

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC130275.7 - phase: 0 /pseudo
(833 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
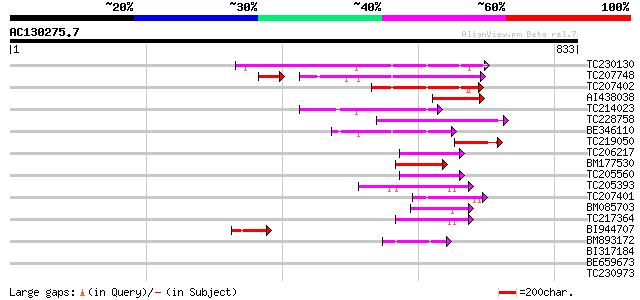
Score E
Sequences producing significant alignments: (bits) Value
TC230130 similar to UP|AHM7_ARATH (Q9SH30) Potential copper-tran... 162 6e-40
TC207748 similar to UP|AHM1_ARATH (Q9M3H5) Potential cadmium/zin... 109 3e-31
TC207402 similar to UP|Q941L1 (Q941L1) Copper-transporting P-typ... 96 9e-20
AI438038 94 2e-19
TC214023 similar to GB|BAA23769.1|2668492|D89981 metal-transport... 94 3e-19
TC228758 similar to UP|Q7Y051 (Q7Y051) Paa2 P-type ATPase, parti... 79 6e-15
BE346110 weakly similar to PIR|B96660|B9 protein F2K11.18 [impor... 69 7e-12
TC219050 similar to UP|AHM2_ARATH (O64474) Potential cadmium/zin... 66 7e-11
TC206217 similar to UP|ACA9_ARATH (Q9LU41) Potential calcium-tra... 65 2e-10
BM177530 homologue to GP|16508164|g type IIB calcium ATPase {Med... 64 4e-10
TC205560 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transpor... 63 5e-10
TC205393 UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-ATPase, partial... 62 1e-09
TC207401 similar to UP|Q941L1 (Q941L1) Copper-transporting P-typ... 61 2e-09
BM085703 homologue to SP|Q9LIK7|ACA Potential calcium-transporti... 59 1e-08
TC217364 plasma membrane Ca2+-ATPase 55 1e-07
BI944707 similar to PIR|B96660|B9 protein F2K11.18 [imported] - ... 47 5e-05
BM893172 PIR|T52414|T5 H+-exporting ATPase (EC 3.6.3.6) plasma ... 44 3e-04
BI317184 similar to GP|13374852|emb metal-transporting ATPase-li... 42 0.001
BE659673 40 0.004
TC230973 homologue to UP|Q7Y068 (Q7Y068) Plasma membrane H+-ATPa... 39 0.007
>TC230130 similar to UP|AHM7_ARATH (Q9SH30) Potential copper-transporting
ATPase 3 , partial (43%)
Length = 1409
Score = 162 bits (410), Expect = 6e-40
Identities = 115/393 (29%), Positives = 203/393 (51%), Gaps = 20/393 (5%)
Frame = +1
Query: 333 VPDMEPWFHLALV----VLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLETLSR 388
+P F LAL V++ CPCAL L+TP A+ A G+L+KGG LE+ +
Sbjct: 76 IPSSMDSFELALQFGISVMVIACPCALGLATPTAVMVGTGVGASQGVLIKGGQALESAHK 255
Query: 389 IKTVAFDKTGTITRGEFSVTDFSAGDDINNETLLYWISSIESKSSHPVAGALVDYARLHS 448
+ + FDKTGT+T G+ + + + +++ E S HP+A A+V+YA+
Sbjct: 256 VDCIVFDKTGTLTVGKPVIVRTELLTKMVLQEFYELVAAAEVNSEHPLAKAVVEYAKRFR 435
Query: 449 IKPVPE--NVENFQNFPGEGIFGTIDGRDIYIGNKRV-GVRAICKRDNCEVQFQRPEIST 505
+ P +F + G G+ ++ ++I +GNK + I D+ E E
Sbjct: 436 DEENPSWPEARDFVSITGHGVKASVHNKEIIVGNKSLFADHNIAIPDDAEYILAEAEKMA 615
Query: 506 KKN---NCEETLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKN 562
+ + + GV ++ D + GA E + LK + ++S+M+TGD+ A + ++
Sbjct: 616 QTGIVVSINGKVAGVLAVSDPLKPGAQEVISILKSMKIKSIMVTGDNFGTASSIAREV-- 789
Query: 563 AIDIVHAELLPHEKAKLIENFKKEG-PIAMIGDGINDAPALATADIGISMGISGSALANE 621
