

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130806.3 - phase: 0
(320 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
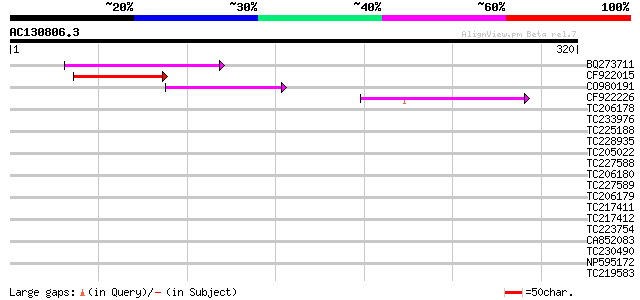
Score E
Sequences producing significant alignments: (bits) Value
BQ273711 85 5e-17
CF922015 77 1e-14
CO980191 46 3e-05
CF922226 41 8e-04
TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 40 0.001
TC233976 39 0.003
TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like prot... 39 0.004
TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana geno... 38 0.007
TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, pa... 37 0.009
TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L1... 37 0.012
TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 37 0.016
TC227589 35 0.035
TC206179 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, part... 35 0.045
TC217411 similar to UP|Q8L7S8 (Q8L7S8) At5g26740, partial (26%) 35 0.059
TC217412 similar to UP|Q6L724 (Q6L724) ATP-dependent RNA helicas... 34 0.077
TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequen... 34 0.10
CA852083 33 0.17
TC230490 similar to UP|Q8LEE4 (Q8LEE4) Zinc finger protein, part... 33 0.17
NP595172 polyprotein [Glycine max] 32 0.29
TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich p... 32 0.50
>BQ273711
Length = 409
Score = 84.7 bits (208), Expect = 5e-17
Identities = 36/90 (40%), Positives = 50/90 (55%)
Frame = -2
Query: 32 AEMRMLETFMRTHPPTFKGRYDPDGAQTWLKEIERVFRVMQYTEDQKVRFGTHQLAEEGD 91
AE R L F + HPP F G YDP+GA+ WL E E++F M E+ KV + T L E +
Sbjct: 270 AEYRGLMAFRKNHPPKFSGDYDPEGARLWLAETEKIFEAMGCLEEHKVLYATFMLQGEAE 91
Query: 92 DWWVSLLPTLKQDGAAVTWAVFRREFLSRY 121
+WW + P+ G + W F+ +FL Y
Sbjct: 90 NWWKFVKPSFVAPGGVIPWNAFKDKFLENY 1
>CF922015
Length = 172
Score = 76.6 bits (187), Expect = 1e-14
Identities = 32/53 (60%), Positives = 42/53 (78%)
Frame = -3
Query: 37 LETFMRTHPPTFKGRYDPDGAQTWLKEIERVFRVMQYTEDQKVRFGTHQLAEE 89
L+ F R +PPTFKG YDP+GA+ WL+EIE++FRVM+ + QKV F TH LA+E
Sbjct: 164 LDRFQRNNPPTFKGGYDPEGAEAWLREIEKIFRVMECQDHQKVLFATHMLADE 6
>CO980191
Length = 802
Score = 45.8 bits (107), Expect = 3e-05
Identities = 25/68 (36%), Positives = 35/68 (50%)
Frame = +1
Query: 89 EGDDWWVSLLPTLKQDGAAVTWAVFRREFLSRYFPEDVRGKKEIEFLELKQGNMSMTEYA 148
EG+ + + L +T VF+ FL ++ ED+ K+ EFLELK GNM + Y
Sbjct: 307 EGNTYEIKLASHGTT*NRVIT*EVFKEIFLEKHLHEDI*NHKKTEFLELKHGNMIVANYL 486
Query: 149 AKFVELAK 156
F EL K
Sbjct: 487 VNFDELPK 510
>CF922226
Length = 667
Score = 40.8 bits (94), Expect = 8e-04
Identities = 25/102 (24%), Positives = 46/102 (44%), Gaps = 7/102 (6%)
Frame = -3
Query: 199 NSCRIYEDDTKAHYKIVNERKTK-------GQQSRPKPYSAPADKGKQRMVEDRRPKKKD 251
+S + E T + K +NERK K G +R K + ++ K++ + + +
