

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.5 + phase: 0
(569 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
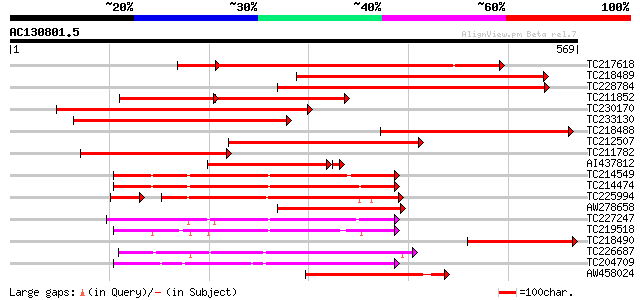
Score E
Sequences producing significant alignments: (bits) Value
TC217618 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {A... 457 e-140
TC218489 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (35%) 430 e-121
TC228784 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (45%) 426 e-119
TC211852 homologue to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (... 267 e-119
TC230170 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (39%) 378 e-105
TC233130 similar to UP|Q9LNN0 (Q9LNN0) F8L10.9 protein, partial ... 358 3e-99
TC218488 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (21%) 344 5e-95
TC212507 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {A... 333 9e-92
TC211782 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (25%) 236 2e-62
AI437812 214 1e-57
TC214549 homologue to UP|O82135 (O82135) Cdc2, complete 216 2e-56
TC214474 homologue to UP|CDC2_VIGAC (Q41639) Cell division contr... 213 2e-55
TC225994 similar to UP|Q7XUF4 (Q7XUF4) OJ991113_30.14 protein, p... 182 1e-50
AW278658 193 2e-49
TC227247 UP|Q6T2Z8 (Q6T2Z8) Cyclin-dependent kinases CDKB, complete 183 2e-46
TC219518 similar to UP|Q9FYT8 (Q9FYT8) Cyclin-dependent kinase B... 181 8e-46
TC218490 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (7%) 179 4e-45
TC226687 homologue to UP|MMK2_MEDSA (Q40353) Mitogen-activated p... 177 1e-44
TC204709 similar to UP|Q8H0X4 (Q8H0X4) Serine/threonine-protein ... 170 2e-42
AW458024 166 3e-41
>TC217618 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Arabidopsis
thaliana;} , partial (60%)
Length = 1325
Score = 457 bits (1176), Expect(2) = e-140
Identities = 219/292 (75%), Positives = 252/292 (86%), Gaps = 2/292 (0%)
Frame = +1
Query: 207 QIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTS 266
Q+KCYM QLLSGLEHCH+R VLHRDIKGSNLLIDNEGILKIADFGLA+FFDP +PMTS
Sbjct: 133 QVKCYMKQLLSGLEHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFFDPKKKHPMTS 312
Query: 267 RVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGS 326
RVVTLWYRPPELLLG+T YGVG+DLWSAGCIL ELL GKP MPGRTEVEQLHKI+KLCGS
Sbjct: 313 RVVTLWYRPPELLLGSTSYGVGVDLWSAGCILAELLAGKPTMPGRTEVEQLHKIFKLCGS 492
Query: 327 PSDEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDAL 386
PSDEYWKK +LPNATL+KP++PYKR I ETFK FP S+LPLI+ LLAIDP +R T S AL
Sbjct: 493 PSDEYWKKYRLPNATLYKPQQPYKRNILETFKDFPSSSLPLIETLLAIDPDDRGTTSAAL 672
Query: 387 RSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAAS--KAQVDGSKKHRTRERS 444
SEFFTTEPYAC+PS+LPKYPP+KE+D K RD+E RRQ+A S VDG+++ R R R
Sbjct: 673 NSEFFTTEPYACEPSNLPKYPPTKELDIKLRDEEARRQKALSGKTNAVDGARRVRVR*RG 852
Query: 445 MKAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQLGFPLGSSH 496
+ A+P PEAN E+Q+N+DR R++THANAKSKSEKFPPPHQDG +G+PL S+
Sbjct: 853 L-AIPGPEANVEIQNNVDRWRVVTHANAKSKSEKFPPPHQDGAVGYPLDDSN 1005