I+ V AE P +KA+ +++ + G +AM+GDGIND+PAL AD+G+++G +G+ +A E
Sbjct: 790 GIENVIAEAKPDQKAEKVKDLQASGYTVAMVGDGINDSPALVAADVGMAIG-AGTDIAIE 966
Query: 622 TSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLV------- 674
++ +LM +++ V AI L+RKT ++ N ++G+ +L + IA L
Sbjct: 967 AADIVLMKSNLEDVITAIDLSRKTFSRIRLNYFWALGYN--LLGIPIAAGALFPSTRFRL 1140
Query: 675 --WLAVLTDVGTCLLVILNSMLILQENHKYEKR 705
W+A + + V+ S+L+ KY +R
Sbjct: 1141PPWIAGAAMAASSVSVVCCSLLL-----KYYRR 1224
>TC207748 similar to UP|AHM1_ARATH (Q9M3H5) Potential
cadmium/zinc-transporting ATPase HMA1 , partial (43%)
Length = 1479
Score = 109 bits (273), Expect(2) = 3e-31
Identities = 82/292 (28%), Positives = 142/292 (48%), Gaps = 19/292 (6%)
Frame = +1
Query: 426 SSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNKRVGV 485
+++E ++HP+ A+VD++ + + +VE+F+ FPG G+ T++ + G ++
Sbjct: 241 AAMEKGTTHPIGRAVVDHSEGKDLPSI--SVESFEYFPGRGLTATVNSIESGTGGAKLLK 414
Query: 486 RAICKRDN----CEVQFQRPEISTKKNNCE------------ETLVGVFSLVDACRSGAL 529
++ D C+ + + +I N V + L D R G
Sbjct: 415 ASLGSIDFITSFCQSEVESEKIKEAVNTSSYGSEYVHAALSVNQKVTLIHLEDRPRPGVS 594
Query: 530 EAMEELK-LLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKE-- 586
++EL+ R +MLTGD A V S + I+ H L P +K +++ ++
Sbjct: 595 NVIQELQDEAKFRVMMLTGDHESSARRVASAV--GINEFHCNLKPEDKLSHVKDISRDMG 768
Query: 587 GPIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTT 646
G + M+G+GINDAPALA A +GI + SA A ++ +L+ I VP I +R+TT
Sbjct: 769 GGLIMVGEGINDAPALAAATVGIVLAHRASATAIAVADVLLLRESISAVPFCIAKSRQTT 948
Query: 647 RKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQE 698
+ +NV +++ ++ G+ +WL VL G LLV LNS+ L E
Sbjct: 949 SLIKQNVALALTSILMASLPSVLGFLPLWLTVLLHEGGTLLVCLNSVRALNE 1104
Score = 45.1 bits (105), Expect(2) = 3e-31
Identities = 21/38 (55%), Positives = 27/38 (70%)
Frame = +2
Query: 366 ALTKAAISGLLLKGGDYLETLSRIKTVAFDKTGTITRG 403
A++ A G+LLKGG L+ L+ TVAFDKTGT+T G
Sbjct: 14 AISSCARKGILLKGGHVLDALATCHTVAFDKTGTLTTG 127
>TC207402 similar to UP|Q941L1 (Q941L1) Copper-transporting P-type ATPase,
partial (19%)
Length = 721
Score = 95.5 bits (236), Expect = 9e-20
Identities = 61/174 (35%), Positives = 108/174 (62%), Gaps = 10/174 (5%)
Frame = +1
Query: 532 MEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPIA- 590
+E L+ +GV VM+TGD+ + A V ++ I V AE++P KA ++ +F+K+G IA
Sbjct: 13 IEGLQKMGVTPVMVTGDNWRTARAVAKEV--GIQDVRAEVMPAGKADVVRSFQKDGSIAA 186
Query: 591 MIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLV 650
M+GDGIND+PALA AD+G+++G +G+ +A E + +LM N++ V AI L+RKT ++
Sbjct: 187 MVGDGINDSPALAAADVGMAIG-AGTDIAIEAAEYVLMRNNLEDVITAIDLSRKTFSRIR 363
Query: 651 ENVIISVGFKCAILALAIAG---YPLV------WLAVLTDVGTCLLVILNSMLI 695
N + ++ + ++A+ +A YP + W+A + + V+ +S+L+
Sbjct: 364 LNYVFAMAYN--VVAIPVAAGVFYPSLGIKLPPWVAGACMALSSVSVVCSSLLL 519
>AI438038
Length = 232
Score = 94.4 bits (233), Expect = 2e-19
