

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC133139.3 - phase: 0
(185 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
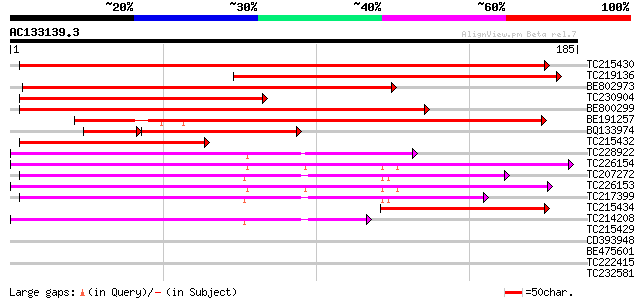
Sequences producing significant alignments: (bits) Value
TC215430 similar to UP|UMPK_ARATH (O04905) Uridylate kinase (UK... 221 2e-58
TC219136 similar to UP|KAD_PRUAR (O24464) Adenylate kinase (ATP... 179 5e-46
BE802973 similar to SP|O04905|UMPK Uridylate kinase (EC 2.7.4.-)... 141 2e-34
TC230904 similar to UP|Q6NMK6 (Q6NMK6) At4g25280, partial (35%) 139 6e-34
BE800299 137 4e-33
BE191257 similar to SP|O04905|UMPK Uridylate kinase (EC 2.7.4.-)... 133 5e-32
BQ133974 82 2e-21
TC215432 homologue to UP|UMPK_ARATH (O04905) Uridylate kinase (... 87 4e-18
TC228922 similar to UP|KADD_ARATH (Q9FIJ7) Probable adenylate ki... 85 2e-17
TC226154 homologue to UP|KADB_ORYSA (Q08480) Adenylate kinase B ... 77 5e-15
TC207272 similar to UP|Q9SNJ4 (Q9SNJ4) ESTs AU065232(E60855), pa... 75 2e-14
TC226153 homologue to UP|KADB_ORYSA (Q08480) Adenylate kinase B ... 74 3e-14
TC217399 similar to UP|KADC_ARATH (Q9ZUU1) Probable adenylate ki... 71 3e-13
TC215434 similar to UP|UMPK_ARATH (O04905) Uridylate kinase (UK... 60 6e-10
TC214208 similar to UP|Q8L7W7 (Q8L7W7) AT3g01820/F28J7_15, parti... 54 6e-08
TC215429 similar to UP|UMPK_ARATH (O04905) Uridylate kinase (UK... 39 0.001
CD393948 similar to SP|Q08480|KADB Adenylate kinase B (EC 2.7.4.... 35 0.026
BE475601 29 1.5
TC222415 similar to UP|Q8W033 (Q8W033) Aldehyde dehydrogenase ,... 28 3.2
TC232581 similar to UP|Q7Y229 (Q7Y229) At1g56720, partial (45%) 28 3.2
>TC215430 similar to UP|UMPK_ARATH (O04905) Uridylate kinase (UK) (Uridine
monophosphate kinase) (UMP kinase) (UMP/CMP kinase) ,
partial (93%)
Length = 1156
Score = 221 bits (563), Expect = 2e-58
Identities = 107/173 (61%), Positives = 135/173 (77%)
Frame = +3
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQCA IVE FG+ HLSAGDLLR + S SE G MI I+EG+IVPS VT++
Sbjct: 267 GGPGSGKGTQCANIVENFGYTHLSAGDLLRAEIKSGSENGTMIQNMIKEGKIVPSEVTIK 446
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
L+ + MQ N KFLIDGFPR+EENR AFE +TG EP FVL+F+CPEEEM +R+LSRNQG
Sbjct: 447 LLQKAMQESGNDKFLIDGFPRNEENRAAFEKVTGIEPAFVLFFECPEEEMERRLLSRNQG 626
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVF 176
R DDNI+TI+KR KVF +LPVI++Y +G++ +I+A +E+FE V+ +F
Sbjct: 627 REDDNIETIRKRFKVFLESSLPVINYYDAKGKVRKIDAARPIEEVFETVKGIF 785
>TC219136 similar to UP|KAD_PRUAR (O24464) Adenylate kinase (ATP-AMP
transphosphorylase) , partial (43%)
Length = 1196