Score = 59.7 bits (143), Expect(2) = e-140
Identities = 28/43 (65%), Positives = 36/43 (83%)
Frame = +2
Query: 169 LQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCY 211
L+GLVTSR+S SLYLVF+YMEHDLAGL+A+ ++F+E Q Y
Sbjct: 2 LEGLVTSRISSSLYLVFEYMEHDLAGLSAAVGVKFSEPQYVLY 130
>TC218489 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (35%)
Length = 763
Score = 430 bits (1106), Expect = e-121
Identities = 207/252 (82%), Positives = 225/252 (89%)
Frame = +2
Query: 289 IDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREP 348
IDLWS GCILGELL GKPI+PG TEVEQLHKIYKL GSPSDEY KKSK+ NATLFKPR P
Sbjct: 8 IDLWSVGCILGELLAGKPILPGPTEVEQLHKIYKL*GSPSDEYRKKSKMHNATLFKPRHP 187
Query: 349 YKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPP 408
YKRCI ETFK FPPSALPLID LLAIDP ER++A+DALRSEFFTTEPYACDPSSLPKYPP
Sbjct: 188 YKRCITETFKDFPPSALPLIDTLLAIDPAERKSATDALRSEFFTTEPYACDPSSLPKYPP 367
Query: 409 SKEMDAKRRDDEVRRQRAASKAQVDGSKKHRTRERSMKAMPAPEANAELQSNIDRRRLIT 468
+KEMDAKRRDDE RR RAA KA VDG+KKHRTR+R+ KA PAPE NAELQSNIDRRR IT
Sbjct: 368 TKEMDAKRRDDEARRSRAAGKAHVDGAKKHRTRDRAAKAAPAPEGNAELQSNIDRRR*IT 547
Query: 469 HANAKSKSEKFPPPHQDGQLGFPLGSSHHIDPDTIPADISFTSTTYTYSKEPFQAWSGPI 528
HANAKSKSEK PPPH+DGQLGFPLGSS+HIDPD P+D+SF ST+YT+SKEPF+AWSG I
Sbjct: 548 HANAKSKSEKLPPPHEDGQLGFPLGSSNHIDPDIDPSDVSFGSTSYTFSKEPFEAWSGAI 727
Query: 529 GNAADIGVSKRK 540
GN A I V+KR+
Sbjct: 728 GNTASISVTKRE 763
>TC228784 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (45%)
Length = 1095
Score = 426 bits (1094), Expect = e-119
Identities = 201/273 (73%), Positives = 231/273 (83%)
Frame = +1
Query: 269 VTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPS 328
VTLWYRPPELLLGAT+Y VG+DLWSAGCIL ELL GKPIMPGRTEVEQLHKI+KLCGSPS
Sbjct: 1 VTLWYRPPELLLGATEYSVGVDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPS 180
Query: 329 DEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRS 388
DEYWKKSKLP+AT+FKP++ YKRCI ET+K FPPS+LPL+D LLAI+P ER TA+ AL S
Sbjct: 181 DEYWKKSKLPHATIFKPQQSYKRCIAETYKDFPPSSLPLMDTLLAINPDERLTATAALHS 360
Query: 389 EFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQVDGSKKHRTRERSMKAM 448
EFFTT+PYAC+PSSLPKYPPSKEMDAK RD+E RR RAA KA DG KK R RER + +
Sbjct: 361 EFFTTKPYACEPSSLPKYPPSKEMDAKLRDEEARRLRAAGKANADGVKKSRPRERVGRGI 540
Query: 449 PAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQLGFPLGSSHHIDPDTIPADIS 508
PEANAELQ+NIDRRRLITH+NAKSKSEKFPPPHQDG LG+PLGSSHH+DP P D+
Sbjct: 541 AVPEANAELQANIDRRRLITHSNAKSKSEKFPPPHQDGALGYPLGSSHHMDPVFDPPDVP 720
Query: 509 FTSTTYTYSKEPFQAWSGPIGNAADIGVSKRKK 541
F+ST ++ K Q WSGP+ + + +GV +RKK
Sbjct: 721 FSSTNFSQPKANIQTWSGPLVDPSGVGVPRRKK 819
>TC211852 homologue to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (39%)
Length = 706
Score = 267 bits (682), Expect(2) = e-119
Identities = 124/135 (91%), Positives = 129/135 (94%)
Frame = +2
Query: 207 QIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTS 266
Q+KCYM+QLLSGLEHCHNR VLHRDIKGSNLLIDNEG LKIADFGLAS FDPN+ +PMTS
Sbjct: 299 QVKCYMHQLLSGLEHCHNRHVLHRDIKGSNLLIDNEGTLKIADFGLASIFDPNHKHPMTS 478