Sbjct: 440 DSVSLDEVQTALNSKELNERKEKKSSASGEGLTARGKTFKKDSEFDKKKQKPENQKNGEG 261
Query: 252 ALAEIICFNCGEKGHKSNACPSEIKRCFRCGKKGHKSNAEII 293
+ +I C++C ++GH CP K +K NA I+
Sbjct: 260 NIFKIRCYHCKKEGHTRKVCPERQKNGGSNNRKKDSGNAAIV 135
>TC206178 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (69%)
Length = 1138
Score = 40.4 bits (93), Expect = 0.001
Identities = 15/32 (46%), Positives = 23/32 (71%)
Frame = +2
Query: 257 ICFNCGEKGHKSNACPSEIKRCFRCGKKGHKS 288
+C+NC E GH +++CP+E C CGK GH++
Sbjct: 455 LCWNCKEPGHMASSCPNE-GICHTCGKAGHRA 547
Score = 40.0 bits (92), Expect = 0.001
Identities = 22/100 (22%), Positives = 45/100 (45%), Gaps = 5/100 (5%)
Frame = +2
Query: 207 DTKAHYKIVNERKTKGQQSRPKPYSAP--ADKGKQRMVEDRRPKKKDALAEIICFNCGEK 264
+ K +K+ ++ +++ + P +D+ R RR ++ + +C NC
Sbjct: 185 ERKKKWKMSSDSRSRSRSRSRSPMDRKIRSDRFSYRDAPYRRDSRRGFSRDNLCKNCKRP 364
Query: 265 GHKSNACPSEIKRCFRCGKKGH---KSNAEIICFNIQHEG 301
GH + CP+ + C CG GH + + +C+N + G
Sbjct: 365 GHYARECPN-VAICHNCGLPGHIASECTTKSLCWNCKEPG 481
Score = 36.6 bits (83), Expect = 0.016
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +2
Query: 255 EIICFNCGEKGHKSNACPSEIKRCFRCGKKGH 286
+++C NC + GH S C + C CG +GH
Sbjct: 833 DVVCRNCQQLGHMSRDCMGPLMICHNCGGRGH 928
Score = 35.4 bits (80), Expect = 0.035
Identities = 18/56 (32%), Positives = 27/56 (48%), Gaps = 9/56 (16%)
Frame = +2
Query: 255 EIICFNCGEKGHKSNAC------PSEIKRCFRCGKKGH---KSNAEIICFNIQHEG 301
E IC CG+ GH++ C P +++ C C K+GH + E C N + G
Sbjct: 506 EGICHTCGKAGHRARECSAPPMPPGDLRLCNNCYKQGHIAAECTNEKACNNCRKTG 673
Score = 29.3 bits (64), Expect = 2.5
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +2
Query: 256 IICFNCGEKGHKSNACPS 273
+IC NCG +GH + CPS
Sbjct: 896 MICHNCGGRGHLAYECPS 949
Score = 27.3 bits (59), Expect = 9.5
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = +2
Query: 255 EIICFNCGEKGHKSNACPSEIKRCFRCGKKGH 286
E C NC + GH + CP++ C C GH
Sbjct: 641 EKACNNCRKTGHLARDCPND-PICNLCNVSGH 733
>TC233976
Length = 763
Score = 38.9 bits (89), Expect = 0.003
Identities = 31/113 (27%), Positives = 47/113 (41%), Gaps = 13/113 (11%)
Frame = +1
Query: 205 EDDTKAHYKIVNERKTKGQQ----SRPKPYSAPADK-----GKQRMVEDRRPKKKDALAE 255
+ + A YK N K ++ SR KPYSAP + G++ L
Sbjct: 25 QQEKVAFYKNANASHGKEKKPMTHSRAKPYSAPLEYENHYGGQRTSGGHHLAGGSSQLVN 204
Query: 256 IICFNCGEKGHKSNACPSEIKRCFRCGKKGHKSN----AEIICFNIQHEGRAS 304
+ G G + A + RC +CG+ GH ++ E+ CFN Q +G S
Sbjct: 205 RVSQPAGRGGSGAPAIVTTPLRCRKCGRLGHNAHECTYREVTCFNYQGKGHLS 363
>TC225188 similar to UP|Q9FYB7 (Q9FYB7) Splicing factor-like protein, partial
(62%)
Length = 1131
Score = 38.5 bits (88), Expect = 0.004
Identities = 21/67 (31%), Positives = 34/67 (50%), Gaps = 7/67 (10%)
Frame = +1
Query: 258 CFNCGEKGHKSNACPSEI--KRCFRCGKKGH-----KSNAEIICFNIQHEGRASHRGRSP 310