Identities = 47/76 (61%), Positives = 59/76 (76%)
Frame = +2
Query: 622 TSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTD 681
T N I+MSNDI K+PEAI+LAR RK+VE +++S+ K AIL LAI G+ LVW AV D
Sbjct: 2 TGNIIIMSNDIMKIPEAIKLARNARRKVVEYIVLSIMTKAAILELAIDGHSLVWTAVEAD 181
Query: 682 VGTCLLVILNSMLILQ 697
VGTCLL+I NSML+L+
Sbjct: 182 VGTCLLLISNSMLLLR 229
>TC214023 similar to GB|BAA23769.1|2668492|D89981 metal-transporting P-type
ATPase {Arabidopsis thaliana;} , partial (21%)
Length = 828
Score = 93.6 bits (231), Expect = 3e-19
Identities = 66/215 (30%), Positives = 106/215 (48%), Gaps = 5/215 (2%)
Frame = +2
Query: 426 SSIESKSSHPVAGALVDYARLHSIKPVPENVENFQNFPGEGIFGTIDGRDIYIGNKRVGV 485
+++E+ S HPV A+VD A+ + F PG G TI + + +G +
Sbjct: 26 AAVETNSVHPVGKAIVDAAQAANCHNAKVKDGTFLEEPGSGAVATIYDKKVSVGT----L 193
Query: 486 RAICKRDNCEVQFQRPEISTKKN----NCEETLVGVFSLVDACRSGALEAMEELKLLGVR 541
I + Q E S ++ ++TL G+ D R A + ++ L +
Sbjct: 194 EWITRHGVINSIHQEVEKSNNQSFVYVGVDDTLAGLIYFEDEIREDARDVVDRLSKQNIG 373
Query: 542 SVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPI-AMIGDGINDAP 600
ML+GD A +V S + + V +E+ P EK K I +K+ I AM+GDGINDA
Sbjct: 374 VYMLSGDKRNAAEHVASLVGIPKEKVLSEVKPDEKKKFINELQKDNNIVAMVGDGINDAA 553
Query: 601 ALATADIGISMGISGSALANETSNAILMSNDIRKV 635
ALA++ +GI++G G A+E S+ +LM N + +V
Sbjct: 554 ALASSHVGIALG-GGVGAASEVSSIVLMRNQLSQV 655
>TC228758 similar to UP|Q7Y051 (Q7Y051) Paa2 P-type ATPase, partial (19%)
Length = 845
Score = 79.3 bits (194), Expect = 6e-15
Identities = 55/195 (28%), Positives = 102/195 (52%), Gaps = 2/195 (1%)
Frame = +3
Query: 540 VRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGP-IAMIGDGIND 598
+++V+L+GD + V + D V A L P +K+ I + K G +AM+GDGIND
Sbjct: 9 IKTVLLSGDREEAVATVADTVGIENDFVKASLSPQQKSGFISSLKAAGHHVAMVGDGIND 188
Query: 599 APALATADIGISM-GISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVENVIISV 657
AP+LA AD+GI++ + A++ ++ IL+ N I +V +A+ LA+ T K+ +N+ +V
Sbjct: 189 APSLAVADVGIALQNEAQENAASDAASIILLGNKISQVVDALDLAQATMGKVYQNLSWAV 368
Query: 658 GFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTKGSKYGKFL 717
+ + +A + +T + L+ L+S+ ++ + + S K G +
Sbjct: 369 AYNVVAIPIAAGVLLPHFDFAMTPSLSGGLMALSSIFVVGNSLLLQLHGSQISRKVGSTI 548
Query: 718 EDKTKPLLNKQSNID 732
E +++ SN D
Sbjct: 549 E-----IISSHSNTD 578
>BE346110 weakly similar to PIR|B96660|B9 protein F2K11.18 [imported] -
Arabidopsis thaliana, partial (12%)
Length = 562
Score = 69.3 bits (168), Expect = 7e-12
Identities = 53/190 (27%), Positives = 95/190 (49%), Gaps = 7/190 (3%)
Frame = +2
Query: 474 RDIYIGNKRVGVRAICKRDNC-EVQFQRPEISTKKNNC-----EETLVGVFSLVDACRSG 527
R + +GNKR+ C C EV+ E C + + G FS+ D +
Sbjct: 8 RTVVVGNKRL--MHACNVPICSEVEKYISENEILATTCIVVSIDGKIAGAFSVTDPVKPE 181
Query: 528 ALEAMEELKLLGVRSVMLTGDSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEG 587