Query: 267 RVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGS 326
RVVTLWYRPPELLLGATDY VG+DLWSAGCILGELL GKPIMPGRTEVEQLHKIYKLCGS
Sbjct: 479 RVVTLWYRPPELLLGATDYSVGVDLWSAGCILGELLAGKPIMPGRTEVEQLHKIYKLCGS 658
Query: 327 PSDEYWKKSKLPNAT 341
PSDEYWKKSKLPNAT
Sbjct: 659 PSDEYWKKSKLPNAT 703
Score = 180 bits (457), Expect(2) = e-119
Identities = 88/99 (88%), Positives = 94/99 (94%)
Frame = +1
Query: 111 DKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQ 170
DKIGQGTYSNVYKA D MTGK+VALKKVRFDN EPES+KFMAREI+ILRRLDHPNV+KLQ
Sbjct: 1 DKIGQGTYSNVYKAKDMMTGKIVALKKVRFDNWEPESVKFMAREILILRRLDHPNVVKLQ 180
Query: 171 GLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIK 209
GLVTSRMSCSLYLVFDYMEHDLAGLAASP IRFTE Q++
Sbjct: 181 GLVTSRMSCSLYLVFDYMEHDLAGLAASPGIRFTEPQVR 297
>TC230170 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (39%)
Length = 826
Score = 378 bits (970), Expect = e-105
Identities = 185/258 (71%), Positives = 213/258 (81%), Gaps = 1/258 (0%)
Frame = +1
Query: 48 QKKKSDDSVQRSRRSKPNPRLSNPPKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADTF 107
+++++ +R R SKPNPRLSNPP H+ GEQVAAGWPSWL+ V GEA+ G PR+ADTF
Sbjct: 19 EERRARTRGERRRSSKPNPRLSNPPNHVHGEQVAAGWPSWLSKVAGEAINGLTPRRADTF 198
Query: 108 EKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVI 167
EKIDKIGQGTYSNVYKA D++TGK+VALKKVRFDNLEPES+KFMAREI+ILRRLDHPNVI
Sbjct: 199 EKIDKIGQGTYSNVYKARDTLTGKIVALKKVRFDNLEPESVKFMAREILILRRLDHPNVI 378
Query: 168 KLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRV 227
KL+GLVTSRMSCSLYLVF+YM HDLAGLA +P I+FTESQ+KCYM+QL SGL
Sbjct: 379 KLEGLVTSRMSCSLYLVFEYMVHDLAGLATNPAIKFTESQVKCYMHQLFSGLGTLSQPSC 558
Query: 228 LHRDI-KGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATDYG 286
I +G NLLIDN+GIL I DFGL SFFDPN+ +PMTS VTLWY P LLLGAT+Y
Sbjct: 559 FCIXI*RGPNLLIDNDGILXIGDFGLXSFFDPNHKHPMTSXXVTLWYXXPXLLLGATEYX 738
Query: 287 VGIDLWSAGCILGELLVG 304
V +DLW+AGCIL ELL G
Sbjct: 739 VRMDLWNAGCILAELLAG 792
>TC233130 similar to UP|Q9LNN0 (Q9LNN0) F8L10.9 protein, partial (30%)
Length = 767
Score = 358 bits (920), Expect = 3e-99
Identities = 171/218 (78%), Positives = 192/218 (87%)
Frame = +1
Query: 65 NPRLSNPPKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKA 124
+P + + PK + GEQVAAGWPSWL AV GEA+ GW+PR+AD+FEK+DKIGQGTYSNVY+A
Sbjct: 112 HPGIGSVPKAMEGEQVAAGWPSWLAAVAGEAIKGWLPRRADSFEKLDKIGQGTYSNVYRA 291
Query: 125 IDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLV 184
D KVVALKKVRFDNLEPES++FMAREI ILRRLDHPNVIKL+GLVTSRMSCSLYLV
Sbjct: 292 RDLEQRKVVALKKVRFDNLEPESVRFMAREIHILRRLDHPNVIKLEGLVTSRMSCSLYLV 471
Query: 185 FDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGI 244
F+YMEHDLAGLA+ P ++FTE+Q+KCYM QLL GL+HCHN VLHRDI GSNLLIDN GI
Sbjct: 472 FEYMEHDLAGLASHPKLKFTEAQVKCYMQQLLQGLDHCHNCGVLHRDINGSNLLIDNNGI 651
Query: 245 LKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGA 282
LKIADFGLAS FDPN P+TSRVVTLWYRPPELLLGA
Sbjct: 652 LKIADFGLASVFDPNQTQPLTSRVVTLWYRPPELLLGA 765
>TC218488 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (21%)
