGFSGFA 318
Query: 101 AILEEEEAAQLANCTQHVL---GNSLD*VQMMSTQLGKREGLVKIPL*LTLIQVFGRNRR 157
A L ++EA +A V G L S K + VKI +
Sbjct: 319 ARLSKKEATSIAQKPGVVSVFPGPVLKLHTTRSWDFLKYQTQVKIDTKPNAVSKSSSVIG 498
Query: 158 VLTT---EE*VQFH*DGVGILLRKFHVTV------YLNGLPRKLIGARFFNKAYEAFHGK 208
+L T E F G+G + ++ T Y + RKLIGAR++ +
Sbjct: 499 ILDTGIWPEAASFSDKGMGPVPSRWKGTCMKSQDFYSSNCNRKLIGARYYADPND----- 663
Query: 209 LPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTD 268
S TARD G GTH TA G V NA+ +G+ G KGGSP SR+A Y+ C +
Sbjct: 664 --SGDNTARDSNGHGTHVAGTAAGVMVTNASYYGVATGCAKGGSPESRLAVYRVCSNF-- 831
Query: 269 VVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVA 328
C G+ +LAA D AI DG DL+SVS G P++ +D IS+GAFHA+ IL+V
Sbjct: 832 --GCRGSSILAAFDDAIADGVDLLSVSLGASTGFRPDLT-SDPISLGAFHAMEHGILVVC 1002
Query: 329 SAGNEGPTPGSVTNVAPWVFTVAASTLDRDF-SSVMTINNKTLTGASLFVNLPP---NQD 384
SAGN+GP+ ++ N APW+ TVAAST+DR+F S+++ +NK + G + +NL P +
Sbjct: 1003SAGNDGPSSYTLVNDAPWILTVAASTIDRNFLSNIVLGDNKIIKGKA--INLSPLSNSPK 1176
Query: 385 FLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVAC-DREGKINSIAEGQEALSAGA 443
+ +I AK + + V+A+ C P +LD +KV GK+V C D+ K ++ + + G
Sbjct: 1177YPLIYGESAKANSTSLVEARQCHPNSLDGNKVKGKIVVCDDKNDKYSTRKKVATVKAVGG 1356
Query: 444 VGVI-MRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPE-DIKTNATIRMSPA 501
+G++ + +Q E + S + A I++ G I + +N +
Sbjct: 1357IGLVHITDQNEA----------IASNYGDFPATVISSKDGVTILQYINSTSNPVATILAT 1506
Query: 502 NALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGF 561
++ KPAP++ +FSSRGP+ + ILKPD+ APGVNILAA+ + + + +V ++
Sbjct: 1507TSVLDYKPAPLVPNFSSRGPSSLSSNILKPDIAAPGVNILAAW--IGNGTEVVPKGKKPS 1680
Query: 562 PFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
+ I GTSM+CPHV G A +KT +P WS ++IKSAIMT+
Sbjct: 1681LYKIISGTSMACPHVSGLASSVKTRNPTWSASSIKSAIMTS 1803
>TC217006 UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1, complete
Length = 2422
Score = 276 bits (707), Expect = 1e-74
Identities = 209/580 (36%), Positives = 293/580 (50%), Gaps = 19/580 (3%)
Frame = +3
Query: 42 LQCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAA 101
L+ YIVY G ++ D +A + +L + S+ E + + + + + +GF A
Sbjct: 111 LKSYIVYTGNSMN-------DEASALTLYSSMLQEVADSNAEPK-LVQHHFKRSFSGFVA 266
Query: 102 ILEEEEAAQLANCTQHVLGNSLD*VQMMSTQLGKREGLVKIPL*LT--------LIQVFG 153
+L EEEA ++A + V Q+ +T + + PL +I VF
Sbjct: 267 MLTEEEADRMARHDRVVAVFPNKKKQLHTT---RSWDFIGFPLQANRAPAESDVIIAVFD 437
Query: 154 RNRRVLTTEE*VQFH*DGVGILLRKFHVTVYLNG---LPRKLIGARFFNKAYEAFHGKLP 210
E F+ G G K+ T + K+IGA+ + + F K
Sbjct: 438 SG----IWPESESFNDKGFGPPPSKWKGTCQTSKNFTCNNKIIGAKIYK--VDGFFSK-- 593
Query: 211 SSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVV 270
++ RD G GTH STA GN V A++ G+G GT +GG ++R+A YK CW
Sbjct: 594 DDPKSVRDIDGHGTHVASTAAGNPVSTASMLGLGQGTSRGGVTKARIAVYKVCW----FD 761
Query: 271 DCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASA 330
