

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130801.12 + phase: 0
(124 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
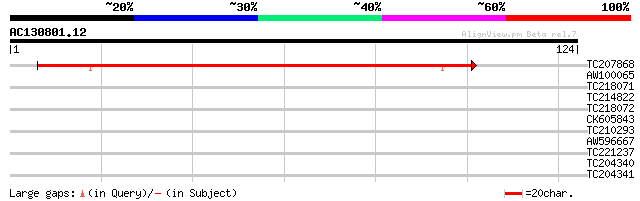
Score E
Sequences producing significant alignments: (bits) Value
TC207868 similar to UP|Q6NLY1 (Q6NLY1) At3g06350, partial (81%) 96 3e-21
AW100065 30 0.073
TC218071 similar to UP|Q01979 (Q01979) Proline-rich protein, par... 27 1.8
TC214822 UP|Q8S3F8 (Q8S3F8) S-adenosylmethionine decarboxylase, ... 27 1.8
TC218072 weakly similar to UP|APG_BRANA (P40603) Anter-specific ... 27 3.2
CK605843 26 5.4
TC210293 weakly similar to UP|Q9LW35 (Q9LW35) Alcohol dehydrogen... 26 5.4
AW596667 weakly similar to GP|27464179|gb expansin {Glycine max}... 26 5.4
TC221237 25 7.0
TC204340 homologue to UP|TCMO_SOYBN (Q42797) Trans-cinnamate 4-m... 25 9.2
TC204341 UP|TCMO_SOYBN (Q42797) Trans-cinnamate 4-monooxygenase ... 25 9.2
>TC207868 similar to UP|Q6NLY1 (Q6NLY1) At3g06350, partial (81%)
Length = 1860
Score = 96.3 bits (238), Expect = 3e-21
Identities = 53/103 (51%), Positives = 68/103 (65%), Gaps = 7/103 (6%)
Frame = +2
Query: 7 VHPLDGRLFV--RAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLACAVFGEVQSYKNLV 64
+ PL G+LFV AGGAGKALA+GAK +GAR+VI + +D +R LA A+ G+ + +L
Sbjct: 1148 ISPLAGKLFVVIGAGGAGKALAYGAKAKGARVVIANRTYDHARKLAYAIGGDALALADLD 1327
Query: 65 NFQPVKGAILANATPIGMHPDTDRIPVAE-----YQLVFDAVY 102
N+ P G ILAN T IGM P D PV++ Y LVFDAVY
Sbjct: 1328 NYHPEDGMILANTTSIGMQPKVDETPVSKHALKYYSLVFDAVY 1456
>AW100065
Length = 354
Score = 30.4 bits (67), Expect(2) = 0.073
Identities = 17/23 (73%), Positives = 18/23 (77%), Gaps = 2/23 (8%)
Frame = +3
Query: 9 PLDGRLFVR--AGGAGKALAFGA 29
PL GRLFV AGGAG ALAFG+
Sbjct: 285 PLAGRLFVLVGAGGAGIALAFGS 353
Score = 20.4 bits (41), Expect(2) = 0.073
Identities = 8/10 (80%), Positives = 9/10 (90%)
Frame = +1
Query: 1 MVKHLQVHPL 10
MV+HL VHPL
Sbjct: 262 MVEHL*VHPL 291
>TC218071 similar to UP|Q01979 (Q01979) Proline-rich protein, partial (6%)
Length = 791
Score = 27.3 bits (59), Expect = 1.8
Identities = 11/30 (36%), Positives = 17/30 (56%)
Frame = +2
Query: 20 GAGKALAFGAKTRGARIVIFDIDFDKSRSL 49
G G AFG+K R +R+ +DFD+ +
Sbjct: 194 GWGFGAAFGSKYRSSRVTFQGVDFDRKEKV 283
>TC214822 UP|Q8S3F8 (Q8S3F8) S-adenosylmethionine decarboxylase, complete
Length = 1826
Score = 27.3 bits (59), Expect = 1.8
Identities = 9/23 (39%), Positives = 17/23 (73%)
Frame = -3
Query: 102 YLHFYIKLGGSKAVLFPSFWCST 124
+ HF + LG ++++FP+FW +T
Sbjct: 1611 FQHFRVDLGEPQSLVFPNFW*TT 1543
>TC218072 weakly similar to UP|APG_BRANA (P40603) Anter-specific proline-rich
protein APG (Protein CEX) (Fragment), partial (10%)
Length = 708
Score = 26.6 bits (57), Expect = 3.2
Identities = 11/30 (36%), Positives = 16/30 (52%)
Frame = +3
