

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130804.6 + phase: 0 /pseudo
(99 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
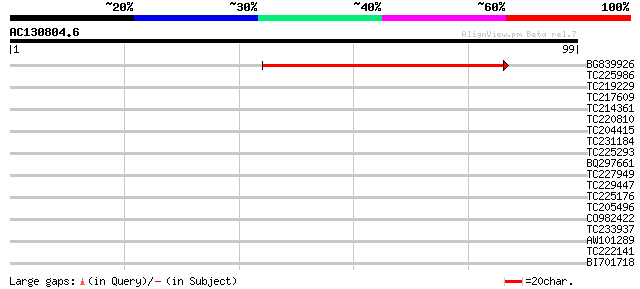
Score E
Sequences producing significant alignments: (bits) Value
BG839926 55 7e-09
TC225986 weakly similar to UP|Q9W1I6 (Q9W1I6) CG18426-PA (LD1203... 29 0.33
TC219229 weakly similar to UP|Q9SD98 (Q9SD98) N-hydroxycinnamoyl... 28 0.73
TC217609 homologue to UP|Q6QI38 (Q6QI38) LRRGT00170, partial (4%) 27 1.6
TC214361 similar to UP|Q8SA69 (Q8SA69) Transcription factor EIL1... 27 2.1
TC220810 similar to PIR|T00552|T00552 lysophospholipase homolog ... 27 2.1
TC204415 similar to UP|Q8BPQ9 (Q8BPQ9) Mus musculus 0 day neonat... 27 2.1
TC231184 26 2.8
TC225293 weakly similar to GB|AAM64538.1|21592589|AY086975 cinna... 26 2.8
BQ297661 26 2.8
TC227949 similar to UP|Q9ZV88 (Q9ZV88) F9K20.26 protein (At1g787... 26 2.8
TC229447 weakly similar to UP|Q7PC53 (Q7PC53) Chitinase B, parti... 26 3.6
TC225176 similar to UP|IF33_ARATH (Q9C5Z2) Eukaryotic translatio... 26 3.6
TC205496 similar to UP|Q9FLC7 (Q9FLC7) Similarity to chalcone-fl... 25 4.7
CO982422 25 6.1
TC233937 similar to GB|AAP37838.1|30725632|BT008479 At4g24220 {A... 25 6.1
AW101289 25 6.1
TC222141 homologue to UP|HMGI_HUMAN (P17096) High mobility group... 25 6.1
BI701718 similar to SP|Q9LD55|IF3A Eukaryotic translation initia... 25 6.1
>BG839926
Length = 750
Score = 54.7 bits (130), Expect = 7e-09
Identities = 27/43 (62%), Positives = 30/43 (68%)
Frame = +1
Query: 45 DGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEFEMEANK 87
DGV PRA MN+QR+RRFR AKD AEE R R+EFE E K
Sbjct: 430 DGVAPRAXMNQQRSRRFRAAKDAADAAAEETRXREEFEKEGRK 558
>TC225986 weakly similar to UP|Q9W1I6 (Q9W1I6) CG18426-PA (LD12035p),
partial (13%)
Length = 1553
Score = 29.3 bits (64), Expect = 0.33
Identities = 16/37 (43%), Positives = 20/37 (53%), Gaps = 4/37 (10%)
Frame = +2
Query: 50 RAKMNKQRTRRFRTAKDNEM----REAEEERLRKEFE 82
R + R R + KD E+ REAE ER+RKE E
Sbjct: 5 RKSRERDRDRELKREKDRELERYEREAERERVRKERE 115
>TC219229 weakly similar to UP|Q9SD98 (Q9SD98)
N-hydroxycinnamoyl/benzoyltransferase-like protein,
partial (43%)
Length = 953
Score = 28.1 bits (61), Expect = 0.73
Identities = 16/58 (27%), Positives = 31/58 (52%)
Frame = -2
Query: 24 ETLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEF 81
E L+A++NR + + PP DI+G K+ R+ +++E + + + +L K F
Sbjct: 943 E*LYAQYNRQMTSLPPKELDINGNFNLPPPKKEEKRK----RESEYKSSRQRKLEKVF 782
>TC217609 homologue to UP|Q6QI38 (Q6QI38) LRRGT00170, partial (4%)
Length = 925
Score = 26.9 bits (58), Expect = 1.6
Identities = 17/57 (29%), Positives = 26/57 (44%)
Frame = +1
Query: 19 KQKIHETLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEE 75
+ +IH +H + I P G TP+ K ++ RR +D+E E EEE
Sbjct: 298 RSEIHHQVHYFSHLAPICATFKEPKGFGPTPKKKKKSKKMRRDYEEEDDEEEEEEEE 468
>TC214361 similar to UP|Q8SA69 (Q8SA69) Transcription factor EIL1, partial
(4%)
Length = 519
Score = 26.6 bits (57), Expect = 2.1
Identities = 19/66 (28%), Positives = 31/66 (46%)
Frame = -1
Query: 22 IHETLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLRKEF 81