C AD+LAA D AI DG D+I+VS GG + N F D I+IGAFHA+ +L V SA
Sbjct: 762 GCTDADILAAFDDAIADGVDIITVSLGGFSDEN---YFRDGIAIGAFHAVRNGVLTVTSA 932
Query: 331 GNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTINNK-TLTGASLFVNLPPNQDFLIII 389
GN GP P S++N +PW +VAAST+DR F + + + NK T G S+ + + II
Sbjct: 933 GNSGPRPSSLSNFSPWSISVAASTIDRKFVTKVELGNKITYEGTSINTFDLKGELYPIIY 1112
Query: 390 STDA--KFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVI 447
DA K + +++C G+LD V GK+V C+ K AGAVG +
Sbjct: 1113GGDAPNKGEGIDGSSSRYCSSGSLDKKLVKGKIVLCESRSK------ALGPFDAGAVGAL 1274
Query: 448 MRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITP-----EDIKTNATIRMSPAN 502
++ Q D L P S + D S+ S TP + +T TI
Sbjct: 1275IQGQGFRDLPPSLPLPG--SYLALQDGASVYDYINSTRTPIATIFKTDETKDTI------ 1430
Query: 503 ALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFP 562
APV+ASFSSRGPN V P ILKPD+ APGV+ILA++S + S++ DNR
Sbjct: 1431-------APVVASFSSRGPNIVTPEILKPDLVAPGVSILASWSPASPPSDVEGDNRT-LN 1586
Query: 563 FNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
FNI GTSM+CPHV G A +K+ HP WSPAAI+SA+MTT
Sbjct: 1587FNIISGTSMACPHVSGAAAYVKSFHPTWSPAAIRSALMTT 1706
>TC215520 GB|AAG38994.1|11611651|AF160513 subtilisin-type protease precursor
{Glycine max;} , complete
Length = 2392
Score = 253 bits (645), Expect = 2e-67
Identities = 167/420 (39%), Positives = 237/420 (55%), Gaps = 7/420 (1%)
Frame = +2
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIK 249
RK+IGARF+ E +TARDF G GTH STA G V A+ +G+ GT +
Sbjct: 605 RKIIGARFYPNPEE----------KTARDFNGHGTHVSSTAVGVPVSGASFYGLAAGTAR 754
Query: 250 GGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFT 309
GGSP SR+A YK C + C G+ +LA D AI+DG D++S+S GG T + + T
Sbjct: 755 GGSPESRLAVYKVCGAFG---SCPGSAILAGFDDAIHDGVDILSLSLGGFGGTKTD-LTT 922
Query: 310 DEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDF-SSVMTINNK 368
D I+IGAFH++ R IL+V +AGN+G P +V N APW+ TVAAST+DRD S V+ NN+
Sbjct: 923 DPIAIGAFHSVQRGILVVCAAGNDG-EPFTVLNDAPWILTVAASTIDRDLQSDVVLGNNQ 1099
Query: 369 TLTGASL-FVNLPPNQDFLIIISTDAKFANVTDV-DAQFCRPGTLDPSKVNGKVVACDRE 426
+ G ++ F L + D+ +I + A AN++++ DA+ C P +LDP KV GK+V CD +
Sbjct: 1100VVKGRAINFSPLLNSPDYPMIYAESAARANISNITDARQCHPDSLDPKKVIGKIVVCDGK 1279
Query: 427 GKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPK---GS 483
I + + + G+ + + + G S YY +T K G
Sbjct: 1280NDIYYSTDEKIVIVKALGGIGLVHITDQSG----------SVAFYYVDFPVTEVKSKHGD 1429
Query: 484 EITPEDIKTNATIRMSPAN-ALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILA 542
I T+ + A + KPAP + FSSRGP+ + +LKPD+ APGVNILA
Sbjct: 1430AILQYINSTSHPVGTILATVTIPDYKPAPRVGYFSSRGPSLITSNVLKPDIAAPGVNILA 1609
Query: 543 AYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
A+ + ++ V R+ + I GTSM+ PHV G A +K +P WS +AIKSAIMT+
Sbjct: 1610AW--FGNDTSEVPKGRKPSLYRILSGTSMATPHVSGLACSVKRKNPTWSASAIKSAIMTS 1783
Score = 34.7 bits (78), Expect = 0.13