Score = 179 bits (455), Expect = 5e-46
Identities = 86/107 (80%), Positives = 96/107 (89%)
Frame = +3
Query: 74 NRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQGRIDDNIDTIK 133
N FLIDGFPRS+ENRIAFE I G EP VL+FDCPEEEMVKRVLSRNQGRIDDNI+TIK
Sbjct: 3 NHLFLIDGFPRSQENRIAFEQIIGAEPHMVLFFDCPEEEMVKRVLSRNQGRIDDNINTIK 182
Query: 134 KRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVFAACE 180
RL+VFE+LNLPVID+YA++G+L+RINAVGT DEIFE VRPVF ACE
Sbjct: 183 NRLQVFESLNLPVIDYYAKKGKLYRINAVGTVDEIFEHVRPVFEACE 323
>BE802973 similar to SP|O04905|UMPK Uridylate kinase (EC 2.7.4.-) (UK)
(Uridine monophosphate kinase) (UMP kinase) (UMP/CMP
kinase)., partial (65%)
Length = 568
Score = 141 bits (355), Expect = 2e-34
Identities = 69/122 (56%), Positives = 85/122 (69%)
Frame = +2
Query: 5 GPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVRL 64
GPG GKGT CA IV+ + HL+AGDLLR + S SE G MI I +G +VPS VT+ L
Sbjct: 80 GPGRGKGTHCANIVQHLAYTHLTAGDLLRAEIKSGSENGTMIQTMINDGNMVPSEVTMNL 259
Query: 65 ILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQGR 124
+ + M N +FLIDG PR+EENR AF+ +TG EP FVL+ DCPEEEM +R+ RNQGR
Sbjct: 260 LHKAMHESGNDEFLIDGCPRNEENRAAFDRVTGIEPAFVLFVDCPEEEMERRLACRNQGR 439
Query: 125 ID 126
D
Sbjct: 440 ED 445
>TC230904 similar to UP|Q6NMK6 (Q6NMK6) At4g25280, partial (35%)
Length = 421
Score = 139 bits (351), Expect = 6e-34
Identities = 67/81 (82%), Positives = 73/81 (89%)
Frame = +1
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQC +IVETFGFKHLSAGDLLR+ MVSDSEYG+MI+ TI EGRIVPS VTV+
Sbjct: 178 GGPGSGKGTQCGKIVETFGFKHLSAGDLLRREMVSDSEYGSMIMNTIGEGRIVPSEVTVK 357
Query: 64 LILREMQYGDNRKFLIDGFPR 84
LILREM+ DN KFLIDGFPR
Sbjct: 358 LILREMESSDNHKFLIDGFPR 420
>BE800299
Length = 496
Score = 137 bits (344), Expect = 4e-33
Identities = 70/134 (52%), Positives = 93/134 (69%)
Frame = +2
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGP SGKGTQCA IVE G+ HLSAGDLLR + S E G +I I+EG+ PS VT++
Sbjct: 77 GGPRSGKGTQCANIVENCGYTHLSAGDLLRTEIKSGPENGTLIQNMIKEGKSGPSEVTIK 256
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
L+ + +Q N +FLIDG PR+EE+R A E +T EP +VL +C +M +R+LS N G
Sbjct: 257 LLQKAVQESGNDRFLIDGDPRNEEDRAA*ERVT*IEPAYVLVVECAGGKMERRLLSWNHG 436
Query: 124 RIDDNIDTIKKRLK 137
DDN++TI+KR+K
Sbjct: 437 *EDDNVETIRKRVK 478
>BE191257 similar to SP|O04905|UMPK Uridylate kinase (EC 2.7.4.-) (UK)
(Uridine monophosphate kinase) (UMP kinase) (UMP/CMP
kinase)., partial (69%)
Length = 472
Score = 133 bits (334), Expect = 5e-32
Identities = 75/157 (47%), Positives = 104/157 (65%), Gaps = 3/157 (1%)
Frame = -1
