

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133341.5 - phase: 1 /pseudo
(808 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
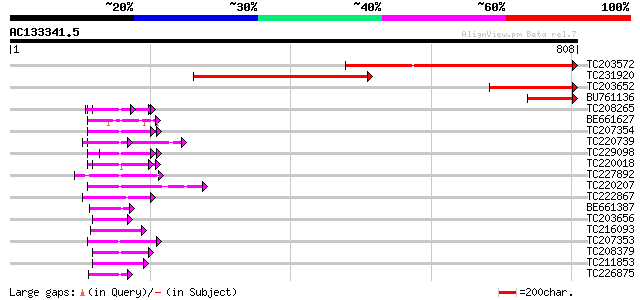
Sequences producing significant alignments: (bits) Value
TC203572 587 e-168
TC231920 368 e-102
TC203652 similar to UP|Q6K884 (Q6K884) Transducin-like, partial ... 240 2e-63
BU761136 136 3e-32
TC208265 similar to UP|Q6S7B0 (Q6S7B0) TAF5, partial (36%) 64 3e-10
BE661627 similar to GP|2443881|gb contains beta-transducin motif... 59 7e-09
TC207354 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat prot... 59 9e-09
TC220739 weakly similar to UP|PRP4_HUMAN (O43172) U4/U6 small nu... 56 6e-08
TC229098 similar to PIR|T46032|T46032 WD-40 repeat regulatory pr... 55 1e-07
TC220018 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana geno... 52 1e-06
TC227892 similar to UP|Q94AB4 (Q94AB4) AT3g13340/MDC11_13, parti... 49 9e-06
TC220207 similar to UP|Q6H975 (Q6H975) STY-L protein, partial (43%) 48 2e-05
TC222867 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Frag... 48 2e-05
BE661387 46 6e-05
TC203656 similar to UP|Q9SYX2 (Q9SYX2) Phytochrome A supressor s... 46 6e-05
TC216093 similar to UP|Q6Z0Y9 (Q6Z0Y9) Sec13p, partial (45%) 46 7e-05
TC207353 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat prot... 45 2e-04
TC208379 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana geno... 45 2e-04
TC211853 similar to UP|O82506 (O82506) F2P3.13 protein, partial ... 44 2e-04
TC226875 similar to UP|Q94JT6 (Q94JT6) At1g78070/F28K19_28, part... 43 5e-04
>TC203572
Length = 1410
Score = 587 bits (1513), Expect = e-168
Identities = 294/332 (88%), Positives = 308/332 (92%), Gaps = 2/332 (0%)
Frame = +3
Query: 479 RDTLAPQISSS--VMANTLPTSNADVAMPSSAMSGSINIPGSSVRSGLRSHFSHSRTPVS 536
RD L+ QI SS +MA+ TSN D AMPSSA+ GSI+IPGSS+RSGLR+HFS SR PVS
Sbjct: 3 RDNLSQQIGSSSYIMASNHSTSNVDAAMPSSAIPGSISIPGSSMRSGLRNHFSQSRVPVS 182
Query: 537 ESGNLAASINTPHDGSDIQTIMSRIQSELATSVAAAAATELPCTVKLRVWSHDIKNPCSP 596
ESGNLAASINTPHDG D+QTI+SRIQSELATSVAA AA ELPCTVKLRVWSHDIKNPC+P
Sbjct: 183 ESGNLAASINTPHDGFDMQTIVSRIQSELATSVAATAA-ELPCTVKLRVWSHDIKNPCAP 359
Query: 597 LNADRCRLIIPHAVLCSEMGAHFSPCGRFLAACVACMLPHIEADPGLQTPVHQESGIATS 656
LNADRCRL +PHAVLCSEMGAHFSPCGRFLAACVACMLPHIEADPGLQTPVHQ+ G+ATS
Sbjct: 360 LNADRCRLTVPHAVLCSEMGAHFSPCGRFLAACVACMLPHIEADPGLQTPVHQDPGVATS 539