Identities = 25/70 (35%), Positives = 38/70 (53%)
Frame = +2
Query: 43 QCYIVYLGAHVHGPTPSSVDLETATYSHYDLLGSILGSHEEAEEAIIYSYNKQINGFAAI 102
+ YIVY+GA T +S+ E H +L S+L +E A ++ +Y +GFAA
Sbjct: 143 EVYIVYMGAA--DSTKASLKNE-----HAQILNSVLRRNENA---LVRNYKHGFSGFAAR 292
Query: 103 LEEEEAAQLA 112
L +EEA +A
Sbjct: 293 LSKEEANSIA 322
>TC226349 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(56%)
Length = 1773
Score = 219 bits (558), Expect = 3e-57
Identities = 128/300 (42%), Positives = 182/300 (60%), Gaps = 5/300 (1%)
Frame = +1
Query: 308 FTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI-N 366
+ D ++IGAF A+ + IL+ SAGN GP P S++NVAPW+ TV A TLDRDF + + + N
Sbjct: 7 YRDSVAIGAFSAMEKGILVSCSAGNSGPGPYSLSNVAPWITTVGAGTLDRDFPAYVALGN 186
Query: 367 NKTLTGASLF-VNLPPNQDFLIIISTDAKFANVTD--VDAQFCRPGTLDPSKVNGKVVAC 423
+G SL+ N P+ ++ + NV++ ++ C GTL P KV GK+V C
Sbjct: 187 GLNFSGVSLYRGNALPDSSLPLVYA-----GNVSNGAMNGNLCITGTLSPEKVAGKIVLC 351
Query: 424 DREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGS 483
DR G + +G SAGA+G+++ N +G+ L+A+ H++ A ++ G
Sbjct: 352 DR-GLTARVQKGSVVKSAGALGMVLSN-TAANGEELVADAHLL------PATAVGQKAGD 507
Query: 484 EITPEDIK-TNATIRMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILA 542
I + T+++ G +P+PV+A+FSSRGPN + P ILKPD+ APGVNILA
Sbjct: 508 AIKKYLVSDAKPTVKIFFEGTKVGIQPSPVVAAFSSRGPNSITPQILKPDLIAPGVNILA 687
Query: 543 AYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
+S + L DNRR FNI GTSMSCPHV G A LIK+ HP+WSPAA++SA+MTT
Sbjct: 688 GWSKAVGPTGLPVDNRR-VDFNIISGTSMSCPHVSGLAALIKSAHPDWSPAAVRSALMTT 864
>TC233666 weakly similar to UP|Q9FJF3 (Q9FJF3) Serine protease-like protein,
partial (30%)
Length = 678
Score = 216 bits (551), Expect = 2e-56
Identities = 114/229 (49%), Positives = 150/229 (64%), Gaps = 1/229 (0%)
Frame = +1
Query: 223 GTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGADVLAAID 282
G+HTLST GG FV A +FG+GNGT +GGSPR+RVATYK CW D +CF AD++AA D
Sbjct: 4 GSHTLSTIGGTFVPGANVFGLGNGTAEGGSPRARVATYKVCWPPIDGNECFDADIMAAFD 183
Query: 283 QAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTN 342
AI+DG D++S+S GG N F D +SIGAFHA + I ++ SAGN GPTP +V N
Sbjct: 184 MAIHDGVDVLSLSLGG----NATDYFDDGLSIGAFHANMKGIPVICSAGNYGPTPATVFN 351
Query: 343 VAPWVFTVAASTLDRDFSSVMTINN-KTLTGASLFVNLPPNQDFLIIISTDAKFANVTDV 401
VAPW+ TV ASTLDR F SV+ ++N + GASL +P ++ + +I + DAK AN
Sbjct: 352 VAPWILTVGASTLDRQFDSVVELHNGQRFMGASLSKAMPEDKLYPLINAADAKAANKPVE 531
Query: 402 DAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRN 450
+A C GT+DP K GK++ C R G + + AL AGA +I+ N
Sbjct: 532 NATLCMRGTIDPEKARGKILVCLR-GVTARVEKSLVALEAGAAXMILCN 675
>BE021282 similar to GP|21593457|gb| subtilisin-like serine protease
{Arabidopsis thaliana}, partial (5%)
Length = 402
Score = 197 bits (502), Expect = 9e-51
Identities = 96/132 (72%), Positives = 112/132 (84%)
Frame = +1
Query: 285 IYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVA 344