Query: 22 GFKHLSAGDLLRKAMVSDSEYGAMILE--TIREGRI-VPSAVTVRLILREMQYGDNRKFL 78
G+ H+SAGD LR + S SE I++ +IR R+ VPS VT++L+ MQ N K+L
Sbjct: 472 GYNHVSAGDSLRAEIKSGSE----IVQ*FSIRSRRMNVPSEVTIKLVQPAMQESGNDKYL 305
Query: 79 IDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQGRIDDNIDTIKKRLKV 138
IDG PR+EE+R AFE +TG EP VL+FDCPE+EM +R+LSR+QGR D N+ ++KR KV
Sbjct: 304 IDGSPRNEESRAAFEIVTGIEPASVLFFDCPEKEMDRRLLSRHQGREDGNLAPLRKRFKV 125
Query: 139 FEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPV 175
F +LPVID+Y G++ I E+ E V+ +
Sbjct: 124 FLESSLPVIDYYDAHGKVRNIYHARPIPEVSEPVQAI 14
>BQ133974
Length = 407
Score = 82.0 bits (201), Expect(2) = 2e-21
Identities = 40/52 (76%), Positives = 45/52 (85%)
Frame = +1
Query: 44 AMILETIREGRIVPSAVTVRLILREMQYGDNRKFLIDGFPRSEENRIAFEHI 95
+MI+ TI EGRIVPS VTV+LILREM+ DN KFLIDGFPRS+ENRIAFE I
Sbjct: 139 SMIMNTIGEGRIVPSEVTVKLILREMESSDNHKFLIDGFPRSQENRIAFEQI 294
Score = 36.6 bits (83), Expect(2) = 2e-21
Identities = 16/19 (84%), Positives = 18/19 (94%)
Frame = +3
Query: 25 HLSAGDLLRKAMVSDSEYG 43
HLSAGDLLR+ MVS+SEYG
Sbjct: 3 HLSAGDLLRREMVSESEYG 59
>TC215432 homologue to UP|UMPK_ARATH (O04905) Uridylate kinase (UK) (Uridine
monophosphate kinase) (UMP kinase) (UMP/CMP kinase) ,
partial (36%)
Length = 452
Score = 87.0 bits (214), Expect = 4e-18
Identities = 41/62 (66%), Positives = 49/62 (78%)
Frame = +1
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GGPGSGKGTQCA IV+ FG+ HLSAGDLLR + S SE G MI I+EG+IVPS VT++
Sbjct: 259 GGPGSGKGTQCANIVQNFGYTHLSAGDLLRAEIKSGSENGTMIQNMIKEGKIVPSEVTIK 438
Query: 64 LI 65
L+
Sbjct: 439 LL 444
>TC228922 similar to UP|KADD_ARATH (Q9FIJ7) Probable adenylate kinase 2,
chloroplast precursor (ATP-AMP transphosphorylase) ,
partial (75%)
Length = 1133
Score = 84.7 bits (208), Expect = 2e-17
Identities = 45/135 (33%), Positives = 75/135 (55%), Gaps = 2/135 (1%)
Frame = +3
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
M+SG P SGKGTQC I +G H++AGDLLR + + S+ G + + +G++VP +
Sbjct: 240 MISGAPASGKGTQCHLITNKYGLVHIAAGDLLRAEIATGSDNGKRAKQYMEKGQLVPDEI 419
Query: 61 TVRLILREMQYGDNRK--FLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVL 118
V ++ + D+++ +L+DG+PRS A E + G P L + E+ +V+RV+
Sbjct: 420 VVMMVKERLLKPDSKENGWLLDGYPRSLSQATALEAL-GFRPHIFLLLEVSEDVLVERVV 596
Query: 119 SRNQGRIDDNIDTIK 133
R + I +K
Sbjct: 597 GRRLDPVTGKIYHLK 641
>TC226154 homologue to UP|KADB_ORYSA (Q08480) Adenylate kinase B (ATP-AMP
transphosphorylase) , partial (86%)
Length = 1116
Score = 77.0 bits (188), Expect = 5e-15
Identities = 58/217 (26%), Positives = 99/217 (44%), Gaps = 33/217 (15%)