Query: 657 PTRHPISAHQVMYELRIYSLEEATFGLVLASRAIRAAHCLTSIQFSPTSEHILLAYGRRH 716
PTRHPISAHQVMYELRIYSLEEATFG VLASRAIRAAHCLTSIQFSPTSEHILLAYGRRH
Sbjct: 540 PTRHPISAHQVMYELRIYSLEEATFGSVLASRAIRAAHCLTSIQFSPTSEHILLAYGRRH 719
Query: 717 GSLLKSIVIDGETTLPIYTVLEVYRVSDMELVRVLPSAEDEVNVACFHPFPGGGLVYGTK 776
GSLLKSIVIDGETTLPIYTVLEVYRVSDMELVRVLPSAEDEVNVACFHPF GGGLVYGTK
Sbjct: 720 GSLLKSIVIDGETTLPIYTVLEVYRVSDMELVRVLPSAEDEVNVACFHPFAGGGLVYGTK 899
Query: 777 EGKLRILHYDGARPVNGTGPSYFPEETIVGVS 808
EGKLR+L YDGA VNG GPSYFPEETIVGVS
Sbjct: 900 EGKLRVLQYDGAHAVNGAGPSYFPEETIVGVS 995
>TC231920
Length = 777
Score = 368 bits (945), Expect = e-102
Identities = 183/258 (70%), Positives = 210/258 (80%), Gaps = 3/258 (1%)
Frame = +3
Query: 262 SSMTEATSIGYLQYPPPAVFVTNVHPTEHVTLSSEPTNVSLPFFLVPPYTVDESRAELQH 321
SSMTEATS+GYLQYPPPAVFVTN+HPTEH++LSSE VSLPF+++P YTVDESRAELQH
Sbjct: 3 SSMTEATSLGYLQYPPPAVFVTNIHPTEHISLSSELPYVSLPFYVMPAYTVDESRAELQH 182
Query: 322 ASHDAGSGRIQIESSAVAQFQADTNSTEQHDTTVSPMDTVSEIPTNSQAGTEYPAHTAFS 381
ASHD GSG +QIES+A+ Q AD ++ ++TTVSPMDT SE+P++SQ G EYPAHTAFS
Sbjct: 183 ASHDVGSGSMQIESAAMVQLHADPSAAAHYETTVSPMDTFSEMPSSSQTGAEYPAHTAFS 362
Query: 382 NGMGIGIGNLTMDGMETDETRPAEGSQHRNPTDASSLNGMLHGLSRQTANHGVHPEDG-- 439
NGMGIG+ NLTMDGMETDETRPAEGSQH N T+ SLNGMLHGLSRQTAN GV E G
Sbjct: 363 NGMGIGLSNLTMDGMETDETRPAEGSQHGNLTNTYSLNGMLHGLSRQTANRGVLSEFGQF 542
Query: 440 HPFVSSRDPSGWELPFLQGWMMGQSQAGLPSMLPHTGVSRDTLAPQI-SSSVMANTLPTS 498
H F SRDPSGWE+PFL GW+MGQSQ G+PSMLPH G SRD L+ I SSS+ A+ TS
Sbjct: 543 HQFFPSRDPSGWEIPFLHGWIMGQSQVGVPSMLPHMGASRDNLSQHIGSSSIKASNPSTS 722
Query: 499 NADVAMPSSAMSGSINIP 516
N D A SS + GSI+IP
Sbjct: 723 NVDAANASSGIPGSISIP 776
>TC203652 similar to UP|Q6K884 (Q6K884) Transducin-like, partial (13%)
Length = 683
Score = 240 bits (613), Expect = 2e-63
Identities = 119/125 (95%), Positives = 121/125 (96%)
Frame = +1
Query: 684 VLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYTVLEVYRVS 743
VLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYTVLEVY+VS
Sbjct: 4 VLASRAIRAAHCLTSIQFSPTSEHILLAYGRRHGSLLKSIVIDGETTLPIYTVLEVYKVS 183
Query: 744 DMELVRVLPSAEDEVNVACFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTGPSYFPEET 803
DMELVRVLPSAEDEVNVACFHPF GGGLVYGTKEGKLR+L YDGA VNGTGPSYFPEET
Sbjct: 184 DMELVRVLPSAEDEVNVACFHPFAGGGLVYGTKEGKLRVLQYDGAHAVNGTGPSYFPEET 363
Query: 804 IVGVS 808
IVGVS
Sbjct: 364 IVGVS 378
>BU761136
Length = 471
Score = 136 bits (343), Expect = 3e-32
Identities = 65/71 (91%), Positives = 66/71 (92%)