I DG D+IS+SAGG PE IFTDE+SIGAFHA+ARN +LVASAGN+GPTPG+V NVA
Sbjct: 1 IDDGVDIISLSAGGSYVVTPEGIFTDEVSIGAFHAIARNRILVASAGNDGPTPGTVLNVA 180
Query: 345 PWVFTVAASTLDRDFSSVMTINNKTLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQ 404
PWVFT+AASTLDRDFSS +TINN+ +TGASLFVNLPPN+ F +I++TDAK AN T DA+
Sbjct: 181 PWVFTIAASTLDRDFSSNLTINNRQITGASLFVNLPPNKAFSLILATDAKLANATFRDAE 360
Query: 405 FCRPGTLDPSKV 416
CRPGTLDP KV
Sbjct: 361 LCRPGTLDPEKV 396
>TC226351 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(48%)
Length = 1490
Score = 196 bits (497), Expect = 3e-50
Identities = 113/267 (42%), Positives = 164/267 (61%), Gaps = 6/267 (2%)
Frame = +3
Query: 190 RKLIGARFFNKAYEAFHGKLPSSQQT--ARDFVGPGTHTLSTAGGNFVQNATIFGIGNGT 247
RKLIGARFF+K EA G + ++++ ARD G GTHT STA G+ V +A++FG +GT
Sbjct: 732 RKLIGARFFSKGVEAILGPINETEESRSARDDDGHGTHTASTAAGSVVSDASLFGYASGT 911
Query: 248 IKGGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVI 307
+G + R+RVA YK CW CF +D+LAAI++AI D +++S+S GG +
Sbjct: 912 ARGMATRARVAAYKVCWK----GGCFSSDILAAIERAILDNVNVLSLSLGGGMSD----Y 1067
Query: 308 FTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTI-N 366
+ D ++IGAF A+ IL+ SAGN GP+P S++NVAPW+ TV A TLDRDF + + + N
Sbjct: 1068YRDSVAIGAFSAMENGILVSCSAGNAGPSPYSLSNVAPWITTVGAGTLDRDFPAYVALGN 1247
Query: 367 NKTLTGASLF-VNLPPNQDFLIIISTDAKFANVTD--VDAQFCRPGTLDPSKVNGKVVAC 423
+G SL+ N P+ + + NV++ ++ C GTL P KV GK+V C
Sbjct: 1248GLNFSGVSLYRGNAVPDSPLPFVYA-----GNVSNGAMNGNLCITGTLSPEKVAGKIVLC 1412
Query: 424 DREGKINSIAEGQEALSAGAVGVIMRN 450
DR G + +G SAGA+G+++ N
Sbjct: 1413DR-GLTARVQKGSVVKSAGALGMVLSN 1490
>TC228968 similar to GB|BAB09208.1|9758669|AB012245 subtilisin-like protease
{Arabidopsis thaliana;} , partial (20%)
Length = 1008
Score = 187 bits (476), Expect = 9e-48
Identities = 115/305 (37%), Positives = 167/305 (54%), Gaps = 6/305 (1%)
Frame = +3
Query: 232 GNFVQNAT-IFGIGNGTIKGGSPRSRVATYKACWSLTDVVD-----CFGADVLAAIDQAI 285
G V NA+ I G GT GG+P +R+A YKACW + C D+L AID AI
Sbjct: 12 GRVVPNASAIGGFAKGTALGGAPLARLAIYKACWPIKGKSKHEGNICTNIDMLKAIDDAI 191
Query: 286 YDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAP 345
DG D++S+S G + E D I+ GA HA+ +NI++V SAGN GP P +++N AP
Sbjct: 192 GDGVDVLSISIGFSAPISYE---EDVIARGALHAVRKNIVVVCSAGNSGPLPQTLSNPAP 362
Query: 346 WVFTVAASTLDRDFSSVMTINNKTLTGASLFVNLPPNQDFLIIISTDAKFANVTDVDAQF 405
W+ TVAAST+DR F + + +N + G S+ N + ++++ D + + ++ F
Sbjct: 363 WIITVAASTVDRSFHAPIKLNGTIIEGRSITPLHMGNSFYPLVLARDVEHPGLPSNNSGF 542
Query: 406 CRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHV 465
C TL P+K GK+V C R G+ + +G E AG VG I+ N +++GK + ++PH
Sbjct: 543 CLDNTLQPNKARGKIVLCMR-GQGERLKKGLEVQRAGGVGFILGNN-KLNGKDVPSDPHF 716
Query: 466 VSTINYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKVQ 525
+ S+ + TP N ++ P + KPAP MASFSSRGPN V
Sbjct: 717 IPATGVSYENSLKLIQYVHSTP-----NPMAQILPGTTVLETKPAPSMASFSSRGPNIVD 881
