

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130807.10 - phase: 0 /pseudo
(530 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
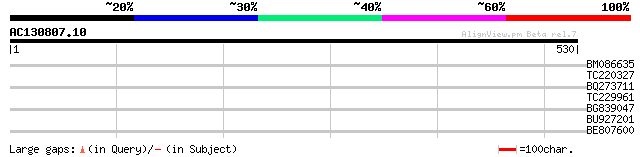
Score E
Sequences producing significant alignments: (bits) Value
BM086635 31 1.6
TC220327 similar to UP|Q8RY07 (Q8RY07) Serine/threonine protein ... 31 1.6
BQ273711 30 3.5
TC229961 similar to GB|AAP68250.1|31711788|BT008811 At3g23640 {A... 29 5.9
BG839047 28 7.7
BU927201 28 7.7
BE807600 similar to GP|24461867|gb CTV.22 {Poncirus trifoliata},... 28 7.7
>BM086635
Length = 428
Score = 30.8 bits (68), Expect = 1.6
Identities = 16/29 (55%), Positives = 23/29 (79%)
Frame = +3
Query: 323 VSLIVLL*LLL*ILVLLIVSLLLNVHISW 351
V+L++LL LLL +L+LL++ LLL VH W
Sbjct: 156 VALLLLLLLLLLLLLLLLLLLLLLVHSVW 242
>TC220327 similar to UP|Q8RY07 (Q8RY07) Serine/threonine protein kinase pk23,
partial (20%)
Length = 780
Score = 30.8 bits (68), Expect = 1.6
Identities = 20/43 (46%), Positives = 29/43 (66%)
Frame = +2
Query: 320 EVLVSLIVLL*LLL*ILVLLIVSLLLNVHISWVWFYLI*KEKW 362
E+ + L++LL LLL +L+LL LL V + + +F L *KEKW
Sbjct: 455 EIFLLLLLLLLLLLSLLLLL**YLLKLVCVFFSFFNLN*KEKW 583
>BQ273711
Length = 409
Score = 29.6 bits (65), Expect = 3.5
Identities = 27/78 (34%), Positives = 38/78 (48%), Gaps = 4/78 (5%)
Frame = -1
Query: 23 ESSPNIQGKV*SRWCPEVA*GSGEDFQGHAVL*SAKGAVRDAYVG*GGR*LV---GKFCY 79
E +Q ++*S C V G+DF+G + * A G++ D Y GG LV F
Sbjct: 238 EPPAKVQWRL*SGRC*VVVS*DGKDFRGDGLP*GA*GSLCDIYAPRGG*KLVEVRETFVC 59
Query: 80 LCWKRMELQ*L-GRCSGE 96
W+R L+ L G+ GE
Sbjct: 58 CTWRRDSLECLQGQVLGE 5
>TC229961 similar to GB|AAP68250.1|31711788|BT008811 At3g23640 {Arabidopsis
thaliana;} , partial (19%)
Length = 717
Score = 28.9 bits (63), Expect = 5.9
Identities = 14/27 (51%), Positives = 17/27 (62%)
Frame = +2
Query: 2 FNSNRNWSW*WRSEDAGDLFEESSPNI 28
+NS + WSW WR G L +SSPNI
Sbjct: 257 YNSTKCWSWFWRVLKVGLL--KSSPNI 331
>BG839047
Length = 610
Score = 28.5 bits (62), Expect = 7.7
Identities = 15/35 (42%), Positives = 24/35 (67%)
Frame = +2
Query: 322 LVSLIVLL*LLL*ILVLLIVSLLLNVHISWVWFYL 356
L SL++LL LLL +L+LL++ LL + + WF +
Sbjct: 467 LFSLLLLLLLLLLLLLLLLLCSLLYYMLCFKWFII 571
>BU927201
Length = 444
Score = 28.5 bits (62), Expect = 7.7
Identities = 15/27 (55%), Positives = 21/27 (77%)
Frame = -2
Query: 322 LVSLIVLL*LLL*ILVLLIVSLLLNVH 348
LV ++LL LLL +LVLL++ LL+ VH
Sbjct: 170 LVERLLLLLLLLLVLVLLVLMLLIIVH 90
>BE807600 similar to GP|24461867|gb CTV.22 {Poncirus trifoliata}, partial
(4%)
Length = 414
Score = 28.5 bits (62), Expect = 7.7
Identities = 21/57 (36%), Positives = 37/57 (64%)
Frame = -2
Query: 322 LVSLIVLL*LLL*ILVLLIVSLLLNVHISWVWFYLI*KEKWLLKLQLRV**LLLLFV 378
++ L++LL LLL +L+LL++ LLL + + +L+ L+ + LR+ *LLLL +
Sbjct: 407 ILLLLLLLLLLLLLLLLLLLLLLLELLPQKLMLHLL-----LVDILLRIL*LLLLLL 252
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.370 0.167 0.608
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,692,214
Number of Sequences: 63676
Number of extensions: 283733
Number of successful extensions: 4018
Number of sequences better than 10.0: 14
Number of HSP's better than 10.0 without gapping: 1557
Number of HSP's successfully gapped in prelim test: 128
Number of HSP's that attempted gapping in prelim test: 2246
Number of HSP's gapped (non-prelim): 1805
length of query: 530
length of database: 12,639,632
effective HSP length: 102
effective length of query: 428
effective length of database: 6,144,680
effective search space: 2629923040
effective search space used: 2629923040
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC130807.10