Frame = +3
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
+L G PGSGKGTQ I + + HL+ GD+LR A+ + + G E + +G +V +
Sbjct: 51 LLFGPPGSGKGTQSPIIKDEYCLCHLATGDMLRAAVAAKTPLGVKAKEAMDKGELVSDDL 230
Query: 61 TVRLILREMQYGDNRK-FLIDGFPRSEENRIAFEHI---TGTEPDFVLYFDCPEEEMVKR 116
V +I M+ +K F++DGFPR+ + + G + D VL F + + +R
Sbjct: 231 VVGIIDEAMKKPSCQKGFILDGFPRTVVQAQKLDEMLQKQGVKVDKVLNFAIDDAILEER 410
Query: 117 VLSR----NQGRI-------------------------DDNIDTIKKRLKVFEALNLPVI 147
+ R + GR DD +K RL+ F PVI
Sbjct: 411 ITGRWIHPSSGRTYHTKFSPPKVLGVDDVTGEPLIQRKDDTAAVLKSRLEAFHKQTEPVI 590
Query: 148 DHYARRGRLHRINAVGTEDEIFEQVRPVFAACEVYIQ 184
D+Y+++G + ++A E+ +V V + E+ ++
Sbjct: 591 DYYSKKGLVANLHAEKPPKEVTVEVEKVLS*MEITLR 701
>TC207272 similar to UP|Q9SNJ4 (Q9SNJ4) ESTs AU065232(E60855), partial (72%)
Length = 1446
Score = 75.1 bits (183), Expect = 2e-14
Identities = 50/203 (24%), Positives = 90/203 (43%), Gaps = 43/203 (21%)
Frame = +2
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PG GKGT +R+ G H++ GDL+R + S + + E +++G++V + +R
Sbjct: 218 GCPGVGKGTYASRLSNLLGVPHIATGDLVRDELASSDPLSSQLSEIVKQGQLVSDEIIIR 397
Query: 64 LILREMQYGDNR---KFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSR 120
L+ + + G+ + F++DGFPR+ + E + T+ D V+ E+ ++++ L R
Sbjct: 398 LLSKRLVAGEAKGDLGFILDGFPRTIKQAEILEGV--TDIDLVINLKLREDVLLEKCLGR 571
Query: 121 ---NQ-------------------------------------GRIDDNIDTIKKRLKVFE 140
NQ R DD +K+RL+++
Sbjct: 572 RICNQCGGNFNVASINIKAENGSPEIIMAPLLPPENCMSKLITRSDDTEAVVKERLRIYN 751
Query: 141 ALNLPVIDHYARRGRLHRINAVG 163
+ PV + Y RG+L N G
Sbjct: 752 EMTQPVEEFYRSRGKLLEFNLPG 820
>TC226153 homologue to UP|KADB_ORYSA (Q08480) Adenylate kinase B (ATP-AMP
transphosphorylase) , partial (98%)
Length = 917
Score = 74.3 bits (181), Expect = 3e-14
Identities = 57/210 (27%), Positives = 95/210 (45%), Gaps = 33/210 (15%)
Frame = +2
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
+L G PGSGKGTQ I + + HL+ GD+LR A+ + + G E + +G +V +
Sbjct: 233 ILIGPPGSGKGTQSPIIKDEYCLCHLATGDMLRAAVAAKTPLGVKAKEAMDKGELVSDDL 412
Query: 61 TVRLILREMQYGDNRK-FLIDGFPRSEENRIAFEHI---TGTEPDFVLYFDCPEEEMVKR 116
V +I M+ +K F++DGFPR+ + + G + D VL F + + +R
Sbjct: 413 VVGIIDEAMKKPSCQKGFILDGFPRTVVQAQKLDEMLQNQGVKVDKVLNFAIDDAILEER 592
Query: 117 VLSR----NQGRI-------------------------DDNIDTIKKRLKVFEALNLPVI 147
+ R + GR DD +K RL+ F PVI
Sbjct: 593 ITGRWIHPSSGRTYHTKFAPPKVLGVDDVTGEPLIQRKDDTAAVLKLRLEAFHKQTEPVI 772
Query: 148 DHYARRGRLHRINAVGTEDEIFEQVRPVFA 177
D+Y+++G + ++A E+ +V V +
Sbjct: 773 DYYSKKGLVANLHAEKPPKEVTVEVEKVLS 862