Length = 581
Score = 344 bits (883), Expect = 5e-95
Identities = 166/193 (86%), Positives = 176/193 (91%)
Frame = +2
Query: 373 AIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRRDDEVRRQRAASKAQV 432
AIDPVER+TASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAK+RDDE+RR RAA KAQ
Sbjct: 2 AIDPVERKTASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKQRDDEMRRLRAAGKAQA 181
Query: 433 DGSKKHRTRERSMKAMPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQLGFPL 492
DG KKHRTR R+ KA PAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQ+GFPL
Sbjct: 182 DGPKKHRTRNRAAKAFPAPEANAELQSNIDRRRLITHANAKSKSEKFPPPHQDGQVGFPL 361
Query: 493 GSSHHIDPDTIPADISFTSTTYTYSKEPFQAWSGPIGNAADIGVSKRKKYTAGEAWDLLK 552
GSSHHIDPDT+P D+SFTST+YTYSKEPFQAWSGPIGNAADIGV KRKK+TA +A DL K
Sbjct: 362 GSSHHIDPDTVPTDVSFTSTSYTYSKEPFQAWSGPIGNAADIGVPKRKKHTAADALDLSK 541
Query: 553 PHTGAPKDKFKGK 565
P A KDK KGK
Sbjct: 542 PQKDALKDKVKGK 580
>TC212507 similar to GB|AAO64868.1|29028978|BT005933 At5g50860 {Arabidopsis
thaliana;} , partial (34%)
Length = 590
Score = 333 bits (855), Expect = 9e-92
Identities = 158/196 (80%), Positives = 174/196 (88%)
Frame = +2
Query: 220 EHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELL 279
EHCH+R VLHRDIKGSNLLIDNEGILKIADFGLA+F+DP MTSRVVTLWYRPPELL
Sbjct: 2 EHCHSRGVLHRDIKGSNLLIDNEGILKIADFGLATFYDPKIKQAMTSRVVTLWYRPPELL 181
Query: 280 LGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPN 339
LGAT YGVGIDLWSAGCIL ELL GKPIMPGRTEVEQLHKI+KLCGSPS+EYW+K +LPN
Sbjct: 182 LGATVYGVGIDLWSAGCILAELLAGKPIMPGRTEVEQLHKIFKLCGSPSEEYWRKYRLPN 361
Query: 340 ATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACD 399
AT+FKP++PYKRCI ET+K FPPS+LPLI+ LLAI P +R TAS AL SEFFTT PYAC+
Sbjct: 362 ATIFKPQQPYKRCILETYKDFPPSSLPLIETLLAIYPDDRCTASSALNSEFFTTYPYACE 541
Query: 400 PSSLPKYPPSKEMDAK 415
PSSLPKYPP K +D K
Sbjct: 542 PSSLPKYPPIK*LDVK 589
>TC211782 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (25%)
Length = 482
Score = 236 bits (603), Expect = 2e-62
Identities = 109/151 (72%), Positives = 132/151 (87%)
Frame = +3
Query: 72 PKHLRGEQVAAGWPSWLTAVCGEALTGWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGK 131
PK + GEQVAAGWP+WL++V GEA+ GWIPR A+TFE++ KIGQGTYS VYKA D + K
Sbjct: 30 PKAIEGEQVAAGWPAWLSSVAGEAIKGWIPRSANTFERLHKIGQGTYSTVYKARDVINQK 209
Query: 132 VVALKKVRFDNLEPESIKFMAREIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHD 191
VALKKVRFDNL+PES+KFM REI +LRRLDHPN+IKL+GL+TS+MS SLYLVF+YMEHD
Sbjct: 210 FVALKKVRFDNLDPESVKFMTREIHVLRRLDHPNIIKLEGLITSQMSRSLYLVFEYMEHD 389
Query: 192 LAGLAASPVIRFTESQIKCYMNQLLSGLEHC 222
L GLA++P I+F+E Q+KCYM QLLSGL+HC
Sbjct: 390 LTGLASNPDIKFSEPQLKCYMQQLLSGLDHC 482
>AI437812
Length = 421
Score = 214 bits (546), Expect(2) = 1e-57
Identities = 100/125 (80%), Positives = 109/125 (87%)
Frame = +1
Query: 199 PVIRFTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDP 258
P ++FTE+Q+KCYM QLL GL+HCH+ VLHRDIKGSNLLIDN GILKIADFGLASFFDP
Sbjct: 7 PGLKFTEAQVKCYMQQLLRGLDHCHSCGVLHRDIKGSNLLIDNSGILKIADFGLASFFDP 186