IH+ R L+A P PD+ + K N++R RR + +E EEE R++
Sbjct: 327 IHQLNLKISKRLLMAFTPEDPDVGFLV**PKFNRRRRRRELL*TLSRKKEMEEEERREKK 148
Query: 82 EMEANK 87
E + +
Sbjct: 147 ERKVGR 130
>TC220810 similar to PIR|T00552|T00552 lysophospholipase homolog F12L6.8 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(56%)
Length = 833
Score = 26.6 bits (57), Expect = 2.1
Identities = 21/77 (27%), Positives = 31/77 (39%)
Frame = +2
Query: 8 KFACHLLANTLKQKIHETLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDN 67
K A + NT+ + L F W I P P + D+ P+ + + R R+ K N
Sbjct: 95 KIAEEMKPNTMVISVLSALSRVFPSWRIVPTPDIIDLAFKVPKVR-EEIRANRY-CYKGN 268
Query: 68 EMREAEEERLRKEFEME 84
E LR E+E
Sbjct: 269 PRLRTAYELLRVSTEIE 319
>TC204415 similar to UP|Q8BPQ9 (Q8BPQ9) Mus musculus 0 day neonate eyeball
cDNA, RIKEN full-length enriched library,
clone:E130106O21 product:mitogen activated protein
kinase kinase 6, full insert sequence, partial (12%)
Length = 955
Score = 26.6 bits (57), Expect = 2.1
Identities = 16/41 (39%), Positives = 23/41 (56%), Gaps = 2/41 (4%)
Frame = -3
Query: 54 NKQRT--RRFRTAKDNEMREAEEERLRKEFEMEANKFFLNK 92
N++RT R RT K++E E RLRK + + +FF K
Sbjct: 149 NERRTDEER*RTTKNDE--EFPHNRLRKRYRSTSTRFFFTK 33
>TC231184
Length = 541
Score = 26.2 bits (56), Expect = 2.8
Identities = 11/29 (37%), Positives = 15/29 (50%)
Frame = -2
Query: 33 WLIAPPPTLPDIDGVTPRAKMNKQRTRRF 61
W++ PP DI TP +M +R R F
Sbjct: 171 WMLRPPGGCCDITPQTPSERMELRRRRNF 85
>TC225293 weakly similar to GB|AAM64538.1|21592589|AY086975 cinnamoyl-CoA
reductase-like protein {Arabidopsis thaliana;} , partial
(97%)
Length = 1421
Score = 26.2 bits (56), Expect = 2.8
Identities = 21/67 (31%), Positives = 30/67 (44%), Gaps = 12/67 (17%)
Frame = +2
Query: 16 NTLKQKIHETLHAKF-NRWLIAPPPTLPDIDG--------VTPR---AKMNKQRTRRFRT 63
N LK++ H T A RW ++P P P G VTP +K+++ + RR
Sbjct: 86 NKLKEEHH*TNKATLCRRWFVSPEPAAPSDHGWSYSSSSVVTPSMPPSKISRMKMRRNIW 265
Query: 64 AKDNEMR 70
K E R
Sbjct: 266 RKWKEQR 286
>BQ297661
Length = 422
Score = 26.2 bits (56), Expect = 2.8
Identities = 10/25 (40%), Positives = 18/25 (72%)
Frame = +1
Query: 37 PPPTLPDIDGVTPRAKMNKQRTRRF 61
PPP LP +D + ++++++ R RRF
Sbjct: 76 PPPNLPRLDLLRSQSQISQIRRRRF 150
>TC227949 similar to UP|Q9ZV88 (Q9ZV88) F9K20.26 protein
(At1g78700/F9K20_26), partial (78%)
Length = 1834
Score = 26.2 bits (56), Expect = 2.8
Identities = 10/25 (40%), Positives = 18/25 (72%)
Frame = +2
Query: 37 PPPTLPDIDGVTPRAKMNKQRTRRF 61
PPP LP +D + ++++++ R RRF
Sbjct: 89 PPPNLPRLDLLRSQSQISQIRRRRF 163
>TC229447 weakly similar to UP|Q7PC53 (Q7PC53) Chitinase B, partial (4%)
Length = 582
Score = 25.8 bits (55), Expect = 3.6
Identities = 17/66 (25%), Positives = 31/66 (46%)
Frame = +3
Query: 19 KQKIHETLHAKFNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEERLR 78
+Q+ HET+ + ++ P PTL D + + ++Q + E+ E EEE
Sbjct: 156 QQQNHETV--ELDQQQPIPEPTLEDSNTMAIDEPKHEQEEEPQNAESEQEVEEEEEEEDE 329
Query: 79 KEFEME 84
++ E E
Sbjct: 330 QQEEQE 347
>TC225176 similar to UP|IF33_ARATH (Q9C5Z2) Eukaryotic translation
initiation factor 3 subunit 3 (eIF-3 gamma) (eIF3 p38
subunit) (eIF3h), partial (94%)