A + L +G+ S+++TGD+ A + +++ ID V AE P KA +++ + G
Sbjct: 182 AKRVIFFLHSMGISSMIVTGDNCATATAIANEM--GIDEVFAETDPEGKADKVKDLQMTG 355
Query: 588 -PIAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTT 646
+AM+GDG+ND+P L AD+G++ G +G+ +A + + V I ++R T
Sbjct: 356 MTVAMVGDGMNDSPDLLAADVGMATG-AGTDIAIAAAVIASVKRRFEDVIIDIDISRTTM 532
Query: 647 RKLVENVIIS 656
+++ N I++
Sbjct: 533 SRILLNYILA 562
>TC219050 similar to UP|AHM2_ARATH (O64474) Potential
cadmium/zinc-transporting ATPase 2 , partial (4%)
Length = 589
Score = 65.9 bits (159), Expect = 7e-11
Identities = 35/70 (50%), Positives = 43/70 (61%)
Frame = +1
Query: 654 IISVGFKCAILALAIAGYPLVWLAVLTDVGTCLLVILNSMLILQENHKYEKRESTKGSKY 713
+ S+ K AIL LAI G+PLVW AV+ DVGTCLLVI NSML+L++ H +
Sbjct: 1 VFSIMTKAAILDLAIGGHPLVWAAVVADVGTCLLVIFNSMLLLRKGHNHG---------- 150
Query: 714 GKFLEDKTKP 723
GK TKP
Sbjct: 151 GKCCRSSTKP 180
>TC206217 similar to UP|ACA9_ARATH (Q9LU41) Potential calcium-transporting
ATPase 9, plasma membrane-type (Ca(2+)-ATPase isoform
9) , partial (32%)
Length = 1383
Score = 64.7 bits (156), Expect = 2e-10
Identities = 33/97 (34%), Positives = 57/97 (58%), Gaps = 1/97 (1%)
Frame = +1
Query: 573 PHEKAKLIENFKKEGPI-AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSND 631
P++K L++ +K G + A+ GDG NDAPAL ADIG+SMGI G+ +A E+S+ I++ ++
Sbjct: 136 PNDKLLLVQALRKGGEVVAVTGDGTNDAPALHEADIGLSMGIQGTEVAKESSDIIILDDN 315
Query: 632 IRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAI 668
V + +R R + + + + A L + +
Sbjct: 316 FASVVKVVRWGRSVYANIQKFIQFQLTVNVAALVINV 426
>BM177530 homologue to GP|16508164|g type IIB calcium ATPase {Medicago
truncatula}, partial (13%)
Length = 422
Score = 63.5 bits (153), Expect = 4e-10
Identities = 32/78 (41%), Positives = 50/78 (64%), Gaps = 1/78 (1%)
Frame = +2
Query: 567 VHAELLPHEKAKLIENFKKEGPI-AMIGDGINDAPALATADIGISMGISGSALANETSNA 625
V A P +K +++ KK+G + A+ GDG NDAPAL ADIG+SMGI G+ +A E+S+
Sbjct: 20 VMARSSPLDKLLMVQCLKKKGHVVAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDI 199
Query: 626 ILMSNDIRKVPEAIRLAR 643
+++ ++ V +R R
Sbjct: 200 VILDDNFNSVATVLRWGR 253
>TC205560 similar to GB|BAA97361.1|8843813|AB023042 Ca2+-transporting
ATPase-like protein {Arabidopsis thaliana;} , partial
(27%)
Length = 1356
Score = 63.2 bits (152), Expect = 5e-10
Identities = 31/97 (31%), Positives = 58/97 (58%), Gaps = 1/97 (1%)
Frame = +1
Query: 573 PHEKAKLIENFKKEGPI-AMIGDGINDAPALATADIGISMGISGSALANETSNAILMSND 631
P++K L++ +++G + A+ GDG NDAPAL ADIG++MGI G+ +A E+S+ I++ ++
Sbjct: 28 PNDKLLLVQALRRKGHVVAVTGDGTNDAPALHEADIGLAMGIQGTEVAKESSDIIILDDN 207
Query: 632 IRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAI 668
V + +R R + + + + A L + +
Sbjct: 208 FASVVKVVRWGRSVYANIQKFIQFQLTVNVAALVINV 318
>TC205393 UP|Q9FVE8 (Q9FVE8) Plasma membrane Ca2+-ATPase, partial (55%)
Length = 2009
Score = 62.0 bits (149), Expect = 1e-09