>TC217399 similar to UP|KADC_ARATH (Q9ZUU1) Probable adenylate kinase 1,
chloroplast precursor (ATP-AMP transphosphorylase) ,
partial (83%)
Length = 1188
Score = 70.9 bits (172), Expect = 3e-13
Identities = 48/196 (24%), Positives = 85/196 (42%), Gaps = 43/196 (21%)
Frame = +3
Query: 4 GGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
G PG GKGT +R+ G H++ GDL+R + S+ + + E + +G++V + +
Sbjct: 144 GCPGVGKGTYASRLCNLLGVPHIATGDLVRHELASNGPLSSQLSEIVNQGKLVSDEIIIS 323
Query: 64 LILREMQYGDNR---KFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSR 120
L+ + + G+ + F++DGFPR+ + E + T+ D V+ EE ++ + L R
Sbjct: 324 LLSKRLADGEAKGESGFILDGFPRTIKQAEILEGV--TDIDLVVNLKLQEEALLAKCLGR 497
Query: 121 ---NQ-------------------------------------GRIDDNIDTIKKRLKVFE 140
NQ R DD +K+RL+++
Sbjct: 498 RICNQCGGNFNIASISVKGENGRPGMVMAPLLPPAHCMSKLIARSDDTESVVKERLRIYN 677
Query: 141 ALNLPVIDHYARRGRL 156
+ PV Y RG+L
Sbjct: 678 EKSQPVEGFYRSRGKL 725
>TC215434 similar to UP|UMPK_ARATH (O04905) Uridylate kinase (UK) (Uridine
monophosphate kinase) (UMP kinase) (UMP/CMP kinase) ,
partial (28%)
Length = 540
Score = 60.1 bits (144), Expect = 6e-10
Identities = 27/55 (49%), Positives = 41/55 (74%)
Frame = +1
Query: 122 QGRIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVF 176
QGR DDNI+TI+KR KVF +LPVI++Y +G++ +I+A +E+FE V+ +F
Sbjct: 1 QGREDDNIETIRKRFKVFLESSLPVINYYDAKGKVRKIDAARPIEEVFETVKAIF 165
>TC214208 similar to UP|Q8L7W7 (Q8L7W7) AT3g01820/F28J7_15, partial (44%)
Length = 1300
Score = 53.5 bits (127), Expect = 6e-08
Identities = 34/121 (28%), Positives = 59/121 (48%), Gaps = 3/121 (2%)
Frame = +2
Query: 1 MLSGGPGSGKGTQCARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAV 60
++ G PG+ + R+ + H+S LL + + S I T+ G++VP +
Sbjct: 293 VMIGEPGAKRHQFAQRLSKLLEVPHISMATLLSQDLNPRSSLYQQISHTLDHGKLVPEDI 472
Query: 61 TVRLILREMQYGDNR---KFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRV 117
L+ + ++ G +R F++DGFPR+ +HI D V+ F C EEE+ K+
Sbjct: 473 IFGLLSKRLEDGYSRGETGFILDGFPRTRIQAEILDHI--ARVDLVVNFKCSEEELRKKN 646
Query: 118 L 118
L
Sbjct: 647 L 649
>TC215429 similar to UP|UMPK_ARATH (O04905) Uridylate kinase (UK) (Uridine
monophosphate kinase) (UMP kinase) (UMP/CMP kinase) ,
partial (22%)
Length = 356
Score = 39.3 bits (90), Expect = 0.001
Identities = 17/42 (40%), Positives = 29/42 (68%)
Frame = +1
Query: 135 RLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVF 176
R KVF +LPVI++Y +G++ +I+A +E+FE V+ +F
Sbjct: 1 RFKVFLESSLPVINYYDAKGKVRQIDAARPIEEVFEPVKAIF 126
>CD393948 similar to SP|Q08480|KADB Adenylate kinase B (EC 2.7.4.3) (ATP-AMP