Query: 526 PYILK 530
P ILK
Sbjct: 882 PNILK 896
>TC216758 similar to PIR|T05768|T05768 subtilisin-like proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(35%)
Length = 796
Score = 185 bits (470), Expect = 4e-47
Identities = 114/276 (41%), Positives = 161/276 (58%), Gaps = 2/276 (0%)
Frame = +3
Query: 324 ILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDF-SSVMTINNKTLTGASLFVNLPPN 382
+ + +SAGN+GP+ SVTN+APW+ TV A T+DRDF S V+ + + L+G SL+
Sbjct: 6 VFVSSSAGNDGPSGMSVTNLAPWLTTVGAGTIDRDFPSQVILGDGRRLSGVSLYAGAALK 185
Query: 383 QDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAG 442
++ K + D C +LDP+ V GK+V CDR G +A+G AG
Sbjct: 186 GKMYQLVYP-GKSGILGD---SLCMENSLDPNMVKGKIVICDR-GSSPRVAKGLVVKKAG 350
Query: 443 AVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPE-DIKTNATIRMSPA 501
VG+I+ N +G+ L+ + H++ A ++ +G I TN T +
Sbjct: 351 GVGMILANGIS-NGEGLVGDAHLLP------ACAVGANEGDVIKKYISSSTNPTATLDFK 509
Query: 502 NALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGF 561
+ G KPAPV+ASFS+RGPN + P ILKPD APGVNILAA++ + L +D RR
Sbjct: 510 GTILGIKPAPVIASFSARGPNGLNPQILKPDFIAPGVNILAAWTQAVGPTGLDSDTRR-T 686
Query: 562 PFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKS 597
FNI GTSM+CPHV G A L+K+ HP+WSPAA++S
Sbjct: 687 EFNILSGTSMACPHVSGAAALLKSAHPDWSPAALRS 794
>TC205344 similar to UP|Q9SZV5 (Q9SZV5) Proteinase-like protein (AT4g30020
protein) (AT4g30020/F6G3_50), partial (78%)
Length = 2192
Score = 185 bits (470), Expect = 4e-47
Identities = 146/438 (33%), Positives = 216/438 (48%), Gaps = 26/438 (5%)
Frame = +2
Query: 191 KLIGARFFNKAYEAFHGKLPSSQ-QTARDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIK 249
K++GA+ F +A A P + D G G+HT S A G + G G
Sbjct: 137 KIVGAQHFAQAAIAAGAFNPFIVFDSPLDGDGHGSHTASIAAGRNGIPVRMHGHEFGKAS 316
Query: 250 GGSPRSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAG-GKPNTNPEVIF 308
G +PR+R+A YKA + L F ADV+AAIDQA++DG D++S+S G P +N + F
Sbjct: 317 GMAPRARIAVYKALYRL---FGGFIADVVAAIDQAVHDGVDILSLSVGPNSPPSNTKTTF 487
Query: 309 TDEISIGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDFSSVMTINN- 367
+ A+ + + +AGN GP P S+ + +PW+ TVAA+ DR + + + + N
Sbjct: 488 LNPFDATLLGAVKAGVFVAQAAGNGGPFPKSLVSYSPWIATVAAAIDDRRYKNHLILGNG 667
Query: 368 KTLTGASLFVNLPPNQDFLIIISTDAKF-ANVTDVDAQFC-RPGTLDPSKVNGKVVACDR 425
K L G L + NQ + ++ +TD ++VT C RP L+ + + G ++ C
Sbjct: 668 KILAGLGLSPSTRLNQTYTLVAATDVLLDSSVTKYSPTDCQRPELLNKNLIKGNILLCGY 847
Query: 426 E-------GKINSIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDAR--- 475
I ++E +AL GAVG ++ + G P + I DA
Sbjct: 848 SYNFVIGSASIKQVSETAKAL--GAVGFVLYVENVSPGTKFDPVPVGIPGILITDASKSK 1021
Query: 476 ------SITTPKGSEITPEDIKTNATIRMSPANALNGRKPAPVMASFSSRGPNKV----- 524
+I+TP+ + + I L+ K AP +A FS+RGPN
Sbjct: 1022ELIDYYNISTPRDWTGRVKTFEGTGKIEDGLMPILH--KSAPQVAMFSARGPNIKDFSFQ 1195
Query: 525 QPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIK 584