Frame = +1
Query: 738 EVYRVSDMELVRVLPSAEDEVNVACFHPFPGGGLVYGTKEGKLRILHYDGARPVNGTGPS 797
EVYRVSDMELVRVLPSAEDEVNVACFHPF GGGLVYGTKEGKLR+L YDGA VNG GPS
Sbjct: 1 EVYRVSDMELVRVLPSAEDEVNVACFHPFAGGGLVYGTKEGKLRVLQYDGAHAVNGAGPS 180
Query: 798 YFPEETIVGVS 808
YFPEETIVGVS
Sbjct: 181 YFPEETIVGVS 213
>TC208265 similar to UP|Q6S7B0 (Q6S7B0) TAF5, partial (36%)
Length = 1048
Score = 63.5 bits (153), Expect = 3e-10
Identities = 32/72 (44%), Positives = 43/72 (59%)
Frame = +2
Query: 108 IAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSL 167
++ A SPDG+ +AS D T+ + D +GRCL LIGH W + F ++ASGS
Sbjct: 377 LSLAMSPDGRYMASGDEDGTIMMWDLSSGRCLTPLIGHTSCVWSLAFSS-EGSVIASGSA 553
Query: 168 DQEVRLWDANTS 179
D V+LWD NTS
Sbjct: 554 DCTVKLWDVNTS 589
Score = 50.4 bits (119), Expect = 3e-06
Identities = 31/94 (32%), Positives = 50/94 (52%), Gaps = 1/94 (1%)
Frame = +2
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
FSP G AS+ D T +I + + L+++ GH+ V++H + +A+GS D+ V
Sbjct: 137 FSPVGHYFASSSHDRTARIWSMDRIQPLRIMAGHLSDVDCVQWHA-NCNYIATGSSDKTV 313
Query: 172 RLWDANTSECITSHHFYR-PIASIAFHAKGEIIA 204
RLWD + EC+ +R I S+A G +A
Sbjct: 314 RLWDVQSGECVRVFVGHRGMILSLAMSPDGRYMA 415
Score = 48.9 bits (115), Expect = 9e-06
Identities = 26/90 (28%), Positives = 49/90 (53%), Gaps = 1/90 (1%)
Frame = +2
Query: 119 LASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANT 178
+A+ D TV++ D ++G C++V +GH + P + +ASG D + +WD ++
Sbjct: 284 IATGSSDKTVRLWDVQSGECVRVFVGHRGMILSLAMSP-DGRYMASGDEDGTIMMWDLSS 460
Query: 179 SECITSHHFYRP-IASIAFHAKGEIIAVAS 207
C+T + + S+AF ++G +IA S
Sbjct: 461 GRCLTPLIGHTSCVWSLAFSSEGSVIASGS 550
>BE661627 similar to GP|2443881|gb contains beta-transducin motif
{Arabidopsis thaliana}, partial (13%)
Length = 735
Score = 59.3 bits (142), Expect = 7e-09
Identities = 45/114 (39%), Positives = 62/114 (53%), Gaps = 11/114 (9%)
Frame = +2
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCL---KVLIGHMRTPWVVRFHPLHPKILASGSLD 168
F+PDG+ + S D+ VK+ D G+ L K GH+R+ + FHPL +LA+GS D
Sbjct: 2 FTPDGRWVVSGGFDNVVKVWDLTAGKLLHDFKFHEGHIRS---IDFHPLE-FLLATGSAD 169
Query: 169 QEVRLWDANTSECITSHHFYRP----IASIAFHAKGEIIAVASGH----KLYIW 214
+ V+ WD T E I S RP + SIAFH G A+ +GH K+Y W
Sbjct: 170 RTVKFWDLETFELIGS---ARPEATGVRSIAFHPDGR--ALFTGHEDGLKVYSW 316
>TC207354 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat protein 5,
partial (57%)
Length = 1124
Score = 58.9 bits (141), Expect = 9e-09