Query: 259 NYMNPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLH 318
N P+TSRVVTLWYRPPELLLGAT YG +DLWS GCIL EL GKPIMPGRTEVEQLH
Sbjct: 187 NQAQPLTSRVVTLWYRPPELLLGATYYGTAVDLWSTGCILAELYAGKPIMPGRTEVEQLH 366
Query: 319 KIYKL 323
KI+KL
Sbjct: 367 KIFKL 381
Score = 27.3 bits (59), Expect(2) = 1e-57
Identities = 9/12 (75%), Positives = 12/12 (100%)
Frame = +3
Query: 325 GSPSDEYWKKSK 336
GSPS++YW+KSK
Sbjct: 384 GSPSEDYWRKSK 419
>TC214549 homologue to UP|O82135 (O82135) Cdc2, complete
Length = 1216
Score = 216 bits (550), Expect = 2e-56
Identities = 118/293 (40%), Positives = 181/293 (61%), Gaps = 6/293 (2%)
Frame = +3
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRLDH 163
+ +EK++KIG+GTY VYK D +T + +ALKK+R + E E + A REI +L+ + H
Sbjct: 159 EQYEKVEKIGEGTYGVVYKGRDRVTNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQH 335
Query: 164 PNVIKLQGLVTSRMSCSLYLVFDYMEHDLAG-LAASPVIRFTESQIKCYMNQLLSGLEHC 222
N+++LQ +V S LYLVF+Y++ DL + +SP Q+K ++ Q+L G+ +C
Sbjct: 336 RNIVRLQDVVHDEKS--LYLVFEYLDLDLKKHMDSSPEFAKDPRQVKMFLYQILCGIAYC 509
Query: 223 HNRRVLHRDIKGSNLLIDNE-GILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLG 281
H+ RVLHRD+K NLLID LK+ADFGLA F + T VVTLWYR PE+LLG
Sbjct: 510 HSHRVLHRDLKPQNLLIDRSTNALKLADFGLARAFGIP-VRTFTHEVVTLWYRAPEILLG 686
Query: 282 ATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKLPNA 340
+ Y +D+WS GCI E++ +P+ PG +E+++L KI+++ G+P+++ W + LP+
Sbjct: 687 SRQYSTPVDIWSVGCIFAEMVNQRPLFPGDSEIDELFKIFRIMGTPNEDTWPGVTSLPD- 863
Query: 341 TLFKPREP--YKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
FK P + ++ P+ L L+ +L +DP +R TA AL E+F
Sbjct: 864 --FKSAFPKWQPKDLKNVVPNLEPAGLDLLSSMLYLDPSKRITARSALEHEYF 1016
>TC214474 homologue to UP|CDC2_VIGAC (Q41639) Cell division control protein 2
homolog (p34cdc2) , complete
Length = 1204
Score = 213 bits (542), Expect = 2e-55
Identities = 117/291 (40%), Positives = 179/291 (61%), Gaps = 4/291 (1%)
Frame = +2
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMA-REIIILRRLDH 163
+ +EK++KIG+GTY VYKA D T + +ALKK+R + E E + A REI +L+ + H
Sbjct: 83 EQYEKVEKIGEGTYGVVYKARDRATNETIALKKIRLEQ-EDEGVPSTAIREISLLKEMQH 259
Query: 164 PNVIKLQGLVTSRMSCSLYLVFDYMEHDLAG-LAASPVIRFTESQIKCYMNQLLSGLEHC 222
N+++LQ +V S LYLVF+Y++ DL + +SP Q+K ++ Q+L G+ +C
Sbjct: 260 RNIVRLQDVVHSEKR--LYLVFEYLDLDLKKHMDSSPEFVKDPRQVKMFLYQILCGIAYC 433
Query: 223 HNRRVLHRDIKGSNLLIDNE-GILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLG 281
H+ RVLHRD+K NLLID LK+ADFGLA F + T VVTLWYR PE+LLG
Sbjct: 434 HSHRVLHRDLKPQNLLIDRRTNSLKLADFGLARAFGIP-VRTFTHEVVTLWYRAPEILLG 610
Query: 282 ATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWK-KSKLPNA 340
+ Y +D+WS GCI E++ +P+ PG +E+++L KI+++ G+P+++ W + LP+
Sbjct: 611 SRHYSTPVDVWSVGCIFAEMVNRRPLFPGDSEIDELFKIFRILGTPNEDTWPGVTSLPDF 790
Query: 341 TLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
P+ P K + + L L+ +L +DP +R TA A+ E+F
Sbjct: 791 KSTFPKWPSKD-LANVVPNLDAAGLNLLSSMLCLDPSKRITARSAVEHEYF 940
>TC225994 similar to UP|Q7XUF4 (Q7XUF4) OJ991113_30.14 protein, partial (43%)