Query: 20 GAGKALAFGAKTRGARIVIFDIDFDKSRSL 49
G G AFG+K R +R+ +DFD +
Sbjct: 174 GWGFGTAFGSKYRSSRVTFQGVDFDSKEKV 263
>CK605843
Length = 382
Score = 25.8 bits (55), Expect = 5.4
Identities = 12/40 (30%), Positives = 22/40 (55%)
Frame = +2
Query: 12 GRLFVRAGGAGKALAFGAKTRGARIVIFDIDFDKSRSLAC 51
G+ + +GGA + L+ G R ++V ++ F + SL C
Sbjct: 263 GQGSISSGGASQCLSAGENERREQLVPINVHFFHALSLCC 382
>TC210293 weakly similar to UP|Q9LW35 (Q9LW35) Alcohol dehydrogenase-like
protein, partial (81%)
Length = 1182
Score = 25.8 bits (55), Expect = 5.4
Identities = 13/33 (39%), Positives = 18/33 (54%)
Frame = +3
Query: 18 AGGAGKALAFGAKTRGARIVIFDIDFDKSRSLA 50
A G GKA A GA+++I DID + + A
Sbjct: 303 ASGIGKATATKFINNGAKVIIADIDQELGQETA 401
>AW596667 weakly similar to GP|27464179|gb expansin {Glycine max}, partial
(40%)
Length = 316
Score = 25.8 bits (55), Expect = 5.4
Identities = 12/30 (40%), Positives = 16/30 (53%)
Frame = +2
Query: 49 LACAVFGEVQSYKNLVNFQPVKGAILANAT 78
L+C F E+ Y+NL P +IL AT
Sbjct: 227 LSCGPFFEILCYQNLRKCNPTNPSILITAT 316
>TC221237
Length = 902
Score = 25.4 bits (54), Expect = 7.0
Identities = 15/31 (48%), Positives = 19/31 (60%), Gaps = 4/31 (12%)
Frame = -1
Query: 98 FDAVYLHF---YIK-LGGSKAVLFPSFWCST 124
FD + LHF + K +G SKAVL SF C +
Sbjct: 650 FDKIVLHFVSIFSK*VGSSKAVLHNSFGCKS 558
>TC204340 homologue to UP|TCMO_SOYBN (Q42797) Trans-cinnamate 4-monooxygenase
(Cinnamic acid 4-hydroxylase) (CA4H) (C4H) (P450C4H)
(Cytochrome P450 73) , partial (34%)
Length = 598
Score = 25.0 bits (53), Expect = 9.2
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = +1
Query: 23 KALAFGAKTRGARIVIFDIDFDKSRSLACAVFGE 56
+ + FG++TR V+FDI K + + V+GE
Sbjct: 319 QGVEFGSRTRN---VVFDIFTGKGQDMVFTVYGE 411
>TC204341 UP|TCMO_SOYBN (Q42797) Trans-cinnamate 4-monooxygenase (Cinnamic
acid 4-hydroxylase) (CA4H) (C4H) (P450C4H) (Cytochrome
P450 73) , complete
Length = 1851
Score = 25.0 bits (53), Expect = 9.2
Identities = 12/34 (35%), Positives = 20/34 (58%)
Frame = +3
Query: 23 KALAFGAKTRGARIVIFDIDFDKSRSLACAVFGE 56
+ + FG++TR V+FDI K + + V+GE
Sbjct: 330 QGVEFGSRTRN---VVFDIFTGKGQDMVFTVYGE 422
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.143 0.438
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,990,343
Number of Sequences: 63676
Number of extensions: 50869
Number of successful extensions: 288
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 287
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 287
length of query: 124
length of database: 12,639,632
effective HSP length: 85
effective length of query: 39
effective length of database: 7,227,172
effective search space: 281859708
effective search space used: 281859708
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 53 (25.0 bits)
Medicago: description of AC130801.12