Identities = 32/99 (32%), Positives = 52/99 (52%), Gaps = 2/99 (2%)
Frame = +1
Query: 111 AFSPDGKVLASTHGDHTVKIIDCETGR-CLKVLIGHMRTPWVVRFHPLHPKILASGSLDQ 169
A+S D + S D T++I D G C+K+L GH + V F+P + SGS D+
Sbjct: 283 AWSSDSHYICSASDDRTLRIWDATVGGGCIKILRGHDDAVFCVNFNP-QSSYIVSGSFDE 459
Query: 170 EVRLWDANTSECI-TSHHFYRPIASIAFHAKGEIIAVAS 207
+++WD T +C+ T P+ S+ ++ G +I AS
Sbjct: 460 TIKVWDVKTGKCVHTIKGHTMPVTSVHYNRDGNLIISAS 576
Score = 45.8 bits (107), Expect = 7e-05
Identities = 30/108 (27%), Positives = 50/108 (45%), Gaps = 3/108 (2%)
Frame = +1
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
F+P + S D T+K+ D +TG+C+ + GH V ++ ++ S S D
Sbjct: 415 FNPQSSYIVSGSFDETIKVWDVKTGKCVHTIKGHTMPVTSVHYN-RDGNLIISASHDGSC 591
Query: 172 RLWDANTSECI-TSHHFYRPIASIA-FHAKGEIIAVAS-GHKLYIWHY 216
++WD T + T P S A F G++I A+ L +W+Y
Sbjct: 592 KIWDTETGNLLKTLIEDKAPAVSFAKFSPNGKLILAATLNDTLKLWNY 735
Score = 37.4 bits (85), Expect = 0.027
Identities = 28/86 (32%), Positives = 43/86 (49%), Gaps = 4/86 (4%)
Frame = +2
Query: 94 SAKYCPLL-PAPRST-IAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWV 151
S+K PLL P+P S +A++FSP ++G V G+CLK+ GH+ +
Sbjct: 632 SSKIRPLLSPSPNSPPMASSFSPPLSTTLLSYGTMGV-------GKCLKIYSGHVNRVYC 790
Query: 152 V--RFHPLHPKILASGSLDQEVRLWD 175
+ F + K + GS D V +WD
Sbjct: 791 ITSTFSVTNGKYIVGGSEDHCVYIWD 868
>TC220739 weakly similar to UP|PRP4_HUMAN (O43172) U4/U6 small nuclear
ribonucleoprotein Prp4 (U4/U6 snRNP 60 kDa protein) (WD
splicing factor Prp4) (hPrp4), partial (34%)
Length = 1206
Score = 56.2 bits (134), Expect = 6e-08
Identities = 44/144 (30%), Positives = 62/144 (42%), Gaps = 2/144 (1%)
Frame = +1
Query: 111 AFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQE 170
AF P GK L + D T ++ D ETG L + GH R+ + + FH + AS LD
Sbjct: 340 AFHPSGKYLGTASFDKTWRLWDIETGDELLLQEGHSRSVYGLAFHN-DGSLAASCGLDSL 516
Query: 171 VRLWDANTSECITS-HHFYRPIASIAFHAKGEIIAV-ASGHKLYIWHYDKKGEASYSPIF 228
R+WD T I + +P+ I+F G +A + IW KK P
Sbjct: 517 ARVWDLRTGRSILALEGHVKPVLGISFSPNGYHLATGGEDNTCRIWDLRKKKSFYTIPAH 696
Query: 229 VLKTRRSLRAVHFHPHAAPYLLTA 252
+ V F PH +L+TA
Sbjct: 697 ----SNLISQVKFEPHEGYFLVTA 756
Score = 42.7 bits (99), Expect = 6e-04
Identities = 25/71 (35%), Positives = 35/71 (49%)
Frame = +1
Query: 105 RSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILAS 164
RS AF DG + AS D ++ D TGR + L GH++ + F P + LA+
Sbjct: 448 RSVYGLAFHNDGSLAASCGLDSLARVWDLRTGRSILALEGHVKPVLGISFSP-NGYHLAT 624
