

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133571.7 + phase: 0
(79 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
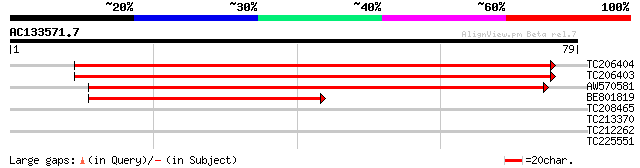
TC206404 similar to UP|Q6Z1L1 (Q6Z1L1) Stromal cell-derived fact... 135 3e-33
TC206403 similar to UP|SDF2_ARATH (Q93ZE8) Stromal cell-derived ... 134 6e-33
AW570581 95 6e-21
BE801819 60 2e-10
TC208465 similar to GB|AAM16215.1|20334918|AY094059 At1g65700/F1... 26 4.2
TC213370 25 5.5
TC212262 similar to UP|PODK_FLAPR (Q42736) Pyruvate, phosphate d... 25 7.1
TC225551 similar to UP|Q9AYT8 (Q9AYT8) Mangrin, partial (68%) 25 9.3
>TC206404 similar to UP|Q6Z1L1 (Q6Z1L1) Stromal cell-derived factor 2-like
protein, partial (65%)
Length = 864
Score = 135 bits (341), Expect = 3e-33
Identities = 63/67 (94%), Positives = 64/67 (95%)
Frame = +2
Query: 10 EGSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 69
EGSGKTWKQDQR RLQHIDT GY +SHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY
Sbjct: 236 EGSGKTWKQDQRIRLQHIDTGGYLHSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 415
Query: 70 LPVTESK 76
LPVTESK
Sbjct: 416 LPVTESK 436
>TC206403 similar to UP|SDF2_ARATH (Q93ZE8) Stromal cell-derived factor
2-like protein precursor (SDF2-like protein), partial
(88%)
Length = 1148
Score = 134 bits (338), Expect = 6e-33
Identities = 62/67 (92%), Positives = 64/67 (94%)
Frame = +2
Query: 10 EGSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 69
EGSGKTWKQDQ+ RLQHIDT GY +SHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY
Sbjct: 716 EGSGKTWKQDQKIRLQHIDTGGYLHSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVY 895
Query: 70 LPVTESK 76
LPVTESK
Sbjct: 896 LPVTESK 916
>AW570581
Length = 232
Score = 95.1 bits (235), Expect = 6e-21
Identities = 46/64 (71%), Positives = 47/64 (72%)
Frame = +2
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEGVYLP 71
SGKTW D R RL H DT GY +SHD KYSR AGGQ E C VREKRADN W A E VYLP
Sbjct: 41 SGKTWNHDHRMRLHHKDTGGYLHSHD*KYSRNAGGQ*EACRVREKRADNAW*AYERVYLP 220
Query: 72 VTES 75
VTES
Sbjct: 221 VTES 232
>BE801819
Length = 118
Score = 60.5 bits (145), Expect = 2e-10
Identities = 25/33 (75%), Positives = 29/33 (87%)
Frame = +2
Query: 12 SGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIA 44
SGKTWKQDQ+ RL HIDT GY +SH+KKYSRI+
Sbjct: 20 SGKTWKQDQKIRLHHIDTGGYLHSHNKKYSRIS 118
>TC208465 similar to GB|AAM16215.1|20334918|AY094059 At1g65700/F1E22_3
{Arabidopsis thaliana;} , complete
Length = 684
Score = 25.8 bits (55), Expect = 4.2
Identities = 13/32 (40%), Positives = 18/32 (55%)
Frame = +3
Query: 35 SHDKKYSRIAGGQQEVCGVREKRADNVWLAAE 66
SH++ YS G QQ V G+ R DN+ + E
Sbjct: 246 SHERVYSTKEGVQQLVLGLYIIRGDNISVVGE 341
>TC213370
Length = 510
Score = 25.4 bits (54), Expect = 5.5
Identities = 10/20 (50%), Positives = 12/20 (60%)
Frame = +3
Query: 6 ETIGEGSGKTWKQDQRFRLQ 25
E GEG K W +D R R+Q
Sbjct: 126 ERFGEGHAKKWWEDNRERIQ 185
>TC212262 similar to UP|PODK_FLAPR (Q42736) Pyruvate, phosphate dikinase,
chloroplast precursor (Pyruvate, orthophosphate
dikinase) , partial (17%)
Length = 983
Score = 25.0 bits (53), Expect = 7.1
Identities = 15/52 (28%), Positives = 23/52 (43%)
Frame = +1
Query: 10 EGSGKTWKQDQRFRLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNV 61
E +TW++ Q + S Y H K +R+ Q CG R +A N+
Sbjct: 391 EEKEQTWQKWQPLAYLFLLDSLYQQKHAKSINRMERSYQMACGRRYLKACNL 546
>TC225551 similar to UP|Q9AYT8 (Q9AYT8) Mangrin, partial (68%)
Length = 1482
Score = 24.6 bits (52), Expect = 9.3
Identities = 12/45 (26%), Positives = 18/45 (39%)
Frame = -3
Query: 23 RLQHIDTSGYFYSHDKKYSRIAGGQQEVCGVREKRADNVWLAAEG 67
R QH+ +S KK + C V E +++WL G
Sbjct: 967 REQHLRNCNRIFSLKKKQKKCGSSLSHCCLVSEASNNSMWLTGLG 833
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.131 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,722,944
Number of Sequences: 63676
Number of extensions: 39353
Number of successful extensions: 105
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 104
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 105
length of query: 79
length of database: 12,639,632
effective HSP length: 55
effective length of query: 24
effective length of database: 9,137,452
effective search space: 219298848
effective search space used: 219298848
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 52 (24.6 bits)
Medicago: description of AC133571.7