CFNCG GH + C + +C+RCG++GH K++ + + + R+ R RSP
Sbjct: 364 CFNCGIDGHWARDCKAGDWKNKCYRCGERGHIEKNCKNSPKKLSQRGRRLSRSPVRSRSP 543
Query: 311 ANRSSAE 317
S +
Sbjct: 544 RRGRSRD 564
>TC228935 similar to UP|Q9FHC2 (Q9FHC2) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K24M7, partial (36%)
Length = 1224
Score = 37.7 bits (86), Expect = 0.007
Identities = 15/46 (32%), Positives = 23/46 (49%), Gaps = 6/46 (13%)
Frame = +1
Query: 247 PKKKDALAEIICFNCGEKGHKSNACP------SEIKRCFRCGKKGH 286
P+K + IC C +GH++ CP + K C+ CG+ GH
Sbjct: 313 PEKAEWEKNKICLRCRRRGHRAKNCPEVLDGAKDAKYCYNCGENGH 450
Score = 34.7 bits (78), Expect = 0.059
Identities = 14/38 (36%), Positives = 19/38 (49%), Gaps = 7/38 (18%)
Frame = +1
Query: 258 CFNCGEKGHKSNAC-------PSEIKRCFRCGKKGHKS 288
C+NCGE GH C ++ CF C ++GH S
Sbjct: 424 CYNCGENGHALTQCLHPLQEGGTKFAECFVCNQRGHLS 537
Score = 31.2 bits (69), Expect = 0.66
Identities = 27/106 (25%), Positives = 44/106 (41%), Gaps = 18/106 (16%)
Frame = +1
Query: 216 NERKTKGQQS---RPKPYSAPADKGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACP 272
+++K K + S R +P P + + + R P K + CF C H + CP
Sbjct: 154 DKKKKKNKNSAFKRKRPEPKPGSRKRHLL---RVPGMKPGES---CFICKAMDHIAKLCP 315
Query: 273 SEI-----KRCFRCGKKGH--KSNAEII--------CFNIQHEGRA 303
+ K C RC ++GH K+ E++ C+N G A
Sbjct: 316 EKAEWEKNKICLRCRRRGHRAKNCPEVLDGAKDAKYCYNCGENGHA 453
>TC205022 similar to UP|Q9FYA7 (Q9FYA7) Splicing factor RSZ33, partial (69%)
Length = 1623
Score = 37.4 bits (85), Expect = 0.009
Identities = 21/54 (38%), Positives = 30/54 (54%), Gaps = 2/54 (3%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSEI--KRCFRCGKKGHKSNAEIICFNIQHEGRASHRGRS 309
CFNCG GH + C + +C+RCG++GH E C N + ++ RGRS
Sbjct: 314 CFNCGLDGHWARDCKAGDWKNKCYRCGERGH---IERNCKN-SPKKLSTRRGRS 463
Score = 27.3 bits (59), Expect = 9.5
Identities = 17/71 (23%), Positives = 28/71 (38%), Gaps = 10/71 (14%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSEIKRCFRCGKKGHKSNAEIICFNIQHEGRAS----------HRG 307
C+ CGE+GH C + K+ ++G + + + GR+ R
Sbjct: 380 CYRCGERGHIERNCKNSPKKL--STRRGRSYSRSPVRSHSPRRGRSRDRSYSRDRSYSRS 553
Query: 308 RSPANRSSAEV 318
RSP R + V
Sbjct: 554 RSPVRREESPV 586
>TC227588 similar to PIR|T00837|T00837 glycine-rich protein T13L16.11 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(10%)
Length = 1300
Score = 37.0 bits (84), Expect = 0.012
Identities = 17/34 (50%), Positives = 22/34 (64%), Gaps = 3/34 (8%)
Frame = +1
Query: 258 CFNCGEKGHKSNACPSEIKR---CFRCGKKGHKS 288
CFNCGE+GH + C S +KR C+ CG GH +
Sbjct: 61 CFNCGEEGHAAVNC-SAVKRKKPCYVCGCLGHNA 159
Score = 35.8 bits (81), Expect = 0.027
Identities = 17/41 (41%), Positives = 22/41 (53%), Gaps = 9/41 (21%)
Frame = +1
Query: 255 EIICFNCGEKGHKSNACP---SEI------KRCFRCGKKGH 286