Query: 165 GSLDQEVRLWD 175
G D R+WD
Sbjct: 625 GGEDNTCRIWD 657
Score = 36.2 bits (82), Expect = 0.059
Identities = 24/112 (21%), Positives = 45/112 (39%), Gaps = 2/112 (1%)
Frame = +1
Query: 105 RSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILAS 164
+ + +FSP+G LA+ D+T +I D + + H V+F P L +
Sbjct: 574 KPVLGISFSPNGYHLATGGEDNTCRIWDLRKKKSFYTIPAHSNLISQVKFEPHEGYFLVT 753
Query: 165 GSLDQEVRLWDANTSECI--TSHHFYRPIASIAFHAKGEIIAVASGHKLYIW 214
S D ++W + + S H + + G I+ V+ + +W
Sbjct: 754 ASYDMTAKVWSGRDFKPVKTLSGHEAKVTSVDVLGDGGSIVTVSHDRTIKLW 909
>TC229098 similar to PIR|T46032|T46032 WD-40 repeat regulatory protein tup1
homolog - Arabidopsis thaliana {Arabidopsis thaliana;} ,
partial (59%)
Length = 580
Score = 55.1 bits (131), Expect = 1e-07
Identities = 32/108 (29%), Positives = 55/108 (50%), Gaps = 3/108 (2%)
Frame = +3
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
F+P ++ S D TV++ D ++G+CLKVL H V F+ ++ S S D
Sbjct: 78 FNPQSNIIVSGSFDETVRVWDVKSGKCLKVLPAHSDPVTAVDFN-RDGSLIVSSSYDGLC 254
Query: 172 RLWDANTSECITS--HHFYRPIASIAFHAKGEIIAVAS-GHKLYIWHY 216
R+WDA+T C+ + P++ + F + I V + + L +W+Y
Sbjct: 255 RIWDASTGHCMKTLIDDDNPPVSFVKFSPNAKFILVGTLDNTLRLWNY 398
Score = 48.5 bits (114), Expect = 1e-05
Identities = 25/80 (31%), Positives = 43/80 (53%), Gaps = 1/80 (1%)
Frame = +3
Query: 129 KIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANTSECITSHHFY 188
++ D TG +K L GH + V F+P I+ SGS D+ VR+WD + +C+ +
Sbjct: 3 RLWDVPTGSLIKTLHGHTNYVFCVNFNP-QSNIIVSGSFDETVRVWDVKSGKCLKVLPAH 179
Query: 189 R-PIASIAFHAKGEIIAVAS 207
P+ ++ F+ G +I +S
Sbjct: 180 SDPVTAVDFNRDGSLIVSSS 239
Score = 38.5 bits (88), Expect = 0.012
Identities = 19/73 (26%), Positives = 38/73 (52%), Gaps = 2/73 (2%)
Frame = +3
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVV--RFHPLHPKILASGSLDQ 169
FSP+ K + D+T+++ + TG+ LK GH+ + + + F + K + GS +
Sbjct: 333 FSPNAKFILVGTLDNTLRLWNYSTGKFLKTYTGHVNSKYCISSTFSTTNGKYIVGGSEEN 512
Query: 170 EVRLWDANTSECI 182
+ LWD + + +
Sbjct: 513 YIYLWDLQSRKIV 551
>TC220018 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K8K14 (AT5g67320/K8K14_4),
partial (42%)
Length = 1050
Score = 51.6 bits (122), Expect = 1e-06
Identities = 35/113 (30%), Positives = 52/113 (45%), Gaps = 10/113 (8%)
Frame = +1
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPL--------HPKILA 163
+ P G +LAS D T KI + L L H + + +R+ P H +LA