Length = 2770
Score = 182 bits (462), Expect(2) = 1e-50
Identities = 100/251 (39%), Positives = 153/251 (60%), Gaps = 8/251 (3%)
Frame = +2
Query: 153 REIIILRRLDHPNVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYM 212
REI IL +HP+++ ++ +V ++V ++ME+DL GL F+ S+IK +
Sbjct: 788 REINILLSFNHPSIVNVKEVVVDDFD-GTFMVMEHMEYDLKGLMEVKKQPFSMSEIKSLV 964
Query: 213 NQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLW 272
QLL G+++ H+ V+HRD+K SN+L++++G LKI DFGL+ + + + P T VVTLW
Sbjct: 965 RQLLEGVKYLHDNWVIHRDLKSSNILLNHDGELKICDFGLSRQYG-SPLKPYTPVVVTLW 1141
Query: 273 YRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYW 332
YR PELLLGA +Y ID+WS GCI+ EL+ +P+ G++E+EQL KI++ G+P ++ W
Sbjct: 1142 YRAPELLLGAKEYSTSIDMWSVGCIMAELIAKEPLFRGKSELEQLDKIFRTLGTPDEKIW 1321
Query: 333 K-KSKLPNATLFKPREPY----KRCIRETFKGFP---PSALPLIDKLLAIDPVERETASD 384
SKLP A ++P K+ +F G P L+ +LL DP +R TA D
Sbjct: 1322 PGLSKLPGAKANFVKQPINTLRKKFPAASFTGLPVLSELGFDLLKRLLTYDPEKRITAED 1501
Query: 385 ALRSEFFTTEP 395
AL ++F P
Sbjct: 1502 ALLHDWFHEAP 1534
Score = 36.6 bits (83), Expect(2) = 1e-50
Identities = 18/34 (52%), Positives = 22/34 (63%)
Frame = +3
Query: 102 RKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVAL 135
R F+ I KI +GTY VYKA D TG++VAL
Sbjct: 630 RSVCEFKMIKKINEGTYGVVYKARDKKTGELVAL 731
>AW278658
Length = 395
Score = 193 bits (490), Expect = 2e-49
Identities = 85/129 (65%), Positives = 107/129 (82%)
Frame = +3
Query: 269 VTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPS 328
VTLWYRPPELLLG+T YG +DLWS GC+ ELL+GKPI+ GRTEVEQLHKI+KLCGSP
Sbjct: 9 VTLWYRPPELLLGSTAYGASVDLWSVGCVFAELLIGKPILQGRTEVEQLHKIFKLCGSPP 188
Query: 329 DEYWKKSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRS 388
+EYWKK++LP+ATLFKP++PY C+RETFK F S++ L+ LL+++P +R TAS AL
Sbjct: 189 EEYWKKTRLPHATLFKPQQPYDSCLRETFKDFHASSVNLLQTLLSVEPSKRGTASSALSL 368
Query: 389 EFFTTEPYA 397
E+F T+PYA
Sbjct: 369 EYFKTKPYA 395
>TC227247 UP|Q6T2Z8 (Q6T2Z8) Cyclin-dependent kinases CDKB, complete
Length = 1235
Score = 183 bits (464), Expect = 2e-46
Identities = 114/305 (37%), Positives = 167/305 (54%), Gaps = 11/305 (3%)
Frame = +2
Query: 98 GWIPRKADTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIII 157
G + + FEK++K+G+GTY VY+A + TGK+VALKK R E RE+ I
Sbjct: 50 GGVLSAKEAFEKLEKVGEGTYGKVYRAREKATGKIVALKKTRLHEDEEGVPPTTLREVSI 229
Query: 158 LRRLDH-PNVIKLQGLVTSRMS---CSLYLVFDYMEHDLAGLAASPVIRFT-----ESQI 208
LR L P+V++L + + LYLVF+YM+ DL S R T I
Sbjct: 230 LRMLSRDPHVVRLMDVKQGQNKEGKTVLYLVFEYMDTDLKKFIRS--FRQTGQTVPPQTI 403
Query: 209 KCYMNQLLSGLEHCHNRRVLHRDIKGSNLLIDNEGI-LKIADFGLASFFDPNYMNPMTSR 267
K M QL G+ CH +LHRD+K NLL+D + + LKIAD GLA F + T
Sbjct: 404 KSLMYQLCKGVAFCHGHGILHRDLKPHNLLMDPKTMMLKIADLGLARAFTVP-IKKYTHE 580
Query: 268 VVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSP 327
++TLWYR PE+LLGAT Y + +D+WS GCI EL+ + + PG +E++QL I++L G+P
Sbjct: 581 ILTLWYRAPEVLLGATHYSMAVDMWSVGCIFAELVTKQALFPGDSELQQLLHIFRLLGTP 760