+ +LKPD+ APG I AA+SL + N G F + GTSM+ PH+ G A LIK
Sbjct: 1196EADLLKPDILAPGSLIWAAWSL----NGTDEPNYAGEGFAMISGTSMAAPHIAGIAALIK 1363
Query: 585 TLHPNWSPAAIKSAIMTT 602
HP+WSPAAIKSA+MTT
Sbjct: 1364QKHPHWSPAAIKSALMTT 1417
>TC221491 similar to UP|P93204 (P93204) SBT1 protein (Subtilisin-like
protease) , partial (34%)
Length = 872
Score = 157 bits (396), Expect = 2e-38
Identities = 95/224 (42%), Positives = 126/224 (55%)
Frame = +3
Query: 379 LPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEA 438
LPPN I+ + ANV+D C GTL KV GK+V CDR G + +G
Sbjct: 15 LPPNSPLPIVYA-----ANVSDESQNLCTRGTLIAEKVAGKIVICDRGGNAR-VEKGLVV 176
Query: 439 LSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPEDIKTNATIRM 498
SAG +G+I+ N + G+ L+A+ +++ S K +P N T ++
Sbjct: 177 KSAGGIGMILSNNEDY-GEELVADSYLLPAAALGQKSSNELKKYVFSSP-----NPTAKL 338
Query: 499 SPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNR 558
G +P+PV+A+FSSRGPN + P ILKPD+ APGVNILA ++ + L D R
Sbjct: 339 GFGGTQLGVQPSPVVAAFSSRGPNVLTPKILKPDLIAPGVNILAGWTGAVGPTGLTEDTR 518
Query: 559 RGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
FNI GTSMSCPHV G A L+K HP WSPAAI+SA+MTT
Sbjct: 519 H-VEFNIISGTSMSCPHVTGLAALLKGTHPEWSPAAIRSALMTT 647
>BE660772
Length = 826
Score = 152 bits (383), Expect = 5e-37
Identities = 102/300 (34%), Positives = 156/300 (52%), Gaps = 13/300 (4%)
Frame = +2
Query: 254 RSRVATYKACWSLTDVVDCFGADVLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEIS 313
++R+A YK CW L CF +D+LAA+D+A+ DG +IS+S G P+ + D I+
Sbjct: 2 KARIAAYKICWKL----GCFDSDILAAMDEAVSDGVHVISLSVGSSGYA-PQY-YRDSIA 163
Query: 314 IGAFHALARNILLVASAGNEGPTPGSVTNVAPWVFTVAASTLDRDF-SSVMTINNKTLTG 372
+GAF A N+L+ SAGN GP P + N+APW+ TV AST+DR+F + V+ + + G
Sbjct: 164 VGAFGAAKHNVLVSCSAGNSGPGPSTAVNIAPWILTVGASTVDREFPADVILGDGRVFGG 343
Query: 373 ASLFV--NLPPNQDFLIIISTDAKFANVTDVDAQFCRPGTLDPSKVNGKVVACDREGKIN 430
SL+ +LP DF + D +++C G+L+ SKV GK+ CDR G
Sbjct: 344 VSLYYGESLP---DFKL------PLVYAKDCGSRYCYIGSLESSKVQGKIXVCDRGGNAR 496
Query: 431 SIAEGQEALSAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITPEDI 490
+ +G G +G+IM N K+ L + + + ++ + P++I
Sbjct: 497 -VEKGSAVKLTGGLGMIMGNT*SQR*KSFL-QMRIF*QLLWW------------VKPQEI 634
Query: 491 KTNATIRMSPAN----------ALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNI 540
+ + +S L G P +ASFSSRGPN ++ ILKP+V GVNI
Sbjct: 635 RLRSXXSLSQVPNSNDXI*XNWELGGLXLXPXVASFSSRGPNHLRWKILKPEVLVLGVNI 814
>TC229906 weakly similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1,
partial (40%)
Length = 1392
Score = 145 bits (367), Expect = 4e-35
Identities = 96/275 (34%), Positives = 138/275 (49%), Gaps = 6/275 (2%)
Frame = +2
Query: 334 GPTPGSVTNVAPWVFTVAASTLDRDF-SSVMTINNKTLTGASLFVNLPPNQDFLIIISTD 392
GP S+T +PW+ +VAAST+ R F + V N G S+ N+ F ++ + D
Sbjct: 5 GPGLSSITTYSPWILSVAASTIGRKFLTKVQLGNGMVFEGVSINTFDLKNKMFPLVYAGD 184
Query: 393 AKFANVTD----VDAQFCRPGTLDPSKVNGKVVACDREGKINSIAEGQEALSAGAVGVIM 448