Sbjct: 304 WDPTGSLLASCSDDITAKIWSMKQDTYLHDLREHSKEIYTIRWSPTGPGTNNPNHKLVLA 483
Query: 164 SGSLDQEVRLWDANTSECITSHHFYR-PIASIAFHAKGEIIAVAS-GHKLYIW 214
S S D V+LWD + I S +R P+ S+AF G+ + S ++IW
Sbjct: 484 SASFDSTVKLWDVELGKLIYSLDGHRHPVYSVAFSPNGDYLVSGSLDRSMHIW 642
Score = 47.0 bits (110), Expect = 3e-05
Identities = 28/87 (32%), Positives = 45/87 (51%)
Frame = +1
Query: 118 VLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDAN 177
VLAS D TVK+ D E G+ + L GH + V F P + L SGSLD+ + +W
Sbjct: 475 VLASASFDSTVKLWDVELGKLIYSLDGHRHPVYSVAFSP-NGDYLVSGSLDRSMHIWSLR 651
Query: 178 TSECITSHHFYRPIASIAFHAKGEIIA 204
+ + ++ I + ++ +G+ IA
Sbjct: 652 DGKIVKTYTGNGGIFEVCWNKEGDKIA 732
>TC227892 similar to UP|Q94AB4 (Q94AB4) AT3g13340/MDC11_13, partial (68%)
Length = 1179
Score = 48.9 bits (115), Expect = 9e-06
Identities = 38/129 (29%), Positives = 65/129 (49%), Gaps = 2/129 (1%)
Frame = +3
Query: 93 LSAKYCPLLPAPRSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVV 152
LS +C P +++ SPDGK+LA + ++D +TG+ + L GH+ +
Sbjct: 387 LSKHFCFPWPVNHTSL----SPDGKLLAVVGDNPKGLLVDSQTGKTITPLRGHLDFSFAS 554
Query: 153 RFHPLHPKILASGSLDQEVRLWDA-NTSECI-TSHHFYRPIASIAFHAKGEIIAVASGHK 210
+HP +I A+G+ D+ R+WD N S+ + I SI F + G+ +A+A
Sbjct: 555 AWHP-DGRIFATGNQDKTCRVWDVRNLSKSVAVLKGNLGAIRSIRFTSDGQFMAMAEPAD 731
Query: 211 LYIWHYDKK 219
++ YD K
Sbjct: 732 -FVHVYDTK 755
>TC220207 similar to UP|Q6H975 (Q6H975) STY-L protein, partial (43%)
Length = 1244
Score = 48.1 bits (113), Expect = 2e-05
Identities = 52/176 (29%), Positives = 75/176 (42%), Gaps = 5/176 (2%)
Frame = +3
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
FS DGK LAS D V I + +T H VRF P + LA+ S D+ V
Sbjct: 435 FSSDGKWLASAGDDMKVDIWNMDTLETESTPAEHKSVITDVRFRP-NSSQLATASTDKSV 611
Query: 172 RLWD-ANTSECITSHHFY-RPIASIAFH-AKGEIIAVASG-HKLYIWHYDKKGEASYSPI 227
RLWD N S C+ + + I S+ FH K E+ G +++ W+ + S
Sbjct: 612 RLWDTTNPSRCLQEYSGHSSAIMSLDFHPKKTELFCFCDGENEIRYWNIN-------SAT 770
Query: 228 FVLKTRRSLRAVHFHPHAAPYLLTAEVNDLDSSDSSMTEATSIGYLQ-YPPPAVFV 282
T+ + V F P +L A +D S + T I LQ +P P ++
Sbjct: 771 CTRVTKGASAQVRFQPRLGRFL--AAASDKGVSIFDVESDTQIYTLQGHPEPVSYI 932
>TC222867 similar to UP|Q8L830 (Q8L830) WD40-repeat protein (Fragment),
partial (23%)
Length = 591
Score = 47.8 bits (112), Expect = 2e-05
Identities = 30/105 (28%), Positives = 50/105 (47%), Gaps = 1/105 (0%)
Frame = +1
Query: 105 RSTIAAAFSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILAS 164