Query: 328 SDEYWK-KSKLPNATLFKPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDAL 386
+++ W SKL N + P + + L L+ ++L +P +R +A A+
Sbjct: 761 NEDVWPGVSKLMNWHEYPQWNP--QSLSTAVPSLDELGLDLLSQMLKYEPSKRISAKKAM 934
Query: 387 RSEFF 391
+F
Sbjct: 935 EHVYF 949
>TC219518 similar to UP|Q9FYT8 (Q9FYT8) Cyclin-dependent kinase B1-2,
complete
Length = 1333
Score = 181 bits (459), Expect = 8e-46
Identities = 114/314 (36%), Positives = 171/314 (54%), Gaps = 27/314 (8%)
Frame = +2
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDN----------LEPESIKFMARE 154
+ +EK++K+G+GTY VYKA + +G +VALKK R + E ++ +++
Sbjct: 8 EKYEKLEKVGEGTYGKVYKAREKASGSLVALKKTRLEMDEEGVPPTALREVSLLQLLSQS 187
Query: 155 IIILRRLDHPNVIKLQGLVTSRMSCS-------LYLVFDYMEHDLAGLAAS----PVIR- 202
I I+R L +V K+ S+ S S LYLVF+Y++ DL S P R
Sbjct: 188 IYIVRLLSVEHVDKVP---KSQKSSSNPLTKPILYLVFEYLDTDLKKFIDSHRKGPNPRP 358
Query: 203 FTESQIKCYMNQLLSGLEHCHNRRVLHRDIKGSNLLID-NEGILKIADFGLASFFDPNYM 261
I+ ++ QL G+ HCH+ VLHRD+K NLL+D ++GILKIAD GL F +
Sbjct: 359 LPPPLIQSFLFQLCKGVAHCHSHGVLHRDLKPQNLLLDQHKGILKIADLGLGRAFTVP-L 535
Query: 262 NPMTSRVVTLWYRPPELLLGATDYGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIY 321
T +VTLWYR PE+LLG+T Y G+D+WS GCI E++ + + PG +E +QL I+
Sbjct: 536 KSYTHEIVTLWYRAPEVLLGSTHYSTGVDIWSVGCIFAEMVRRQALFPGDSEFQQLIHIF 715
Query: 322 KLCGSPSDEYWKKSKLPNATLFKPREPYKR----CIRETFKGFPPSALPLIDKLLAIDPV 377
K+ G+P++E W P T + Y R + + P + L+ K+L +P
Sbjct: 716 KMLGTPTEENW-----PGVTSLRDWHVYPRWEPQSLAKNVPSLGPDGVDLLSKMLKYNPS 880
Query: 378 ERETASDALRSEFF 391
ER +A AL +F
Sbjct: 881 ERISAKAALDHPYF 922
>TC218490 similar to UP|Q9LDC1 (Q9LDC1) CRK1 protein, partial (7%)
Length = 689
Score = 179 bits (453), Expect = 4e-45
Identities = 83/110 (75%), Positives = 96/110 (86%)
Frame = +2
Query: 460 NIDRRRLITHANAKSKSEKFPPPHQDGQLGFPLGSSHHIDPDTIPADISFTSTTYTYSKE 519
NIDRRRLITHANA SKSEKFPPPH+DGQLGFPLGSS+HIDPD +P+D+S ST+YT+SKE
Sbjct: 2 NIDRRRLITHANA*SKSEKFPPPHEDGQLGFPLGSSNHIDPDIVPSDVSLGSTSYTFSKE 181
Query: 520 PFQAWSGPIGNAADIGVSKRKKYTAGEAWDLLKPHTGAPKDKFKGKAIIA 569
PF+AWSGPIGN A I V+KRKK+TAG+A DL KP+ GA KDK KGK +A
Sbjct: 182 PFEAWSGPIGNTASISVTKRKKHTAGDALDLSKPYKGALKDKVKGKKYVA 331
>TC226687 homologue to UP|MMK2_MEDSA (Q40353) Mitogen-activated protein
kinase homolog MMK2 , partial (97%)
Length = 1783
Score = 177 bits (448), Expect = 1e-44
Identities = 115/313 (36%), Positives = 169/313 (53%), Gaps = 13/313 (4%)
Frame = +3
Query: 110 IDKIGQGTYSNVYKAIDSMTGKVVALKKV--RFDNLEPESIKFMAREIIILRRLDHPNVI 167
I +G+G Y V A+++ TG+ VA+KK+ FDN K REI +LR +DH N++
Sbjct: 351 IRPVGRGAYGIVCAAVNAETGEEVAIKKIGNAFDNRI--DAKRTLREIKLLRHMDHANIM 524
Query: 168 KLQGLVTSRMSCS---LYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHN 224
++ ++ + +YLV + M+ DL + S + T+ + ++ QLL GL++ H+
Sbjct: 525 SIKDIIRPPQKENFNDVYLVSELMDTDLHQIIRSNQ-QLTDDHCRYFLYQLLRGLKYVHS 701
Query: 225 RRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATD 284
VLHRD+K SNLL++ LKIADFGLA + + MT VVT WYR PELLL ++
Sbjct: 702 ANVLHRDLKPSNLLLNANCDLKIADFGLAR--TTSETDFMTEYVVTRWYRAPELLLNCSE 875