Identities = 52/207 (25%), Positives = 95/207 (45%), Gaps = 38/207 (18%)
Frame = +2
Query: 513 TLVGVFSLVDACRSGALEAMEELKLLGVRSVMLTGDSSQVAMYV---------------- 556
T +GV + D R G E++ + G+ M+TGD+ A +
Sbjct: 578 TCIGVIGIKDPVRPGVKESVAM*RSAGITVRMVTGDNINTAKAIARECGILTDDGIAIEG 757
Query: 557 ----QSQLKNAIDI-----VHAELLPHEKAKLIENFKKE--GPIAMIGDGINDAPALATA 605
+ K +++ V A P +K L+++ + +A+ GDG NDAPAL A
Sbjct: 758 PEFREKSQKELLELIPKIQVMARSSPLDKHTLVKHLRTTFGEVVAVTGDGTNDAPALHEA 937
Query: 606 DIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARK---TTRKLVE-----NVI-IS 656
DIG++MGI+G+ +A E+++ I++ ++ + + R +K V+ NV+ +
Sbjct: 938 DIGLAMGIAGTEVAKESADVIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTVNVVALI 1117
Query: 657 VGFKCAIL--ALAIAGYPLVWLAVLTD 681
V F A L + L+W+ ++ D
Sbjct: 1118VNFTSACLTGTAPLTAVQLLWVNMIMD 1198
>TC207401 similar to UP|Q941L1 (Q941L1) Copper-transporting P-type ATPase,
partial (12%)
Length = 532
Score = 60.8 bits (146), Expect = 2e-09
Identities = 38/116 (32%), Positives = 68/116 (57%), Gaps = 7/116 (6%)
Frame = +1
Query: 593 GDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKLVEN 652
GDGIND+PALA AD+G+++G +G+ +A E +N +LM +++ V AI L++KT ++ N
Sbjct: 7 GDGINDSPALAAADVGMAIG-AGTDVAIEAANYVLMRDNLEDVITAIDLSKKTFFRIRLN 183
Query: 653 VIISVGFKCAILALAIAGYPLVWLAVLTD---VGTCLLV----ILNSMLILQENHK 701
+ ++ + + +A AG WL + G C+ + ++ S L+L+ K
Sbjct: 184 YVFAMAYNVVAIPVA-AGVFFPWLGIKLPPWVAGACMALSSVSVVCSSLLLRRYRK 348
>BM085703 homologue to SP|Q9LIK7|ACA Potential calcium-transporting ATPase 13
plasma membrane-type (EC 3.6.3.8) (Ca(2+)-ATPase
isoform, partial (12%)
Length = 411
Score = 58.5 bits (140), Expect = 1e-08
Identities = 32/104 (30%), Positives = 56/104 (53%), Gaps = 11/104 (10%)
Frame = +3
Query: 589 IAMIGDGINDAPALATADIGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRK 648
+A+ GDG NDAPAL ADIG+SMGI G+ +A E+S+ +++ ++ V +R R
Sbjct: 15 VAVTGDGTNDAPALKEADIGLSMGIQGTEVAKESSDIVILDDNFASVVTVLRWGRCVYNN 194
Query: 649 L---------VENVIISVGFKCAILA--LAIAGYPLVWLAVLTD 681
+ V +++ F A+ A + + L+W+ ++ D
Sbjct: 195 IQKFIQFQLTVNVAALAINFVAAVSAGKVPLTAVQLLWVNLIMD 326
>TC217364 plasma membrane Ca2+-ATPase
Length = 1375
Score = 55.1 bits (131), Expect = 1e-07
Identities = 37/128 (28%), Positives = 69/128 (53%), Gaps = 13/128 (10%)
Frame = +3
Query: 567 VHAELLPHEKAKLIENFKK--EGPIAMIGDGINDAPALATADIGISMGISGSALANETSN 624
V A P +K L+++ + + +++ GDG NDAPAL ADIG++MGI+G+ +A E+++
Sbjct: 84 VMARSSPMDKHTLVKHLRTTFQEVVSVTGDGTNDAPALHEADIGLAMGIAGTEVAKESAD 263
Query: 625 AILMSNDIRKVPEAIRLARK---TTRKLVE-----NVI-ISVGFKCAILA--LAIAGYPL 673
I++ ++ + + R +K V+ NV+ + V F A L + L
Sbjct: 264 VIILDDNFSTIVTVAKWGRSVYINIQKFVQFQLTVNVVALIVNFSSACLTGNAPLTAVQL 443
Query: 674 VWLAVLTD 681
+W+ ++ D
Sbjct: 444 LWVNMIMD 467
>BI944707 similar to PIR|B96660|B9 protein F2K11.18 [imported] - Arabidopsis