EI C+ CG+ GH AC EI CF+CG++GH
Sbjct: 409 EISCYKCGQLGHTGLACSRLRDEITSGATPSSCFKCGEEGH 531
Score = 33.5 bits (75), Expect = 0.13
Identities = 13/31 (41%), Positives = 20/31 (63%)
Frame = +1
Query: 258 CFNCGEKGHKSNACPSEIKRCFRCGKKGHKS 288
C+ CG GH + C S+++ CF C K GH++
Sbjct: 127 CYVCGCLGHNARQC-SKVQDCFICKKGGHRA 216
Score = 32.3 bits (72), Expect = 0.29
Identities = 13/36 (36%), Positives = 18/36 (49%), Gaps = 7/36 (19%)
Frame = +1
Query: 258 CFNCGEKGHKSNACPSE-------IKRCFRCGKKGH 286
CF C + GH++ CP + I C +CG GH
Sbjct: 184 CFICKKGGHRAKDCPEKHTSTSKSIAICLKCGNSGH 291
Score = 31.6 bits (70), Expect = 0.50
Identities = 13/29 (44%), Positives = 16/29 (54%)
Frame = +1
Query: 258 CFNCGEKGHKSNACPSEIKRCFRCGKKGH 286
CF CGE+GH + C S IK R + H
Sbjct: 505 CFKCGEEGHFARECTSSIKSGKRNWESSH 591
Score = 28.5 bits (62), Expect = 4.2
Identities = 19/70 (27%), Positives = 28/70 (39%), Gaps = 18/70 (25%)
Frame = +1
Query: 257 ICFNCGEKGHKSNACPSEIK-------RCFRCGKKGH--------KSNAEIICF---NIQ 298
IC CG GH +C ++ +C+ C + GH + EI C+ +
Sbjct: 262 ICLKCGNSGHDIFSCRNDYSQDDLKEIQCYVCKRLGHLCCVNTDDATPGEISCYKCGQLG 441
Query: 299 HEGRASHRGR 308
H G A R R
Sbjct: 442 HTGLACSRLR 471
>TC206180 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (42%)
Length = 655
Score = 36.6 bits (83), Expect = 0.016
Identities = 12/32 (37%), Positives = 18/32 (55%)
Frame = +3
Query: 255 EIICFNCGEKGHKSNACPSEIKRCFRCGKKGH 286
+++C NC + GH S C + C CG +GH
Sbjct: 309 DVVCRNCQQLGHMSRDCMGPLMICHNCGGRGH 404
Score = 29.3 bits (64), Expect = 2.5
Identities = 10/18 (55%), Positives = 13/18 (71%)
Frame = +3
Query: 256 IICFNCGEKGHKSNACPS 273
+IC NCG +GH + CPS
Sbjct: 372 MICHNCGGRGHLAYECPS 425
Score = 27.3 bits (59), Expect = 9.5
Identities = 12/32 (37%), Positives = 16/32 (49%)
Frame = +3
Query: 255 EIICFNCGEKGHKSNACPSEIKRCFRCGKKGH 286
E C NC + GH + CP++ C C GH
Sbjct: 126 EKACNNCRKTGHLARDCPND-PICNLCNVSGH 218
>TC227589
Length = 547
Score = 35.4 bits (80), Expect = 0.035
Identities = 15/33 (45%), Positives = 19/33 (57%), Gaps = 2/33 (6%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPS--EIKRCFRCGKKGHKS 288
CFNCGE GH + C + K C+ CG GH +
Sbjct: 47 CFNCGEDGHAAVNCSAAKRKKPCYVCGGLGHNA 145
Score = 35.0 bits (79), Expect = 0.045
Identities = 17/41 (41%), Positives = 21/41 (50%), Gaps = 9/41 (21%)
Frame = +2
Query: 255 EIICFNCGEKGHKSNAC---PSEI------KRCFRCGKKGH 286
EI C+ CG+ GH AC P EI C +CG+ GH
Sbjct: 395 EISCYKCGQLGHTGLACSKLPDEITSAATPSSCCKCGEAGH 517
Score = 31.6 bits (70), Expect = 0.50
Identities = 12/31 (38%), Positives = 19/31 (60%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSEIKRCFRCGKKGHKS 288
C+ CG GH + C ++ + CF C K GH++
Sbjct: 113 CYVCGGLGHNARQC-TKAQDCFICKKGGHRA 202