N D ++FC ++D V GK+V CD + + +GA G+++
Sbjct: 185 VP--NTADGYNSSTSRFCYVNSVDKHLVKGKIVLCDGNASPKKVGD-----LSGAAGMLL 343
Query: 449 RNQPEVDGKTLLAEPHV-VSTINYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGR 507
D A P +S N+ S N+T + ++ N
Sbjct: 344 GATDVKDAPFTYALPTAFISLRNFKLIHSYMVSL----------RNSTATIFRSDEDNDD 493
Query: 508 KPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQ 567
P + SFSSRGPN + P LKPD+ APGVNILAA+S + ++S D +R +NI+
Sbjct: 494 SQTPFIVSFSSRGPNPLTPNTLKPDLAAPGVNILAAWSPVYTISEFKGD-KRAVQYNIES 670
Query: 568 GTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
GTSM+CPHV A +K+ HPNWSPA IKSA+MTT
Sbjct: 671 GTSMACPHVSAAAAYVKSFHPNWSPAMIKSALMTT 775
>TC216427 similar to UP|O82440 (O82440) Subtilisin-like protease (Fragment),
partial (15%)
Length = 689
Score = 145 bits (366), Expect = 5e-35
Identities = 71/105 (67%), Positives = 83/105 (78%)
Frame = +2
Query: 498 MSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDN 557
+S A G KPAP++A FSSRGP+ VQP ILKPD+TAPGVN++AA++ A SNL +D
Sbjct: 143 LSAAETYIGVKPAPIIAGFSSRGPSSVQPLILKPDITAPGVNVIAAFTQGAGPSNLPSDR 322
Query: 558 RRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
RR FN+QQGTSMSCPHV G AGL+KT HP WSPAAIKSAIMTT
Sbjct: 323 RRSL-FNVQQGTSMSCPHVAGIAGLLKTYHPTWSPAAIKSAIMTT 454
>TC212591 similar to UP|Q9FJF3 (Q9FJF3) Serine protease-like protein, partial
(19%)
Length = 463
Score = 142 bits (357), Expect = 6e-34
Identities = 81/152 (53%), Positives = 95/152 (62%), Gaps = 2/152 (1%)
Frame = +1
Query: 453 EVDGKTLLAEPHVV--STINYYDARSITTPKGSEITPEDIKTNATIRMSPANALNGRKPA 510
E+ G L+A+PH++ S INY D ++ S P + P KPA
Sbjct: 19 ELSGNELIADPHLLPASQINYKDGLAVYAFMNSTKNPLGY-------IYPPKTKLQIKPA 177
Query: 511 PVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTS 570
P MA+FSSRGPN V P ILKPDV APGVNI+AAYS S +NL D RR PF GTS
Sbjct: 178 PAMAAFSSRGPNTVTPEILKPDVIAPGVNIIAAYSEGVSPTNLGFDKRR-VPFITMSGTS 354
Query: 571 MSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
MSCPHV G GL+KTLHP+WSPA IKSA+MTT
Sbjct: 355 MSCPHVAGVVGLLKTLHPDWSPAVIKSALMTT 450
>TC221447 similar to UP|Q75I27 (Q75I27) Putaive subtilisin-like proteinase,
partial (21%)
Length = 491
Score = 136 bits (343), Expect = 2e-32
Identities = 73/161 (45%), Positives = 104/161 (64%), Gaps = 1/161 (0%)
Frame = +1
Query: 217 RDFVGPGTHTLSTAGGNFVQNATIFGIGNGTIKGGSPRSRVATYKACWSLTDVVDCFGAD 276
RD G G+HT +TA G+ V A++FG NGT +G + ++R+ATYK CW + CF +D
Sbjct: 19 RDDDGHGSHTSTTAAGSAVVGASLFGFANGTARGMATQARLATYKVCW----LGGCFTSD 186
Query: 277 VLAAIDQAIYDGADLISVSAGGKPNTNPEVIFTDEISIGAFHALARNILLVASAGNEGPT 336
+ A ID+AI DG +++S+S GG + D I+IG F A A IL+ SAGN GP+
Sbjct: 187 IAAGIDKAIEDGVNILSMSIGGGLMD----YYKDTIAIGTFAATAHGILVSNSAGNGGPS 354
Query: 337 PGSVTNVAPWVFTVAASTLDRDFSSVMTI-NNKTLTGASLF 376
+++NVAPW+ TV A T+DRDF + +T+ N K TG SL+
Sbjct: 355 QATLSNVAPWLTTVGAGTIDRDFPAYITLGNGKMYTGVSLY 477