R + FSP + + + GD T++I G CLK GH + F +I++
Sbjct: 52 RGIWSVEFSPVDQCVVTASGDKTIRIWAISDGSCLKTFEGHTSSVLRALFVTRGTQIVSC 231
Query: 165 GSLDQEVRLWDANTSECITSH-HFYRPIASIAFHAKGEIIAVASG 208
G+ D V+LW T+EC+ ++ H + ++A K E +A G
Sbjct: 232 GA-DGLVKLWTVKTNECVATYDHHEDKVWALAVGRKTEKLATGGG 363
>BE661387
Length = 636
Score = 46.2 bits (108), Expect = 6e-05
Identities = 25/64 (39%), Positives = 34/64 (53%)
Frame = +1
Query: 115 DGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLW 174
D L + DHT K+ D +T C++ L GH V FHP P I+ +GS D VR+W
Sbjct: 115 DKPYLITGSDDHTAKVWDYQTKSCVQTLEGHTHNVSAVCFHPELP-IIITGSEDGTVRIW 291
Query: 175 DANT 178
+ T
Sbjct: 292 HSTT 303
>TC203656 similar to UP|Q9SYX2 (Q9SYX2) Phytochrome A supressor spa1, partial
(33%)
Length = 1951
Score = 46.2 bits (108), Expect = 6e-05
Identities = 24/56 (42%), Positives = 32/56 (56%)
Frame = +2
Query: 119 LASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLW 174
LAST D V++ D +TG+ L + H + W V F PK+ ASGS D V+LW
Sbjct: 638 LASTDYDGAVQMWDADTGQPLSQYMEHQKRAWSVHFSLSDPKMFASGSDDCSVKLW 805
>TC216093 similar to UP|Q6Z0Y9 (Q6Z0Y9) Sec13p, partial (45%)
Length = 548
Score = 45.8 bits (107), Expect = 7e-05
Identities = 31/83 (37%), Positives = 42/83 (50%), Gaps = 4/83 (4%)
Frame = +1
Query: 116 GKVLASTHGDHTVKIIDCE--TGRCLKVLIGHMRTPWVVRF-HPLHPKILASGSLDQEVR 172
GK LA+ DHT+KII + L L GH W V + HP +LAS S D V
Sbjct: 196 GKRLATASSDHTIKIIGVSNTASQHLATLTGHQGPVWQVAWAHPKFGSLLASCSYDGRVI 375
Query: 173 LW-DANTSECITSHHFYRPIASI 194
+W + N +E +H F P +S+
Sbjct: 376 VWKEGNQNEWTQAHVFDDPKSSV 444
>TC207353 weakly similar to UP|WDR5_HUMAN (P61964) WD-repeat protein 5,
partial (58%)
Length = 898
Score = 44.7 bits (104), Expect = 2e-04
Identities = 30/108 (27%), Positives = 49/108 (44%), Gaps = 3/108 (2%)
Frame = +3
Query: 112 FSPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEV 171
F+P + S D T+K+ D +TG+C+ + GH V ++ ++ S S D
Sbjct: 18 FNPQSSYIVSGSFDETIKVWDVKTGKCVHTIKGHTMPVTSVHYN-RDGTLIISASHDGSC 194
Query: 172 RLWDANTSECI-TSHHFYRPIASIA-FHAKGE-IIAVASGHKLYIWHY 216
++WD T + T P S A F G+ I+A L +W+Y
Sbjct: 195 KIWDTGTGNLLKTLIEDKAPAVSFAKFSPNGKFILAATLNDTLKLWNY 338
Score = 41.2 bits (95), Expect = 0.002
Identities = 33/115 (28%), Positives = 56/115 (48%), Gaps = 7/115 (6%)
Frame = +3
Query: 110 AAFSPDGK-VLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVV--RFHPLHPKILASGS 166
A FSP+GK +LA+T D T+K+ + +G+ LK+ GH+ + + F + + + SGS