Score = 31.6 bits (70), Expect = 0.50
Identities = 23/83 (27%), Positives = 35/83 (41%), Gaps = 19/83 (22%)
Frame = +2
Query: 257 ICFNCGEKGHKSNAC-----PSEIK--RCFRCGKKGH--------KSNAEIICFNIQHEG 301
IC CG GH +C P ++K +C+ C + GH + EI C+ G
Sbjct: 248 ICLKCGNSGHDMFSCRNDYSPDDLKEIQCYVCKRVGHLCCVNTDDATPGEISCYKC---G 418
Query: 302 RASHRG----RSPANRSSAEVPT 320
+ H G + P +SA P+
Sbjct: 419 QLGHTGLACSKLPDEITSAATPS 487
>TC206179 similar to UP|Q8LF59 (Q8LF59) DNA-binding protein, partial (16%)
Length = 407
Score = 35.0 bits (79), Expect = 0.045
Identities = 18/78 (23%), Positives = 34/78 (43%)
Frame = +3
Query: 209 KAHYKIVNERKTKGQQSRPKPYSAPADKGKQRMVEDRRPKKKDALAEIICFNCGEKGHKS 268
K K+ ++ +++ + P +D+ R RR ++ + + NC GH +
Sbjct: 165 KKKLKMSSDSRSRSRSRSPMDRKIRSDRFSYRDAPYRRDSRRGFSRDNLGKNCKRPGHYA 344
Query: 269 NACPSEIKRCFRCGKKGH 286
CP+ + C CG GH
Sbjct: 345 RECPN-VAICHNCGLPGH 395
>TC217411 similar to UP|Q8L7S8 (Q8L7S8) At5g26740, partial (26%)
Length = 1275
Score = 34.7 bits (78), Expect = 0.059
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = +3
Query: 246 RPKKKDALAEIICFNCGEKGHKSNACPSEIKR 277
RP D CFNCGE GH+++ CP+ R
Sbjct: 783 RPSSSDRFGGA-CFNCGESGHRASDCPNPSNR 875
>TC217412 similar to UP|Q6L724 (Q6L724) ATP-dependent RNA helicase, partial
(3%)
Length = 868
Score = 34.3 bits (77), Expect = 0.077
Identities = 14/32 (43%), Positives = 18/32 (55%)
Frame = +3
Query: 246 RPKKKDALAEIICFNCGEKGHKSNACPSEIKR 277
RP D CFNCGE GH+++ CP+ R
Sbjct: 279 RPSSSDRFGGT-CFNCGESGHRASDCPNVSNR 371
>TC223754 similar to UP|Q86EQ4 (Q86EQ4) Clone ZZD1536 mRNA sequence, partial
(22%)
Length = 742
Score = 33.9 bits (76), Expect = 0.10
Identities = 17/44 (38%), Positives = 18/44 (40%), Gaps = 15/44 (34%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSEIKR---------------CFRCGKKGH 286
CF CGE GH + C E R CF CGK GH
Sbjct: 317 CFRCGEVGHLARDCGMEGGRFGGGGGSGGGGGKSTCFNCGKPGH 448
Score = 30.4 bits (67), Expect = 1.1
Identities = 14/38 (36%), Positives = 17/38 (43%), Gaps = 9/38 (23%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPSEIKR---------CFRCGKKGH 286
CF CG GH + C + CFRCG+ GH
Sbjct: 230 CFRCGGFGHMARDCATGKGNIGGGGSGGGCFRCGEVGH 343
Score = 28.1 bits (61), Expect = 5.5
Identities = 15/41 (36%), Positives = 16/41 (38%), Gaps = 12/41 (29%)
Frame = +2
Query: 258 CFNCGEKGHKSNAC------------PSEIKRCFRCGKKGH 286
CFNCG GH + C CFRCG GH
Sbjct: 134 CFNCGGFGHLARDCVRGGGGSVGIGGGGGGGSCFRCGGFGH 256
>CA852083
Length = 829
Score = 33.1 bits (74), Expect = 0.17
Identities = 21/72 (29%), Positives = 31/72 (42%), Gaps = 12/72 (16%)
Frame = +1
Query: 216 NERKTKGQQSRPK---PYSAPAD---------KGKQRMVEDRRPKKKDALAEIICFNCGE 263
N+R G + P + P D + K + E K+K+ E+IC CGE
Sbjct: 202 NKRSNAGSDEEEEARDPNAVPTDFTSREAKVWEAKSKATERNWKKRKEE--EMICKLCGE 375