transphosphorylase). [Rice] {Oryza sativa}, partial
(52%)
Length = 549
Score = 34.7 bits (78), Expect = 0.026
Identities = 17/54 (31%), Positives = 29/54 (53%)
Frame = -2
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHYARRGRLHRINAVGTEDEIFEQVRPVFA 177
R DD +K RL+ F PVID+Y+++G + ++A E+ +V V +
Sbjct: 329 RKDDTAAVLKLRLEAFHKPTEPVIDYYSKKGLVANLHAEKPPQEVTVEVEKVLS 168
>BE475601
Length = 426
Score = 28.9 bits (63), Expect = 1.5
Identities = 12/18 (66%), Positives = 14/18 (77%), Gaps = 1/18 (5%)
Frame = +3
Query: 92 FEHITG-TEPDFVLYFDC 108
F H+ G T PDFV+YFDC
Sbjct: 90 FHHLIGSTLPDFVVYFDC 143
>TC222415 similar to UP|Q8W033 (Q8W033) Aldehyde dehydrogenase , partial
(25%)
Length = 553
Score = 27.7 bits (60), Expect = 3.2
Identities = 24/92 (26%), Positives = 39/92 (42%), Gaps = 10/92 (10%)
Frame = +1
Query: 14 CARIVETFGFKHLSAGDLLRKAMVSDSEYGAMILE----------TIREGRIVPSAVTVR 63
C+ ++ TF +L + K + E G ++L+ + R GRIV SA V
Sbjct: 262 CSSLLATFLPTYLDNNAI--KVIQGGPEVGKLLLQQRWDKIFFTGSARVGRIVMSAAAVH 435
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHI 95
L ++ G LID S + +A + I
Sbjct: 436 LTPVTLELGGKCPALIDSLSSSWDKEVAVKRI 531
>TC232581 similar to UP|Q7Y229 (Q7Y229) At1g56720, partial (45%)
Length = 994
Score = 27.7 bits (60), Expect = 3.2
Identities = 47/167 (28%), Positives = 73/167 (43%), Gaps = 5/167 (2%)
Frame = +3
Query: 7 GSGKGTQCARIVETFGF---KHLSAGDLLRKAMVSDSEYGAMILETIREGRIVPSAVTVR 63
GSGK R++ TFG+ ++ + G L K+ V +G ++LE I GR V
Sbjct: 207 GSGKSHVATRVMGTFGYVAPEYANTGLLNEKSDV--YSFGVVLLEAI-TGR---DPVDYG 368
Query: 64 LILREMQYGDNRKFLIDGFPRSEENRIAFEHITGTEPDFVLYFDCPEEEMVKRVLSRNQG 123
+E+ D K ++ G RSEE +P+ + P +KRVL
Sbjct: 369 RPAQEVNMVDWLKTMV-GNRRSEE---------VVDPNIEVK---PSTRALKRVLLTAXR 509
Query: 124 RIDDNIDTIKKRLKVFEALNLPVIDHY--ARRGRLHRINAVGTEDEI 168
+D + + KR K+ + + + + Y AR R HR N G EI
Sbjct: 510 CVDPDSE---KRPKMGQVVRMLESEEYPLAREDRRHRRNR-GVNSEI 638
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.142 0.410
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 6,208,243
Number of Sequences: 63676
Number of extensions: 62632
Number of successful extensions: 250
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 240
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 240
length of query: 185
length of database: 12,639,632
effective HSP length: 91
effective length of query: 94
effective length of database: 6,845,116
effective search space: 643440904
effective search space used: 643440904
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC133139.3