Sbjct: 267 AKFSPNGKFILAATLND-TLKLWNYGSGKFLKIYSGHVNRVYCITSTFSVTNGRYIVSGS 443
Query: 167 LDQEVRLWDANTSECITSHHFYR-PIASIAFHAKGEIIA---VASGHKLYIWHYD 217
D+ V +WD I + + S+ H IA +A + +W D
Sbjct: 444 EDRCVYIWDLQAKNMIQKLEGHTDTVISVTCHPTENKIASAGLAGDRTVRVWVQD 608
>TC208379 similar to UP|Q9FN19 (Q9FN19) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K8K14 (AT5g67320/K8K14_4),
partial (42%)
Length = 1062
Score = 44.7 bits (104), Expect = 2e-04
Identities = 28/87 (32%), Positives = 46/87 (52%)
Frame = +2
Query: 118 VLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDAN 177
VLAS D TVK+ D E G+ L L GH + V F P + + +ASGS D+ + +W
Sbjct: 479 VLASASFDSTVKLWDVELGKLLYSLNGHRDRVYSVAFSP-NGEYIASGSPDRSMLIWSLK 655
Query: 178 TSECITSHHFYRPIASIAFHAKGEIIA 204
+ + ++ I + ++ +G+ IA
Sbjct: 656 EGKIVKTYTGDGGIFEVCWNKEGDKIA 736
>TC211853 similar to UP|O82506 (O82506) F2P3.13 protein, partial (45%)
Length = 649
Score = 44.3 bits (103), Expect = 2e-04
Identities = 26/80 (32%), Positives = 36/80 (44%)
Frame = +1
Query: 119 LASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVRLWDANT 178
LAST D VK+ D TG+ H + W V F + P ASGS D V+LW +
Sbjct: 184 LASTDYDGIVKLWDASTGQEFSQFTEHEKRAWSVDFSAVCPTKFASGSDDCTVKLWSISE 363
Query: 179 SECITSHHFYRPIASIAFHA 198
C+ + + + F A
Sbjct: 364 RNCLGTIRNVANVCCVQFSA 423
>TC226875 similar to UP|Q94JT6 (Q94JT6) At1g78070/F28K19_28, partial (84%)
Length = 1542
Score = 43.1 bits (100), Expect = 5e-04
Identities = 25/63 (39%), Positives = 34/63 (53%)
Frame = +1
Query: 113 SPDGKVLASTHGDHTVKIIDCETGRCLKVLIGHMRTPWVVRFHPLHPKILASGSLDQEVR 172
SPDGK+LA I D TG+ L GH+ + +HP +ILA+G+ D+ R
Sbjct: 775 SPDGKLLAVLGDSTECLIADANTGKITGSLKGHLDYSFSSAWHP-DGQILATGNQDKTCR 951
Query: 173 LWD 175
LWD
Sbjct: 952 LWD 960
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.134 0.404
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,729,898
Number of Sequences: 63676
Number of extensions: 591520
Number of successful extensions: 3230
Number of sequences better than 10.0: 126
Number of HSP's better than 10.0 without gapping: 3091
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3191
length of query: 808
length of database: 12,639,632
effective HSP length: 105
effective length of query: 703
effective length of database: 5,953,652
effective search space: 4185417356
effective search space used: 4185417356
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC133341.5