>TC207702 weakly similar to UP|Q8S896 (Q8S896) Subtilisin-like serine
protease AIR3 (Fragment), partial (20%)
Length = 1025
Score = 134 bits (336), Expect = 2e-31
Identities = 65/75 (86%), Positives = 68/75 (90%)
Frame = +2
Query: 528 ILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMSCPHVVGTAGLIKTLH 587
ILKPDVTAPGVNILAAYS LAS SNL+ DNRRGF FN+ QGTS+SCPHV G AGLIKTLH
Sbjct: 5 ILKPDVTAPGVNILAAYSELASASNLLVDNRRGFKFNVLQGTSVSCPHVAGIAGLIKTLH 184
Query: 588 PNWSPAAIKSAIMTT 602
PNWSPAAIKSAIMTT
Sbjct: 185 PNWSPAAIKSAIMTT 229
>TC226350 similar to PIR|JC7519|JC7519 subtilisin-like serine proteinase -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(34%)
Length = 1312
Score = 133 bits (334), Expect = 3e-31
Identities = 77/166 (46%), Positives = 105/166 (62%), Gaps = 3/166 (1%)
Frame = +1
Query: 440 SAGAVGVIMRNQPEVDGKTLLAEPHVVSTINYYDARSITTPKGSEITP---EDIKTNATI 496
SAGA+G+++ N +G+ L+A+ H++ A ++ G I D K T+
Sbjct: 7 SAGALGMVLSNTA-ANGEELVADAHLLP------ATAVGQKAGDAIKKYLFSDAKP--TV 159
Query: 497 RMSPANALNGRKPAPVMASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTD 556
++ G +P+PV+A+FSSRGPN + P ILKPD+ APGVNILA +S + L D
Sbjct: 160 KILFEGTKLGIQPSPVVAAFSSRGPNSITPQILKPDLIAPGVNILAGWSKAVGPTGLPVD 339
Query: 557 NRRGFPFNIQQGTSMSCPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
NRR FNI GTSMSCPHV G A LIK+ HP+WSPAA++SA+MTT
Sbjct: 340 NRR-VDFNIISGTSMSCPHVSGLAALIKSAHPDWSPAAVRSALMTT 474
>TC232495 similar to UP|Q8W554 (Q8W554) AT3g14240/MLN21_2, partial (40%)
Length = 731
Score = 125 bits (314), Expect = 5e-29
Identities = 73/150 (48%), Positives = 95/150 (62%), Gaps = 2/150 (1%)
Frame = +3
Query: 455 DGKTLLAEPHVVSTINYYDARSITTPKGSEITPE--DIKTNATIRMSPANALNGRKPAPV 512
DG+ L+A+ HV+ A ++ G EI + +T AT + G +PAPV
Sbjct: 6 DGEGLVADCHVLP------ATAVGATAGDEIRSYIGNSRTPATATIVFKGTRLGVRPAPV 167
Query: 513 MASFSSRGPNKVQPYILKPDVTAPGVNILAAYSLLASVSNLVTDNRRGFPFNIQQGTSMS 572
+ASFS+RGPN V P ILKPDV APG+NILAA+ S + +D RR FNI GTSM+
Sbjct: 168 VASFSARGPNPVSPEILKPDVIAPGLNILAAWPDHVGPSGVPSDGRR-TEFNILSGTSMA 344
Query: 573 CPHVVGTAGLIKTLHPNWSPAAIKSAIMTT 602
CPHV G A L+K HP+WSPA+I+SA+MTT
Sbjct: 345 CPHVSGLAALLKAAHPDWSPASIRSALMTT 434
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.138 0.412
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 24,707,328
Number of Sequences: 63676
Number of extensions: 329209
Number of successful extensions: 1837
Number of sequences better than 10.0: 99
Number of HSP's better than 10.0 without gapping: 1743
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1758
length of query: 611
length of database: 12,639,632
effective HSP length: 103
effective length of query: 508
effective length of database: 6,081,004
effective search space: 3089150032
effective search space used: 3089150032
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 62 (28.5 bits)
Medicago: description of AC133779.1