Length = 1273
Score = 25.8 bits (55), Expect = 3.6
Identities = 11/33 (33%), Positives = 18/33 (54%)
Frame = +1
Query: 30 FNRWLIAPPPTLPDIDGVTPRAKMNKQRTRRFR 62
F WL PPP LPD+ +P+ + ++ + R
Sbjct: 67 FKTWL--PPPLLPDLSSKSPQQRRPRRLSESSR 159
>TC205496 similar to UP|Q9FLC7 (Q9FLC7) Similarity to chalcone-flavonone
isomerase, partial (46%)
Length = 966
Score = 25.4 bits (54), Expect = 4.7
Identities = 13/35 (37%), Positives = 19/35 (54%)
Frame = +2
Query: 60 RFRTAKDNEMREAEEERLRKEFEMEANKFFLNKNV 94
R R A +++ E EEE L K E +K+F +V
Sbjct: 512 RDRLAAEDKYEEEEEEALEKVIEFFQSKYFKKLSV 616
>CO982422
Length = 590
Score = 25.0 bits (53), Expect = 6.1
Identities = 11/52 (21%), Positives = 26/52 (49%), Gaps = 6/52 (11%)
Frame = +2
Query: 12 HLLANTLKQKIHETLHAKFNRWLIAPP------PTLPDIDGVTPRAKMNKQR 57
H++ + Q + +L + +N W PP +P + G+T + ++K++
Sbjct: 182 HIIGSYKLQVYNYSLFSCYNPWFSLPPAESGKVDLVPKVQGITGSSTLSKEK 337
>TC233937 similar to GB|AAP37838.1|30725632|BT008479 At4g24220 {Arabidopsis
thaliana;} , partial (47%)
Length = 692
Score = 25.0 bits (53), Expect = 6.1
Identities = 8/15 (53%), Positives = 10/15 (66%)
Frame = +3
Query: 23 HETLHAKFNRWLIAP 37
H +H KF WL+AP
Sbjct: 180 HAAVHGKFTVWLVAP 224
>AW101289
Length = 412
Score = 25.0 bits (53), Expect = 6.1
Identities = 17/74 (22%), Positives = 33/74 (43%), Gaps = 3/74 (4%)
Frame = +1
Query: 12 HLLANTLKQKIHETLHAKFNRWLIAPPPTLPD---IDGVTPRAKMNKQRTRRFRTAKDNE 68
HL + L+ + L + R PPP L + G+ PR++ ++TRR + +
Sbjct: 10 HLHLHPLRPGLPPQLRPQLPR---QPPPLLRPQNRLPGLRPRSRRRHRKTRRHPPRRQQQ 180
Query: 69 MREAEEERLRKEFE 82
R L+++ +
Sbjct: 181HRPRRPRILQRQLQ 222
>TC222141 homologue to UP|HMGI_HUMAN (P17096) High mobility group protein
HMG-I/HMG-Y (HMG-I(Y)) (High mobility group AT-hook 1),
partial (11%)
Length = 564
Score = 25.0 bits (53), Expect = 6.1
Identities = 11/40 (27%), Positives = 20/40 (49%)
Frame = +3
Query: 37 PPPTLPDIDGVTPRAKMNKQRTRRFRTAKDNEMREAEEER 76
PPP + +T R KQR + RT K ++ ++++
Sbjct: 111 PPPATTPVPLLTRRRAERKQRRPKPRTQKGTRRKQQQQQQ 230
>BI701718 similar to SP|Q9LD55|IF3A Eukaryotic translation initiation factor
3 subunit 10 (eIF-3 theta), partial (12%)
Length = 421
Score = 25.0 bits (53), Expect = 6.1
Identities = 14/44 (31%), Positives = 24/44 (53%)
Frame = +3
Query: 55 KQRTRRFRTAKDNEMREAEEERLRKEFEMEANKFFLNKNVKCQN 98
++ ++R R K E EAE+ RL E+E N+ L + + +N
Sbjct: 288 EEESKRLRLQKITE--EAEQRRLATEYEQRKNQRILREIEEREN 413
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.134 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,306,165
Number of Sequences: 63676
Number of extensions: 45128
Number of successful extensions: 351
Number of sequences better than 10.0: 45
Number of HSP's better than 10.0 without gapping: 350
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 351
length of query: 99
length of database: 12,639,632
effective HSP length: 75
effective length of query: 24
effective length of database: 7,863,932
effective search space: 188734368
effective search space used: 188734368
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 51 (24.3 bits)
Medicago: description of AC130804.6