thaliana, partial (9%)
Length = 362
Score = 46.6 bits (109), Expect = 5e-05
Identities = 25/58 (43%), Positives = 35/58 (60%)
Frame = +1
Query: 327 VPAALSVPDMEPWFHLALVVLLSGCPCALILSTPVAIFCALTKAAISGLLLKGGDYLE 384
+P A+ ++ F A+ VL+ CPCAL L+TP A+ A A G+L+KGGD LE
Sbjct: 181 IPKAMDAFELALQF--AISVLVVACPCALGLATPTAVMVASGMGASQGVLIKGGDALE 348
>BM893172 PIR|T52414|T5 H+-exporting ATPase (EC 3.6.3.6) plasma membrane
[imported] - Prunus persica, partial (14%)
Length = 427
Score = 43.9 bits (102), Expect = 3e-04
Identities = 30/103 (29%), Positives = 53/103 (51%), Gaps = 1/103 (0%)
Frame = -2
Query: 548 DSSQVAMYVQSQLKNAIDIVHAELLPHEKAKLIENFKKEGPIA-MIGDGINDAPALATAD 606
D S VA+ + ++ A A + P K ++++ + I M GDG+NDAPAL AD
Sbjct: 318 DESIVALPIDELIEKADGF--AGVFPEHKYEIVKRLQARKHICGMTGDGVNDAPALKKAD 145
Query: 607 IGISMGISGSALANETSNAILMSNDIRKVPEAIRLARKTTRKL 649
IGI++ + A A S+ +L + + A+ +R +++
Sbjct: 144 IGIAVADATDA-ARSASDIVLTEPGLSVIISAVLTSRAIFQRM 19
>BI317184 similar to GP|13374852|emb metal-transporting ATPase-like protein
{Arabidopsis thaliana}, partial (4%)
Length = 407
Score = 41.6 bits (96), Expect = 0.001
Identities = 20/46 (43%), Positives = 31/46 (66%), Gaps = 1/46 (2%)
Frame = +1
Query: 588 PIAMIGDGINDAPALATADIGISM-GISGSALANETSNAILMSNDI 632
P +GDGINDAP+LA AD+GI++ + A++ ++ IL+ N I
Sbjct: 268 PFIKVGDGINDAPSLAVADVGIALQNEAQENAASDAASIILLGNKI 405
>BE659673
Length = 545
Score = 40.0 bits (92), Expect = 0.004
Identities = 24/73 (32%), Positives = 38/73 (51%)
Frame = -2
Query: 626 ILMSNDIRKVPEAIRLARKTTRKLVENVIISVGFKCAILALAIAGYPLVWLAVLTDVGTC 685
+L+ I VP I +R+TT + +NV +++ ++ G+ +WL VL G
Sbjct: 529 LLLRESISAVPFCIAKSRQTTSLIKQNVALALTSILMASLPSVLGFLPLWLTVLLHEGGT 350
Query: 686 LLVILNSMLILQE 698
LLV LNS+ L E
Sbjct: 349 LLVCLNSVRALNE 311
>TC230973 homologue to UP|Q7Y068 (Q7Y068) Plasma membrane H+-ATPase, partial
(22%)
Length = 654
Score = 39.3 bits (90), Expect = 0.007
Identities = 19/44 (43%), Positives = 29/44 (65%), Gaps = 1/44 (2%)
Frame = +3
Query: 569 AELLPHEKAKLIENFKKEGPIA-MIGDGINDAPALATADIGISM 611
A + P K ++++ ++ I M GDG+NDAPAL ADIGI++
Sbjct: 483 AGVFPEHKYEIVKRLQERKHICGMTGDGVNDAPALKKADIGIAV 614
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.137 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 34,003,217
Number of Sequences: 63676
Number of extensions: 437077
Number of successful extensions: 2083
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 2066
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2077
length of query: 833
length of database: 12,639,632
effective HSP length: 105
effective length of query: 728
effective length of database: 5,953,652
effective search space: 4334258656
effective search space used: 4334258656
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 63 (28.9 bits)
Medicago: description of AC130275.7