Query: 285 YGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFK 344
Y ID+WS GCILGE++ +P+ PG+ V QL I +L GSP D + NA +
Sbjct: 876 YTAAIDIWSVGCILGEIITRQPLFPGKDYVHQLRLITELIGSPDDASLGFLRSDNARRYV 1055
Query: 345 PREPY--KRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFFT------TEPY 396
+ P K+ F P A+ L++K+L DP R T +AL + + EP
Sbjct: 1056KQLPQYPKQNFSARFPTMSPGAVDLLEKMLIFDPNRRITVDEALSHPYMSPLHDINEEPV 1235
Query: 397 ACDPSSLPKYPPS 409
P S PS
Sbjct: 1236CTRPFSFDFEQPS 1274
>TC204709 similar to UP|Q8H0X4 (Q8H0X4) Serine/threonine-protein kinase Mak
(Male germ cell-associated kinase)-like protein, partial
(59%)
Length = 2140
Score = 170 bits (430), Expect = 2e-42
Identities = 101/288 (35%), Positives = 166/288 (57%), Gaps = 1/288 (0%)
Frame = +2
Query: 105 DTFEKIDKIGQGTYSNVYKAIDSMTGKVVALKKVRFDNLEPESIKFMAREIIILRRLDHP 164
+ ++ I ++G GT+ +V++AI+ TG+VVA+KK++ E + RE+ LR+++HP
Sbjct: 533 ERYKLIKEVGDGTFGSVWRAINKQTGEVVAIKKMKKKYYSWEECVNL-REVKSLRKMNHP 709
Query: 165 NVIKLQGLVTSRMSCSLYLVFDYMEHDLAGLAASPVIRFTESQIKCYMNQLLSGLEHCHN 224
N++KL+ ++ R S LY VF+YME +L L F+E++++ + Q+ GL + H
Sbjct: 710 NIVKLKEVI--RESDILYFVFEYMECNLYQLMKDREKLFSEAEVRNWCFQVFQGLAYMHQ 883
Query: 225 RRVLHRDIKGSNLLIDNEGILKIADFGLASFFDPNYMNPMTSRVVTLWYRPPELLLGATD 284
R HRD+K NLL+ + +KIADFGLA + + P T V T WYR PE+LL +
Sbjct: 884 RGYFHRDLKPENLLVTKD-FIKIADFGLAR--EISSQPPYTEYVSTRWYRAPEVLLQSYM 1054
Query: 285 YGVGIDLWSAGCILGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKS-KLPNATLF 343
Y +D+W+ G I+ EL +P+ PG +E ++++KI + G+P+ E W KL +
Sbjct: 1055YTSKVDMWAMGAIMAELFSLRPLFPGASEADEIYKICGVIGNPTFESWADGLKLARDINY 1234
Query: 344 KPREPYKRCIRETFKGFPPSALPLIDKLLAIDPVERETASDALRSEFF 391
+ + + A+ LI L + DP +R TAS+AL+ FF
Sbjct: 1235QFPQLAGVHLSALIPSASDDAISLITSLCSWDPCKRPTASEALQHPFF 1378
>AW458024
Length = 426
Score = 166 bits (420), Expect = 3e-41
Identities = 77/144 (53%), Positives = 107/144 (73%)
Frame = +2
Query: 298 LGELLVGKPIMPGRTEVEQLHKIYKLCGSPSDEYWKKSKLPNATLFKPREPYKRCIRETF 357
+GEL G+PI+PG+TEVEQLH+I+KLCGSPS++YW+K + P++T+F+P Y++C+ ETF
Sbjct: 5 VGELYCGRPILPGKTEVEQLHRIFKLCGSPSEDYWRKLRTPHSTVFRPPHHYRQCVAETF 184
Query: 358 KGFPPSALPLIDKLLAIDPVERETASDALRSEFFTTEPYACDPSSLPKYPPSKEMDAKRR 417
K P +A LI+ LL++DP R TA+ AL+SEFF++EP CDPSSLPKYPPSKE+D K
Sbjct: 185 KECPSAATRLIETLLSLDPTLRGTATTALKSEFFSSEPLPCDPSSLPKYPPSKEIDTK-- 358
Query: 418 DDEVRRQRAASKAQVDGSKKHRTR 441
+ AS+ DG K+ + R
Sbjct: 359 -----LWKEASRHGADGGKEQKFR 415
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,629,427
Number of Sequences: 63676
Number of extensions: 375530
Number of successful extensions: 3109
Number of sequences better than 10.0: 726
Number of HSP's better than 10.0 without gapping: 2818
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2853
length of query: 569
length of database: 12,639,632
effective HSP length: 102
effective length of query: 467
effective length of database: 6,144,680
effective search space: 2869565560
effective search space used: 2869565560
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC130801.5