Query: 264 KGHKSNACPSEI 275
GH + CPS +
Sbjct: 376 SGHFTQGCPSTL 411
>TC230490 similar to UP|Q8LEE4 (Q8LEE4) Zinc finger protein, partial (19%)
Length = 574
Score = 33.1 bits (74), Expect = 0.17
Identities = 19/50 (38%), Positives = 26/50 (52%), Gaps = 9/50 (18%)
Frame = +1
Query: 246 RPKKKDAL--AEIICFNCGEKGHKSNACP----SEIKR---CFRCGKKGH 286
R K+ A+ + C NCG +GH+ + CP S + R C CG KGH
Sbjct: 85 RQKRSMAMKGVKFYCQNCGREGHRRHYCPELNDSLVDRRFTCRLCGGKGH 234
Score = 28.1 bits (61), Expect = 5.5
Identities = 18/53 (33%), Positives = 22/53 (40%), Gaps = 1/53 (1%)
Frame = +3
Query: 222 GQQSRPKPYSAP-ADKGKQRMVEDRRPKKKDALAEIICFNCGEKGHKSNACPS 273
G R P + P A K+R RRP K C C ++GH CPS
Sbjct: 324 GHNRRTCPQAVPFAVPSKRRNAASRRPYK--------CRLCKKEGHNIRTCPS 458
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 32.3 bits (72), Expect = 0.29
Identities = 51/249 (20%), Positives = 92/249 (36%), Gaps = 26/249 (10%)
Frame = +1
Query: 51 RYDPDGAQTWLKEIERVFRVMQYTEDQKVRFGTHQLAEEGDDWWVSLLPTLKQDGAAVTW 110
R+D W+ + E+ F + ++ + L ++ W+ L T +W
Sbjct: 310 RFDGKNVMDWIFKAEQFFDYYATPDADRLIIASVHLDQDVVPWYQMLQKTEPFS----SW 477
Query: 111 AVFRREFLSRYFPEDVRGKKEIEFLELKQGNMSMTEYAAKFVELAKFYPHYAAETTEFLK 170
F R + P + F +L Q + ++ EY +F L +AE L
Sbjct: 478 QAFTRALELDFGPSAYDCPRATLF-KLNQ-SATVNEYYMQFTALVNRVDGLSAEA--ILD 645
Query: 171 CIKFENGLRADIKRAIGYQQIRVFSELVNSCRIYEDD------TKAHYKIVNE------- 217
C F +GL+ +I R + + R ++ V +++E+ TK +
Sbjct: 646 C--FVSGLQEEISRDVKAMEPRTLTKAVALAKLFEEKYTSPPKTKTFSNLARNFTSNTSA 819
Query: 218 ----RKTKGQQSRPKPYSAPADKGKQRMVEDRRPK--KKDALAEI-------ICFNCGEK 264
T + PKP P + R + KK + AEI +C+ C EK
Sbjct: 820 TQKYPPTNQKNDNPKPNLPPLLPTPSTKPFNLRNQNIKKISPAEIQLRREKNLCYFCDEK 999
Query: 265 GHKSNACPS 273
++ CP+
Sbjct: 1000FSPAHKCPN 1026
>TC219583 weakly similar to UP|GRP2_NICSY (P27484) Glycine-rich protein 2,
partial (45%)
Length = 995
Score = 31.6 bits (70), Expect = 0.50
Identities = 10/16 (62%), Positives = 13/16 (80%)
Frame = +2
Query: 258 CFNCGEKGHKSNACPS 273
CFNCGE+GH + CP+
Sbjct: 545 CFNCGEEGHFARECPN 592
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.133 0.403
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 13,961,510
Number of Sequences: 63676
Number of extensions: 188972
Number of successful extensions: 1056
Number of sequences better than 10.0: 61
Number of HSP's better than 10.0 without gapping: 971
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1030
length of query: 320
length of database: 12,639,632
effective HSP length: 97
effective length of query: 223
effective length of database: 6,463,060
effective search space: 1441262380
effective search space used: 1441262380
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC130806.3


